siRNA in VIGS: From Molecular Mechanisms to Therapeutic Applications
This article comprehensively examines the critical role of small interfering RNAs (siRNAs) in Virus-Induced Gene Silencing (VIGS), an RNAi-based technology revolutionizing functional genomics and therapeutic development.
siRNA in VIGS: From Molecular Mechanisms to Therapeutic Applications
Abstract
This article comprehensively examines the critical role of small interfering RNAs (siRNAs) in Virus-Induced Gene Silencing (VIGS), an RNAi-based technology revolutionizing functional genomics and therapeutic development. We explore the foundational mechanisms whereby siRNAs mediate sequence-specific gene silencing through the RNA-induced silencing complex (RISC), detail methodological advances in VIGS vector design and delivery systems, analyze optimization strategies to enhance silencing efficiency while minimizing off-target effects, and provide comparative validation against alternative gene silencing technologies. This synthesis provides researchers, scientists, and drug development professionals with current insights into VIGS applications across plant functional genomics and emerging therapeutic paradigms, highlighting both technical challenges and future translational opportunities.
Molecular Foundations: How siRNA Machinery Powers VIGS
Virus-Induced Gene Silencing (VIGS) is a powerful reverse genetics tool that co-opts the plant's innate RNA interference (RNAi) machinery to silence endogenous genes. This natural antiviral defense mechanism uses sequence-specific small interfering RNAs (siRNAs) to guide the degradation of complementary messenger RNA (mRNA), enabling rapid functional genomics studies without the need for stable transformation. This technical guide elucidates the core RNAi mechanisms underpinning VIGS, details current experimental protocols, and presents quantitative data on its application in crop research, with a specific focus on the central role of siRNA in mediating targeted gene silencing for plant science and biotechnology.
RNA interference (RNAi) is a biologically conserved mechanism that mediates sequence-specific gene silencing at the post-transcriptional level [1] [2]. Initially characterized as a defense system against exogenous nucleic acids like viruses and transposons, this pathway is now recognized as a key regulator of endogenous gene expression [3]. In plants, the discovery that this same machinery could be harnessed to target plant genes led to the development of Virus-Induced Gene Silencing (VIGS), a technique first demonstrated using a modified Tobacco mosaic virus vector to silence the phytoene desaturase (PDS) gene in Nicotiana benthamiana [4]. VIGS functions by engineering viral vectors to carry fragments of host plant genes; when infected, the plant's RNAi machinery processes these viral transcripts into siRNAs that then direct the silencing of the corresponding endogenous mRNA [5] [4]. This technology has become an indispensable tool for functional genomics, particularly in species recalcitrant to stable transformation, allowing researchers to link gene sequences to biological functions through observable phenotypic changes.
The Core RNAi Machinery in Plants
The plant RNAi pathway is a sophisticated, multi-layered defense and regulatory system. Its effectiveness in VIGS relies on the precise orchestration of several key components and sub-pathways.
Key Components and Mechanism
The core RNAi mechanism can be simplified into a two-step enzymatic process [1]:
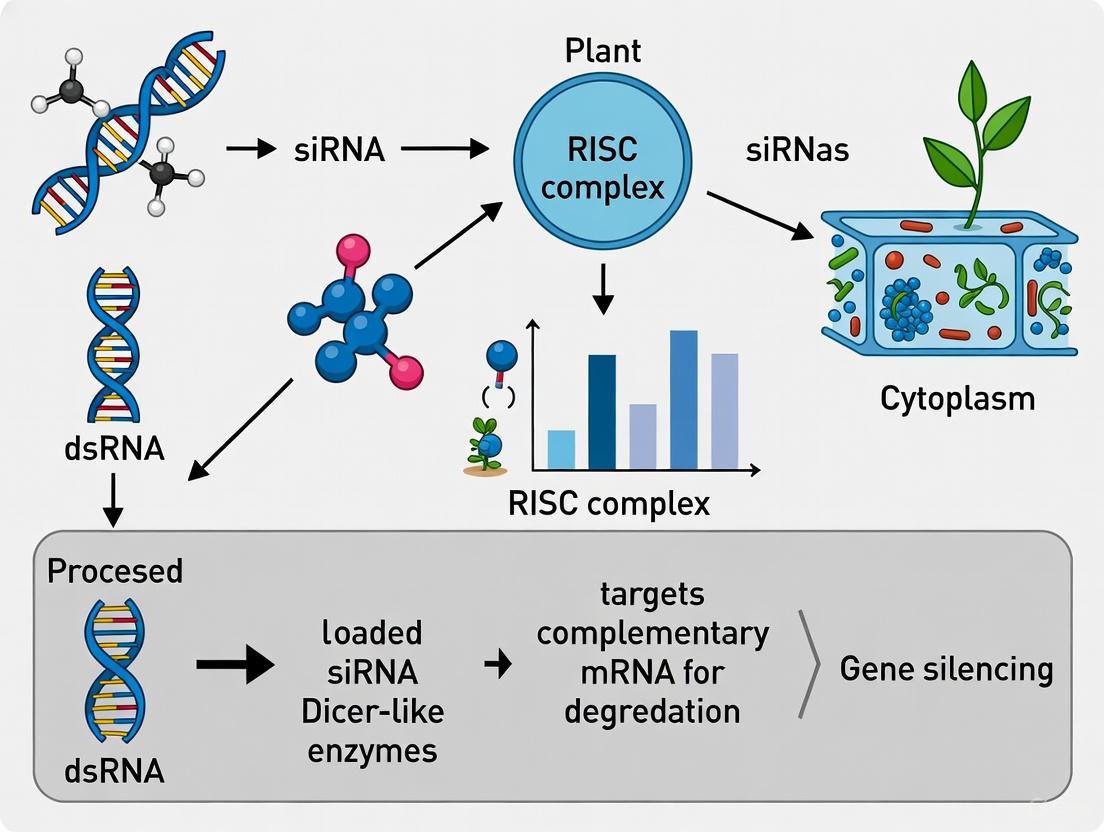
- Initiation and Processing: Double-stranded RNA (dsRNA) triggers, whether of viral origin or from endogenous hairpin transcripts, are recognized and processed by Dicer-like (DCL) ribonuclease enzymes. DCL cleaves long dsRNA molecules into short, 21-24 nucleotide small interfering RNA (siRNA) duplexes with 2-nucleotide 3' overhangs [3] [2].
- Effector Complex Assembly and Silencing: The siRNA duplexes are loaded into the RNA-induced silencing complex (RISC). During RISC assembly, the siRNA passenger strand is degraded, and the guide strand remains to direct RISC to complementary mRNA sequences. The catalytic component of RISC, an Argonaute (AGO) protein, then cleaves the target mRNA, leading to its degradation and consequent gene silencing [1] [3].
The following diagram illustrates the core RNAi pathway that is exploited in VIGS:
Specialized RNA Silencing Pathways
Plants have evolved multiple, overlapping RNA silencing pathways, each with distinct triggers and functional outcomes [3]:
- The microRNA (miRNA) Pathway: miRNAs are derived from endogenous genomic MIR genes transcribed into stem-loop structured precursors. Processed primarily by DCL1, mature miRNAs (typically 21-nt) are incorporated into AGO1 to regulate endogenous gene expression, often targeting transcription factors to control development. While highly specific, they typically act with imperfect complementarity [3] [2].
- The trans-acting siRNA (tasiRNA) Pathway: This pathway links miRNA and siRNA actions. Specific 22-nt miRNAs initiate the process by cleaving non-coding TAS gene transcripts. The cleavage fragments are converted to dsRNA by RDR6 and then processed by DCL4 into 21-nt phased tasiRNAs. These tasiRNAs act in trans to silence target mRNAs, functioning as a robust, amplified silencing signal [3].
- The RNA-directed DNA Methylation (RdDM) Pathway: A plant-specific pathway that leads to transcriptional gene silencing (TGS). It involves 24-nt siRNAs generated by DCL3 from transcripts produced by Pol IV. These siRNAs are loaded into AGO4 and guide de novo DNA methylation to homologous genomic loci, silencing transposons and repetitive elements to maintain genome stability [3] [5].
Table 1: Key Small RNA Pathways in Plants
| Pathway | Small RNA Type | Key Biogenesis Factor | Primary Function | Role in VIGS |
|---|---|---|---|---|
| Exogenic RNAi | siRNA (21-22nt) | DCL2/DCL4, RDR1/6 | Antiviral defense, transgene silencing | Core mechanism: Viral dsRNA processed into siRNAs that direct PTGS of target mRNA [3]. |
| microRNA (miRNA) | miRNA (21-24nt) | DCL1 | Endogenous gene regulation, development | Basis for artificial miRNA (amiRNA) technology; used for highly specific VIGS [6] [3]. |
| trans-acting siRNA (tasiRNA) | tasiRNA (21nt) | DCL4, RDR6 | Amplified silencing of endogenous targets | Basis for syn-tasiR-VIGS; synthetic tasiRNAs from minimal precursors enable potent silencing [7]. |
| RdDM | siRNA (24nt) | DCL3, Pol IV/V | De novo DNA methylation, TGS | Underpins heritable VIGS; siRNAs can trigger epigenetic modifications stably inherited over generations [3] [5]. |
The Molecular Mechanism of VIGS
VIGS cleverly exploits the exogenic RNAi pathway. The process begins with the introduction of a recombinant viral vector carrying a fragment (typically 200-500 base pairs) of the plant gene targeted for silencing.
The Role of siRNA in VIGS
The siRNA is the central effector molecule in the VIGS mechanism. Its production and activity are what make the silencing sequence-specific and systemic. Upon delivery, the viral vector replicates in the host plant, and its dsRNA replication intermediates are recognized by the plant's Dicer-like enzymes as foreign [5]. DCL processes these molecules into a population of 21-24 nucleotide virus-derived siRNAs [4]. A subset of these siRNAs will be complementary to the inserted host gene fragment. These specific siRNAs are then loaded into the RISC. The guide strand of the siRNA directs the RISC to the complementary endogenous mRNA transcript through base-pairing. The AGO protein within RISC then catalyzes the cleavage of the target mRNA, preventing its translation into a functional protein and resulting in a loss-of-function phenotype that can be studied [1] [5]. A key feature of VIGS is the systemic spread of the silencing signal. This is facilitated by the movement of siRNAs themselves, or through the cell-to-cell and long-distance movement of the viral vector, which continuously generates new siRNAs, allowing the silencing effect to reach tissues far from the initial infection site [5] [8].
The integrated workflow below details the step-by-step process from vector design to phenotypic analysis:
Essential VIGS Methodologies and Protocols
The successful application of VIGS depends on robust and reproducible protocols. Below are detailed methodologies for key experimental steps.
Vector Selection and Insert Design
The choice of viral vector is critical and depends on the host plant species.
- Tobacco Rattle Virus (TRV): One of the most widely used vectors, especially in Solanaceous plants like N. benthamiana, tomato, and pepper. Its advantages include efficient systemic movement, mild viral symptoms, and the ability to target meristematic tissues [4]. The TRV system is bipartite, requiring two plasmids: TRV1 (encoding replication and movement proteins) and TRV2 (containing the coat protein and the cloning site for the target insert) [4].
- Cabbage Leaf Curl Virus (CaLCuV): A geminivirus (DNA virus)-based vector suitable for plants like Arabidopsis thaliana and N. benthamiana. It has been successfully engineered to express artificial miRNAs (amiRNAs) for highly specific silencing [6].
- Insert Design Principles: For TRV vectors, a 200-300 bp fragment of the target gene's coding sequence is typically used. Tools like
pssRNAitcan be employed to select fragments with high predicted siRNA density, which may enhance silencing efficiency [8]. The insert should be checked for uniqueness to avoid off-target silencing of homologous genes.
Plant Inoculation Techniques
The delivery method significantly impacts VIGS efficiency and is optimized for the plant species.
- Agrobacterium-Mediated Delivery (Agroinfiltration): The most common method. Recombinant TRV vectors are transformed into Agrobacterium tumefaciens (e.g., strain GV3101). Bacterial cultures are grown to an optimal density (OD600 of 0.5-2.0), resuspended in an induction medium (e.g., with acetosyringone), and incubated for several hours [8] [4]. The suspension is then inoculated into plants.
- Leaf Infiltration: Using a needleless syringe to infiltrate the bacterial suspension into the abaxial side of leaves. Suitable for rosette plants like Arabidopsis and young N. benthamiana [4].
- Stem Injection/Pricking: Using a needle to introduce the agrobacterium into the stem or petiole.
- Vacuum Infiltration: Effective for difficult-to-transform species. A protocol optimized for sunflower involves peeling the seed coat and subjecting the seeds to vacuum infiltration with the agrobacterium suspension, followed by a 6-hour co-cultivation period before sowing. This method achieved infection rates of up to 91% without requiring in vitro culture [8].
- Spray-Based Methods: A transgene-free approach gaining traction. This includes Spray-Induced Gene Silencing (SIGS), where dsRNA targeting a pathogen or pest gene is directly sprayed onto crops [9], and syn-tasiR-VIGS, where infectious viral extracts producing synthetic tasiRNAs are sprayed onto leaves [7].
The Scientist's Toolkit: Essential Research Reagents
Table 2: Key Reagents for VIGS Experiments
| Reagent / Material | Function / Role in VIGS | Example Specifications / Notes |
|---|---|---|
| Viral Vectors | Delivery vehicle for the target gene insert into plant cells. | TRV (pYL192/TRV1, pYL156/TRV2), CaLCuV, CLCrV. Selection depends on host range [6] [4]. |
| Agrobacterium tumefaciens | Biological vector for transferring T-DNA containing viral genomes into the plant cell. | Strain GV3101 is commonly used. Requires appropriate antibiotic resistance (e.g., kanamycin, rifampicin) [8]. |
| Infiltration Buffer | Medium for suspending agrobacterium cells for inoculation. | Typically contains MgCl₂, MES, and acetosyringone (an inducer of Vir genes) [4]. |
| Selection Antibiotics | Maintains plasmid integrity in bacterial cultures. | Kanamycin (for TRV vectors), Gentamicin, Rifampicin (for Agrobacterium strain selection) [8]. |
| Target Gene Insert | Provides the sequence homology for siRNA-guided silencing of the endogenous plant gene. | 100-500 bp fragment, ideally checked for specificity and siRNA prediction [8]. |
| High-Precision Polymerase | Amplifies the target gene insert for cloning with high fidelity. | e.g., Tersus Plus PCR kit [8]. |
| Restriction Enzymes & Ligase | For cloning the target insert into the viral vector's multiple cloning site. | e.g., XbaI and BamHI for TRV2; T4 DNA Ligase [8]. |
Quantitative Data and Applications in Crop Research
VIGS has been successfully applied to characterize genes involved in a wide array of traits, from disease resistance to fruit quality.
Silencing Efficiency and Optimization Parameters
Quantitative data from recent studies highlights key factors for success.
Table 3: Quantitative Factors Influencing VIGS Efficiency
| Factor | Impact on VIGS Efficiency | Experimental Data / Optimal Range |
|---|---|---|
| Plant Genotype | Susceptibility to viral infection and systemic movement varies. | In sunflowers, infection rates ranged from 62% to 91% across 6 genotypes; 'Smart SM-64B' had highest infection but lowest phenotype spread [8]. |
| Agroinoculum Concentration | Affects initial infection load and symptom severity. | Optical density (OD600) typically between 0.5 and 2.0; requires empirical optimization for each species [4]. |
| Inoculation Method | Determines delivery efficiency and scope of initial infection. | Seed vacuum infiltration in sunflowers yielded up to 77% infection and strong silencing (normalized relative expression = 0.01) [8]. |
| Environmental Conditions | Influences plant growth, viral replication, and siRNA amplification. | Temperature (20-22°C), humidity (~45%), and photoperiod (18-h light/6-h dark) are often controlled [8] [4]. |
| Plant Developmental Stage | Younger tissues often show more robust and rapid silencing. | Silencing spreads more actively in young sunflower tissues compared to mature ones [8]. |
Functional Gene Validation in Crops
VIGS has been instrumental in accelerating functional genomics in crops.
- Pepper (Capsicum annuum): VIGS has been used to identify genes governing fruit color (e.g., phytotene desaturase, PDS), pungency (capsaicinoid biosynthesis genes), resistance to bacteria and oomycetes, and tolerance to abiotic stresses like temperature and salt [4].
- Disease Resistance: Silencing the SGT1 gene in N. benthamiana via a geminivirus-based amiRNA system led to the loss of N-mediated resistance to Tobacco mosaic virus, validating SGT1's role in disease resistance [6].
- Crop Protection: Spray-Induced Gene Silencing (SIGS) represents a transgene-free application. For instance, spraying dsRNAs targeting essential genes in fungal pathogens like Botrytis cinerea can significantly inhibit grey mould disease on the surfaces of fruits and vegetables [9].
Advanced VIGS Technologies
The field of VIGS is continuously evolving, with new technologies enhancing its precision and scope.
- Artificial miRNA (amiRNA) VIGS: By modifying endogenous miRNA precursors, researchers can design amiRNAs that target specific genes with high precision and reduced off-target effects compared to long fragment inserts. This "MIR VIGS" approach has been successfully implemented using geminivirus vectors [6].
- syn-tasiR-VIGS: This cutting-edge platform uses synthetic tasiRNA precursors that are significantly shorter than natural TAS genes. These minimal precursors can be expressed from a viral vector and produce highly effective, phased syn-tasiRNAs. This system can be delivered by spraying infectious extracts, creating a potent, transgene-free tool for precision RNAi and even antiviral vaccination in plants [7].
- VIGS for Heritable Epigenetic Modifications: VIGS can be engineered to induce transcriptional gene silencing (TGS) via the RdDM pathway. By targeting viral constructs to gene promoter sequences, researchers can trigger DNA methylation (RdDM) that leads to stable, heritable epigenetic silencing. For example, silencing of the FWA promoter via VIGS in Arabidopsis resulted in transgenerational epigenetic memory of the silent state [5].
Virus-Induced Gene Silencing stands as a testament to how a fundamental biological process—the plant's RNAi-based antiviral defense—can be repurposed as a powerful scientific tool. The centrality of siRNA in this process cannot be overstated; these small RNAs are the sequence-specific guides that direct the silencing machinery to its target, enabling the functional deconvolution of plant genomes. As VIGS technologies evolve—from robust agroinfiltration protocols to advanced, transgene-free spray systems like SIGS and syn-tasiR-VIGS—their impact on functional genomics and crop improvement continues to grow. The integration of VIGS with multi-omics approaches and its ability to induce stable epigenetic changes further promise to accelerate the pace of discovery in plant biology and the development of crops with enhanced agronomic traits.
Small interfering RNA (siRNA) serves as a critical effector molecule in the RNA interference (RNAi) pathway, an evolutionarily conserved mechanism that defends against invasive nucleic acids. This pathway is central to virus-induced gene silencing (VIGS), a process that has revolutionized plant functional genomics and therapeutic development. Within VIGS research, understanding the precise molecular journey from viral infection to gene silencing provides fundamental insights for harnessing this pathway to silence deleterious genes. This technical guide delineates the core biochemical pathway of siRNA biogenesis, detailing the key molecular players, mechanisms, and experimental approaches. We further provide structured data visualization, standardized protocols, and essential research toolkits to equip scientists in leveraging siRNA biogenesis for advanced genetic research and therapeutic development.
RNA interference (RNAi) is a biological process for gene silencing that is initiated by double-stranded RNA (dsRNA) molecules. Discovered in plants and later in Caenorhabditis elegans, RNAi has emerged as a transformative tool for biomedical research and therapeutic development [10] [11]. The core of this pathway involves small non-coding RNAs, of which small interfering RNA (siRNA) is a major class. siRNAs are typically 21 nucleotides in length and are generated from the cleavage of long dsRNA precursors, serving to guide the silencing of complementary target mRNAs [12] [10].
In the context of virus-induced gene silencing (VIGS), the siRNA pathway functions as an adaptive immune defense. When viruses infect a host, they often produce dsRNA intermediates during their replication cycle. The host cell recognizes these dsRNAs as "non-self" and processes them into siRNAs, which then guide the destruction of viral RNA genomes and transcripts, thereby limiting the infection [13]. Researchers have ingeniously co-opted this natural defense mechanism. By engineering viral vectors to carry host gene sequences, infection triggers the production of siRNAs that target the corresponding host mRNA for degradation, leading to specific gene silencing [13]. This principle forms the basis of VIGS, a powerful technique for interrogating gene function without the need for stable transformation.
The Core siRNA Biogenesis Pathway
The transformation of a viral infection signal into a precise gene-silencing command is a multi-step enzymatic process. The pathway can be segmented into three major stages: the initiator phase, the processor phase, and the effector phase.
The Initiator Phase: Recognition of Double-Stranded RNA
The biogenesis of siRNAs begins with the recognition of long double-stranded RNA (dsRNA) molecules. In a natural VIGS context, these dsRNAs are often replicative intermediates formed during viral replication. In experimental applications, dsRNA can be introduced directly or expressed from engineered vectors. This dsRNA acts as the foundational substrate, signaling the activation of the cell's RNAi machinery [10].
The Processor Phase: Dicing by Dicer
The key enzyme responsible for processing long dsRNA into siRNAs is Dicer, an RNase III-like endonuclease. Dicer cleaves the dsRNA into short fragments of 21-23 base pairs in length. Characteristically, the resulting siRNA duplexes possess 2-nucleotide overhangs on their 3' ends and 5'-phosphate groups [10]. This specific structure is a molecular signature that licenses the siRNA for entry into the subsequent silencing complex.
The Effector Phase: RISC Assembly and Slicer Activity
The siRNA duplex is then loaded into a multi-protein complex known as the RNA-induced silencing complex (RISC). A critical step within RISC is "passenger strand" dissociation; the antisense (guide) strand is retained, while the sense (passenger) strand is degraded [10]. The core catalytic component of RISC is an Argonaute protein [12] [14].
- Argonaute Architecture: Argonaute proteins contain two primary domains crucial for their function:
- Target Recognition and Cleavage: The guide siRNA, housed within Argonaute, base-pairs with a perfectly complementary sequence on the target messenger RNA (mRNA). Upon recognition, the catalytic PIWI domain of Argonaute cleaves the target mRNA, leading to its rapid degradation and thus, gene silencing [10] [14].
The diagram below illustrates this coordinated pathway from viral infection to gene silencing.
Key Enzymes and Cofactors in siRNA Biogenesis
Table 1: Core Protein Components of the siRNA Biogenesis Pathway
| Protein/Factor | Function | Key Domains/Features |
|---|---|---|
| Dicer | RNase III enzyme; initiates siRNA pathway by cleaving long dsRNA into siRNA duplexes. | RNase IIIa/IIIb domains, PAZ domain, dsRNA-binding domain. |
| Argonaute (Ago) | Catalytic core of RISC; binds siRNA guide strand and cleaves target mRNA. | PAZ domain (binds siRNA 3' end), MID domain (binds 5' phosphate), PIWI domain (slicer activity). |
| RISC Loading Complex | Multi-protein complex that facilitates the transfer of the siRNA duplex from Dicer to Argonaute. | Includes Dicer and other co-factors (e.g., TRBP in humans). |
Experimental Analysis of siRNA Biogenesis
Studying the siRNA pathway requires methodologies to trigger, observe, and quantify the silencing process. The following section outlines a standard experimental workflow and key reagents.
A Standard Workflow for siRNA-Mediated Gene Silencing
A typical experiment to harness siRNA for gene silencing involves a sequence of deliberate steps, from design to validation.
The Scientist's Toolkit: Essential Reagents for siRNA Research
Table 2: Key Research Reagents and Solutions for siRNA Experiments
| Reagent / Solution | Function / Purpose | Example Applications |
|---|---|---|
| Synthetic siRNA | Chemically synthesized 21-23 nt RNA duplex; provides immediate, transient silencing. | High-throughput screens; rapid validation of gene function; therapeutic development. |
| shRNA Expression Plasmids | DNA vectors that express short hairpin RNAs (shRNAs), which are processed into siRNAs by cellular machinery. | Creation of stable, long-term gene knockdown cell lines. |
| Dicer Enzymes | Recombinant Dicer protein for in vitro processing of long dsRNA into siRNA pools. | Generation of siRNA libraries; biochemical studies of dicing. |
| Lipid-Based Transfection Reagents | Cationic lipids/liposomes that complex with anionic siRNA to facilitate cellular uptake. | Standard in vitro delivery of siRNA into a wide range of cell types. |
| Argonaute-Specific Antibodies | Immunoreagents for detecting, quantifying, or immunoprecipitating Argonaute proteins and their bound RNAs. | Western blot, immunofluorescence, RIP-Chip, and CLIP-seq analyses. |
| RNAiMAX Lipofectamine | A proprietary, widely used lipid formulation optimized for high-efficiency siRNA delivery with low cytotoxicity. | Standardized transfection of adherent and suspension cells. |
| Dynamic PolyConjugates | Advanced polymer-siRNA conjugate delivery system designed for targeted in vivo delivery (e.g., to hepatocytes). | Systemic therapeutic delivery of siRNA. |
Advanced Methodologies and Protocols
Protocol: In Vitro Reconstitution of RISC Activity
This protocol allows for the biochemical dissection of RISC assembly and activity using purified components.
- siRNA Duplex Preparation: Resuspend synthetic siRNA strands in nuclease-free buffer. Anneal the guide and passenger strands by mixing equimolar amounts, heating to 90°C for 1 minute, and gradually cooling to room temperature.
- RISC Assembly: In a reaction buffer (e.g., 30 mM HEPES-KOH pH 7.4, 100 mM KOAc, 2 mM MgOAc), combine the following:
- 1 µM recombinant human Argonaute 2 (Ago2) protein.
- 100 nM siRNA duplex.
- 1 mM ATP.
- An ATP-regenerating system (e.g., creatine phosphate/creatine kinase).
- Incubate at 30°C for 60-90 minutes to allow for RISC loading and passenger strand ejection.
- Target Cleavage Assay: Add a radiolabeled or fluorescently-labeled target RNA substrate (complementary to the siRNA guide) to the assembled RISC. Incubate at 37°C for 30-60 minutes.
- Reaction Termination and Analysis: Stop the reaction by adding proteinase K and SDS. Resolve the cleavage products by denaturing polyacrylamide gel electrophoresis (PAGE). Visualize and quantify the intact substrate and cleavage products using autoradiography, phosphorimaging, or a fluorescence gel scanner.
Protocol: Validating Gene Silencing Efficacy
Following siRNA delivery, robust validation is required to confirm successful knockdown.
mRNA Level Quantification (qRT-PCR):
- RNA Extraction: Harvest cells 24-48 hours post-transfection. Isolate total RNA using a commercial kit (e.g., TRIzol or silica-membrane columns), including a DNase I digestion step to remove genomic DNA contamination.
- Reverse Transcription: Convert equal amounts of RNA (e.g., 1 µg) into cDNA using a high-capacity reverse transcription kit with random hexamers or oligo-dT primers.
- Quantitative PCR: Perform qPCR using gene-specific primers for the target gene and a stable reference gene (e.g., GAPDH, β-Actin). Calculate the percentage of silencing using the comparative ΔΔCt method relative to a non-targeting siRNA control.
Protein Level Analysis (Western Blot):
- Protein Extraction: Harvest cells 48-72 hours post-transfection. Lyse cells in RIPA buffer supplemented with protease inhibitors.
- Gel Electrophoresis and Transfer: Separate proteins by SDS-PAGE and transfer to a PVDF or nitrocellulose membrane.
- Immunodetection: Probe the membrane with a primary antibody against the target protein and a loading control antibody (e.g., β-Tubulin). Use HRP-conjugated secondary antibodies and chemiluminescent substrate for detection. Quantify band intensity using densitometry software.
The siRNA biogenesis pathway represents a fundamental biological process with profound implications for antiviral defense and functional genomics. The detailed mechanistic understanding of how dsRNA is processed into siRNAs by Dicer and loaded into the Argonaute-centered RISC complex to achieve sequence-specific silencing has provided researchers with an unparalleled tool for gene manipulation. Within VIGS research, this pathway is the engine that powers the technique, allowing for the functional annotation of genes in a rapid and specific manner. As delivery technologies, such as lipid nanoparticles and biomimetic systems, continue to advance, the application of siRNA-based strategies in both basic research and clinical therapeutics is poised for significant expansion. The standardized protocols and research tools outlined in this guide provide a foundation for scientists to rigorously exploit this pathway for innovative genetic research and therapeutic development.
The RNA-induced silencing complex (RISC) is a multiprotein complex that serves as the fundamental effector machinery in RNA interference (RNAi) pathways [15]. This ribonucleoprotein complex utilizes small interfering RNAs (siRNAs) as guide sequences to identify and cleave complementary messenger RNA (mRNA) targets, thereby executing sequence-specific gene silencing [15] [16]. In the context of virus-induced gene silencing (VIGS), RISC represents the crucial executive arm that translates the initial detection of viral pathogens into targeted antiviral defense [17] [5]. When plants encounter viral infection, double-stranded RNA replication intermediates trigger the RNAi machinery, leading to siRNA production that guides RISC to dismantle viral RNA genomes [17] [4]. This sophisticated mechanism not only provides plants with adaptive immunity against pathogens but has also been harnessed as a powerful reverse genetics tool for functional genomics studies [5] [4]. Understanding RISC's composition, loading mechanisms, and catalytic functions is therefore essential for advancing both fundamental plant biology and applied biotechnology.
Molecular Architecture of RISC
The RISC complex exhibits a dynamic composition that varies depending on organism, cell type, and the specific class of small RNA (siRNA or miRNA) it incorporates [15]. Despite this variability, certain core components remain conserved across systems, forming the catalytic and functional heart of the complex.
Core Protein Components
Argonaute Proteins (AGO): The catalytic center of RISC contains Argonaute family proteins, which possess endonuclease ("slicer") activity [15] [16]. Among human AGO isoforms, AGO2 uniquely mediates target mRNA cleavage through its RNase H-like catalytic domain containing a conserved DDH motif [16]. In Drosophila, genetic and biochemical studies have identified Ago-2 as essential for RNAi and a core component of RISC [18].
Dicer Enzymes: Although not a permanent RISC component, Dicer initiates RNAi by processing long double-stranded RNA into 21-23 nucleotide siRNA duplexes [15] [16]. Drosophila possesses two Dicers: Dcr-1 processes miRNA precursors, while Dcr-2 preferentially handles siRNA precursors [16].
Accessory Factors: Several additional proteins contribute to RISC function and regulation:
- dFXR: The Drosophila homolog of human Fragile X Mental Retardation Protein associates with RISC and may facilitate target recognition [18].
- VIG (Vasa intronic gene): An RNA-binding protein containing an RGG box domain that co-purifies with RISC [18].
- Tudor-SN: A component that binds hyper-edited double-stranded RNAs and may participate in precursor degradation [16].
Table 1: Core Components of the RNA-Induced Silencing Complex
| Component | Type | Key Function | Organisms Studied |
|---|---|---|---|
| Argonaute (AGO2) | Protein | Catalytic "slicer" enzyme; mRNA cleavage | Human, Drosophila, Plants |
| Dicer (Dcr-2) | Enzyme | Processes dsRNA into siRNAs | Drosophila, conserved in eukaryotes |
| dFXR/FMRP | RNA-binding protein | Potential target recognition; RISC association | Drosophila, Human homolog |
| VIG | RNA-binding protein | RISC component; exact role unclear | Drosophila, conserved homologs |
| Tudor-SN | Nuclease family | Degrades hyper-edited dsRNAs | Drosophila |
siRNA Loading and RISC Assembly
RISC assembly follows a carefully orchestrated sequence where siRNA duplexes are loaded and activated for target recognition:
Duplex Processing: Dicer cleaves long double-stranded RNA into 21-23 nt siRNA duplexes with 2-nt 3' overhangs [15].
Strand Selection: The siRNA duplex is loaded into RISC following the "asymmetry rule," where the strand with less stable 5' pairing is selected as the guide strand, while the passenger strand is degraded [15].
Complex Maturation: The guide strand integrates with AGO2 and accessory proteins to form the mature, active RISC complex [15] [16].
The Catalytic Mechanism of siRNA-Mediated Cleavage
RISC executes its silencing function through a precise catalytic mechanism that ensures specific targeting of complementary mRNA sequences. The process begins when the siRNA-loaded RISC scans cellular mRNAs, searching for sequences complementary to its guide strand [15]. The degree of complementarity between siRNA and target determines the mechanistic outcome, with perfect or near-perfect matches leading to endonucleolytic cleavage.
Molecular Requirements for Cleavage
Two critical conditions must be met for RISC-mediated cleavage to occur:
Complementarity Requirement: A near-perfect complementary match must exist between the guide strand and target mRNA sequence [15].
Catalytic Argonaute: The RISC must contain a catalytically active Argonaute protein (AGO2 in humans, Ago-2 in Drosophila) capable of cleaving the target mRNA [15] [16].
The crystal structure of Argonaute proteins reveals an RNase H-like fold that comprises the catalytic "slicer" activity [16]. The conserved DDH motif (D597, D669, H807 in human AGO2) coordinates magnesium ions that are essential for catalyzing the phosphodiester bond cleavage [16].
Cleavage Site Specificity
RISC-mediated cleavage occurs at a specific position relative to the guide siRNA: between nucleotides 10 and 11 from the 5' end of the guide strand [19]. This precise positioning ensures predictable fragmentation of the target mRNA.
Post-Cleavage mRNA Degradation
Following RISC-mediated cleavage, the mRNA fragments undergo rapid degradation through two major pathways:
5'-to-3' Degradation: XRN1 exonuclease degrades the fragment in cytoplasmic P-bodies [15].
3'-to-5' Degradation: The exosome and Ski complex process the fragment [15].
Both degradation pathways are initiated by removal of the mRNA's poly(A) tail and 5' cap, which destablizes the transcript and exposes it to exonuclease activity [15].
RISC in VIGS: Experimental Analysis
The critical role of RISC in viral RNA targeting during VIGS was definitively demonstrated through elegant experiments using the Cymbidium ringspot virus (CymRSV) system in Nicotiana benthamiana plants [17]. This experimental approach revealed key insights into how RISC executes antiviral defense at the molecular level.
Key Experimental Methodology
The foundational protocol for analyzing RISC-mediated viral RNA cleavage involves several critical steps:
Plant Material and Virus Infection:
- Use 6-leaf-stage Nicotiana benthamiana plants
- Infect with silencing suppressor mutant virus (Cym19stop) which lacks functional p19 protein
- Grow plants at constant 22°C until recovery phenotype appears (10-15 days post-inoculation) [17]
Sensor Construct Preparation:
- Clone 190-205 bp viral RNA fragments into binary vectors downstream of GFP open reading frame
- Generate both positive-sense and negative-sense orientation constructs
- Express sensor constructs in recovered leaves via Agrobacterium-mediated transient expression [17]
RNA and Protein Analysis:
- Extract total RNA from agro-infiltrated leaf tissues using Tri-reagent
- Detect viral siRNAs using ³²P-labeled riboprobes from specific CymRSV regions
- Analyze cleavage products via 3' RACE (rapid amplification of cDNA ends) sequencing [17]
Biochemical Fractionation:
- Isect high-molecular-mass nucleoprotein complexes from recovery leaves
- Cofractionate viral siRNA-containing complexes with miRNA-containing complexes
- Identify RISC components through protein analysis [17]
Critical Findings in VIGS Context
Application of this methodology revealed several fundamental principles of RISC function in VIGS:
- RISC mediates specific cleavage of viral RNA genomes rather than translational inhibition [17]
- Cleavage events do not occur randomly but form "hot spots" asymmetrically distributed along positive and negative viral RNA strands [17]
- Both polarities of viral RNA serve as RISC targets, though minus-sense RNA is cleaved more efficiently [17]
- No direct correlation exists between cleavage site hot spots and virus-generated siRNA abundance profiles [17]
Table 2: Experimental Approaches for Studying RISC in VIGS
| Method | Application | Key Outcome | Reference |
|---|---|---|---|
| Agroinfiltration of Sensor Constructs | Target cleavage analysis | Confirmed sequence-specific viral RNA cleavage | [17] |
| 3' RACE Sequencing | Cleavage site mapping | Identified hot spots for target RNA cleavage | [17] |
| Biochemical Fractionation | RISC complex isolation | Co-fractionation of viral siRNAs with effector complexes | [17] |
| siRNA Profiling | Small RNA analysis | Revealed asymmetric distribution of viral siRNAs | [17] |
The Scientist's Toolkit: Essential Reagents and Materials
Successful investigation of RISC function in VIGS research requires specific biological materials, molecular reagents, and methodological expertise. The following toolkit compiles essential resources based on proven experimental approaches.
Table 3: Essential Research Reagents for RISC-VIGS Studies
| Reagent/Resource | Function/Application | Specific Examples | Experimental Role |
|---|---|---|---|
| Plant Material | VIGS experimental system | Nicotiana benthamiana (6-leaf stage) | Model organism for virus infection studies [17] |
| Viral Vectors | VIGS induction & gene silencing | CymRSV, Cym19stop mutant, TRV, BBWV2 | Delivery of silencing triggers and target sequences [17] [4] |
| Sensor Constructs | Target cleavage analysis | GFP-Cym(+), GFP-Cym(-) binary plasmids | Report on RISC-mediated cleavage efficiency [17] |
| Agrobacterium Strains | Plant transformation | A. tumefaciens for transient expression | Delivery of sensor constructs into plant tissues [17] |
| siRNA Detection Tools | Small RNA analysis | ³²P-labeled riboprobes, LNA oligonucleotides | Detection and quantification of viral siRNAs [17] |
| Biochemical Fractionation | RISC complex isolation | Sucrose gradients, size exclusion chromatography | Isolation of native RISC complexes from plant tissues [17] |
| Antibodies | Protein component detection | Anti-AGO antibodies, anti-p19 sera | Identification of RISC-associated proteins [17] |
| 3' RACE Kit | Cleavage product mapping | Commercial 3' RACE systems | Identification of precise RISC cleavage sites [17] |
Therapeutic and Research Applications
The mechanistic understanding of RISC function has enabled groundbreaking applications in both therapeutic development and functional genomics research. In VIGS-based studies, RISC activity serves as the primary effector mechanism allowing researchers to transiently silence genes of interest through recombinant viral vectors [5] [4]. This approach has been successfully applied to characterize gene functions in numerous plant species, including pepper (Capsicum annuum L.), tomato, barley, and cotton [4].
Beyond plant functional genomics, RISC mechanics have inspired revolutionary therapeutic approaches. siRNA therapeutics harness the endogenous RISC machinery to silence disease-causing genes [19]. Several siRNA-based drugs have received regulatory approval, including:
- Patisiran: Treats hereditary transthyretin-mediated amyloidosis
- Givosiran: Targets acute hepatic porphyria
- Lumasiran: For primary hyperoxaluria type 1
- Inclisiran: Manages hypercholesterolemia [19]
These clinical applications underscore the importance of understanding RISC biochemistry, as therapeutic efficacy depends on efficient RISC loading and target cleavage [19]. Current research focuses on optimizing siRNA design through chemical modifications (2'-O-methyl, 2'-fluoro groups) and advanced delivery systems (lipid nanoparticles, GalNAc conjugates) to enhance RISC engagement and therapeutic outcomes [19].
The RISC complex represents nature's precise gene silencing machinery, executing sequence-specific RNA cleavage through its catalytic Argonaute core and guide siRNA. In VIGS, RISC serves as the ultimate effector that translates viral detection into targeted RNA destruction, providing plants with adaptive antiviral immunity [17] [5]. The detailed mechanistic understanding of RISC function - from siRNA loading and strand selection to target recognition and cleavage - has enabled transformative applications across basic research and clinical medicine [4] [19]. As our knowledge of RISC biology continues to expand, particularly in the context of epigenetic modifications and transgenerational inheritance [5], new opportunities emerge for developing increasingly sophisticated tools for genetic analysis, crop improvement, and human therapeutics. The ongoing characterization of RISC components and their regulatory mechanisms will undoubtedly yield further insights into eukaryotic gene regulation and enhance our ability to manipulate biological systems with precision.
Double-Stranded RNA as the Triggering Molecule for siRNA Production
Double-stranded RNA (dsRNA) serves as the fundamental trigger for one of the most important mechanisms in functional genomics and molecular biology: the production of small interfering RNAs (siRNAs) that guide sequence-specific gene silencing. This process forms the cornerstone of RNA interference (RNAi) pathways, which have been harnessed for revolutionary technologies such as Virus-Induced Gene Silencing (VIGS). Within the context of VIGS research, understanding the molecular journey from dsRNA to functional siRNA is paramount for designing effective gene silencing strategies. This technical guide provides a comprehensive examination of dsRNA-triggered siRNA biogenesis, detailing the underlying mechanisms, key experimental methodologies, and critical parameters for optimizing silencing efficiency. By synthesizing current knowledge with practical applications, this review serves as an essential resource for researchers leveraging siRNA technologies in functional genomics and therapeutic development.
RNA interference (RNAi) represents a conserved biological mechanism for sequence-specific gene silencing that plays crucial roles in antiviral defense, gene regulation, and genome maintenance in eukaryotic organisms [20] [13]. The discovery that double-stranded RNA (dsRNA) serves as the universal trigger for this pathway revolutionized molecular biology and opened new avenues for functional genomics research [21]. When introduced into cells, dsRNA initiates a sophisticated molecular cascade that ultimately leads to the degradation of complementary messenger RNA (mRNA) targets, effectively silencing gene expression.
The significance of dsRNA-triggered silencing extends far beyond its natural biological functions. In plant systems, this mechanism has been ingeniously adapted for Virus-Induced Gene Silencing (VIGS), a powerful technique that leverages recombinant viral vectors to deliver gene fragments and silence corresponding plant genes [4]. VIGS has emerged as a particularly valuable tool for functional genomics in species that are recalcitrant to stable genetic transformation, including many agriculturally important crops such as pepper (Capsicum annuum L.) and cotton (Gossypium hirsutum) [4] [22]. The effectiveness of VIGS and related RNAi technologies hinges entirely on the efficient conversion of dsRNA into small interfering RNAs (siRNAs), which serve as the guide molecules for target recognition.
This technical guide explores the molecular underpinnings of dsRNA-triggered siRNA production, with particular emphasis on its applications within VIGS research. We examine the key enzymes and pathways involved, detail experimental approaches for harnessing this mechanism, and provide practical guidance for optimizing silencing efficiency. By framing these concepts within the context of functional genomics, this review aims to equip researchers with the knowledge necessary to design effective gene silencing strategies for both basic research and applied biotechnology.
Molecular Mechanisms: From dsRNA to Functional siRNA
The transformation of dsRNA into functional siRNA involves a precisely orchestrated series of enzymatic reactions within the plant cell. This process can be divided into distinct stages: initiation, processing, and effector complex assembly, each mediated by specific protein complexes and enzymatic activities.
Initiation and dsRNA Processing
The siRNA biogenesis pathway begins with the recognition and cleavage of long dsRNA molecules by specialized RNase III-type enzymes known as Dicer-like (DCL) proteins [20] [13]. In the model plant Arabidopsis thaliana, four DCL enzymes (DCL1-4) perform specialized functions in processing different types of dsRNA precursors. For canonical antiviral RNAi and VIGS, DCL2 and DCL4 are primarily responsible for processing perfect duplex dsRNAs into 21- to 22-nucleotide siRNA duplexes [23]. These enzymes cleave dsRNA in an ATP-dependent manner, yielding short RNA duplexes with 2-nucleotide overhangs on each 3' end and 5' monophosphate groups [13].
The specific DCL enzyme activated depends on both the origin and structure of the dsRNA trigger. In VIGS, the dsRNA can originate from multiple sources: viral replication intermediates, hairpin RNA structures formed by viral genomes, or dsRNA synthesized by host RNA-dependent RNA polymerases (RDRs) using aberrant viral RNA as templates [4] [23]. The latter mechanism represents a crucial amplification step in the RNAi pathway, enhancing the potency and persistence of silencing through the production of secondary siRNAs [23].
siRNA Duplex Formation and RISC Loading
Following DCL-mediated processing, the resulting siRNA duplexes undergo strand selection and incorporation into the RNA-induced silencing complex (RISC). This complex is centered around Argonaute (AGO) proteins, which are the catalytic components responsible for target recognition and cleavage [20] [13]. Plants possess multiple AGO proteins with specialized functions; in Arabidopsis, ten AGO proteins (AGO1-10) have been identified, with AGO1 and AGO2 playing predominant roles in post-transcriptional gene silencing during antiviral defense [23].
The loading of siRNA into RISC involves the preferential selection of one strand of the siRNA duplex (the guide strand) based on the relative thermodynamic stability of its 5' end [13]. The complementary strand (the passenger strand) is typically degraded. The assembled RISC complex then uses the guide siRNA as a template to scan cellular mRNAs for complementary sequences. Upon finding a complementary target, the AGO protein catalyzes the endonucleolytic cleavage of the mRNA, preventing its translation and marking it for further degradation [20] [13].
Amplification and Systemic Spread
A distinctive feature of RNAi in plants is the capacity for amplification and systemic spread of the silencing signal. This amplification is mediated by host RDRs, which use cleaved mRNA fragments as templates to synthesize additional dsRNA molecules [23]. These secondary dsRNAs are subsequently processed into secondary siRNAs, exponentially expanding the pool of silencing molecules. The silencing signal can then move cell-to-cell through plasmodesmata and systemically through the phloem, enabling whole-plant gene silencing from a localized initiation site [4] [23]. This systemic property is particularly valuable in VIGS applications, where a single inoculation can lead to comprehensive gene silencing throughout the plant.
The following diagram illustrates the complete pathway from dsRNA to mRNA degradation:
Experimental Approaches for Triggering siRNA Production
Several well-established experimental methodologies leverage the dsRNA-to-siRNA pathway to induce gene silencing in plant systems. These approaches vary in their delivery mechanisms, persistence, and applicability to different plant species.
Virus-Induced Gene Silencing (VIGS)
VIGS represents one of the most widely used techniques for triggering siRNA production and gene silencing in plants. This approach utilizes recombinant viral vectors engineered to carry fragments of host target genes [4]. When introduced into plants, these vectors replicate and spread systemically, producing dsRNA intermediates during their replication cycle that are recognized by the host RNAi machinery. The resulting siRNAs then guide the silencing of both viral and homologous endogenous plant genes [4].
Multiple viral vectors have been developed for VIGS applications, with Tobacco Rattle Virus (TRV) emerging as one of the most popular due to its broad host range and efficient systemic movement [4]. The TRV system employs a bipartite design, with TRV1 encoding replication and movement proteins, and TRV2 carrying the coat protein and the insert sequence for the target gene. Other viral vectors used for VIGS include Potato Virus X (PVX), Broad Bean Wilt Virus 2 (BBWV2), Cucumber Mosaic Virus (CMV), and various geminiviruses [4] [24]. The selection of an appropriate viral vector depends on the host plant species, the target tissue, and the desired duration of silencing.
Exogenous Application of dsRNA
An alternative to viral vector-mediated delivery is the direct application of synthetic or biologically produced dsRNA. This approach, known as Spray-Induced Gene Silencing (SIGS), involves topically applying dsRNA solutions to plant surfaces [21] [25]. The exogenous dsRNA is taken up by plant cells and processed into siRNAs, triggering silencing of complementary genes. While the mechanisms of dsRNA uptake in plants are not fully understood, evidence suggests that it may involve pattern recognition receptors that recognize dsRNA as a pathogen-associated molecular pattern (PAMP) [21].
A significant challenge for SIGS is the environmental instability of naked dsRNA, which is rapidly degraded by nucleases and environmental factors [25]. To address this limitation, researchers have developed various carrier systems, particularly nanocarriers composed of biodegradable polymers or layered double hydroxide (LDH) clay nanosheets, that protect dsRNA during application and enhance cellular uptake [25].
Artificial miRNA and Syn-tasiRNA Platforms
Recent advances in RNAi technology have enabled the design of highly specific artificial small RNAs, including synthetic trans-acting siRNAs (syn-tasiRNAs) [24]. These platforms utilize minimal precursor scaffolds (as short as 54 nt) that are processed by the endogenous miRNA machinery to produce precise siRNA species [24]. The compact size of these precursors allows for their efficient incorporation into viral vectors, combining the specificity of designed siRNAs with the systemic delivery capabilities of VIGS.
The following experimental workflow illustrates the primary methods for triggering siRNA production in plants:
Key Parameters for Optimizing Silencing Efficiency
The effectiveness of dsRNA-triggered siRNA production and subsequent gene silencing depends on multiple factors that must be carefully considered in experimental design.
Target Sequence Selection
The selection of an appropriate target sequence within the gene of interest critically influences silencing efficiency. Research has demonstrated that different regions of the same mRNA can vary significantly in their susceptibility to RNAi, likely due to variations in secondary structure and accessibility [22]. Computational tools such as the Sfold program have been developed to predict optimal target sequences based on parameters including free energy of disruption (ΔGdisruption), duplex stability of the siRNA seed region (DSSE), and accessibility to the target site (AIS) [22].
Empirical studies in cotton have validated that sequences with lower ΔGdisruption values (indicating easier disruption of secondary structure) and moderate DSSE values (reflecting appropriate RISC assembly stability) consistently yield higher silencing efficiencies [22]. When designing VIGS constructs, target fragments of 200-500 nucleotides generally provide optimal results, balancing insert stability with silencing specificity [4].
Delivery Optimization
The method of dsRNA delivery significantly impacts the efficiency of siRNA production and systemic silencing. For VIGS, key parameters include the developmental stage of the plant, the concentration of the agroinoculum, and the specific infiltration method [4]. Younger plants with actively growing tissues typically show more efficient silencing, likely due to enhanced viral movement and RNAi activity in meristematic regions [4].
For exogenous dsRNA applications, delivery efficiency depends on overcoming multiple physical and biochemical barriers, including the hydrophobic cuticle, rigid cell wall, and extracellular nucleases [25]. The use of nanocarriers and surfactant compounds can significantly enhance dsRNA uptake by facilitating cuticle penetration and protecting dsRNA from degradation [25]. Recent advances in polymer-based nanocarriers have demonstrated particular promise for enhancing dsRNA stability and cellular delivery [25].
Environmental and Host Factors
Environmental conditions profoundly influence the efficiency of dsRNA-triggered silencing. Temperature, humidity, and photoperiod all affect viral replication and movement in VIGS applications, as well as the activity of the host RNAi machinery [4]. Most VIGS protocols recommend maintaining plants at 20-25°C with appropriate light conditions following inoculation to optimize silencing efficiency.
Host factors, including the plant genotype and the inherent activity of its RNAi pathway components, also significantly impact silencing outcomes [4]. Some plant species and varieties exhibit more robust or persistent silencing than others, reflecting natural variation in the expression or activity of DCL, AGO, and RDR proteins [4]. Additionally, viral suppressors of RNA silencing (VSRs) encoded by many plant viruses can be leveraged to enhance silencing in certain contexts by temporarily inhibiting components of the RNAi pathway [4].
Table 1: Key Parameters for Optimizing dsRNA-Triggered Silencing Efficiency
| Parameter Category | Specific Factors | Optimal Conditions/Considerations |
|---|---|---|
| Target Sequence | Fragment length | 200-500 bp for VIGS inserts |
| GC content | Moderate (30-60%) | |
| Secondary structure | Low ΔGdisruption values preferred | |
| Sequence specificity | Avoid off-target matches to non-target genes | |
| Delivery Method | Plant developmental stage | Younger seedlings (2-4 leaf stage) |
| Agroinoculum concentration | OD₆₀₀ of 0.5-2.0 for most species | |
| Infiltration method | Syringe infiltration or vacuum infiltration | |
| Nanocarrier formulation | Cationic polymers for dsRNA complexation | |
| Environmental Conditions | Temperature | 20-25°C post-inoculation |
| Light intensity | Moderate to high (avoid low light) | |
| Photoperiod | Species-dependent (often 16h light/8h dark) | |
| Humidity | 60-70% relative humidity |
Quantitative Assessment of Silencing Efficiency
Rigorous quantification of silencing efficiency is essential for evaluating and optimizing dsRNA-triggered siRNA production. Multiple methodological approaches provide complementary data on different aspects of the silencing process.
Molecular confirmation of successful siRNA production typically involves small RNA northern blotting or high-throughput sequencing to detect and quantify siRNAs derived from the dsRNA trigger [24]. These methods provide direct evidence that the introduced dsRNA is being processed correctly by the host RNAi machinery. For VIGS applications, the presence of virus-derived siRNAs (vsiRNAs) can be confirmed using virus-specific probes or primers [23].
The functional consequences of siRNA production are most commonly assessed by measuring transcript levels of the target gene using reverse transcription quantitative PCR (RT-qPCR) [24]. Effective silencing typically results in 70-90% reduction in target mRNA levels compared to control treatments. For visible phenotypes, such as photobleaching in phytoene desaturase (PDS) silencing, morphological assessment provides additional confirmation of silencing efficiency [4].
High-throughput approaches for screening siRNA combinations have recently been developed, utilizing encoded microcarriers that enable simultaneous testing of hundreds to thousands of siRNA combinations in a single assay [26]. These advanced platforms employ composite micro-carriers responsive to near-infrared light and magnetic fields to achieve photoporation-facilitated siRNA transfection with single-cell resolution [26]. While initially developed for mammalian systems, similar approaches show promise for plant research applications.
Table 2: Methods for Assessing Silencing Efficiency
| Assessment Method | Parameter Measured | Technical Considerations |
|---|---|---|
| Small RNA Northern Blot | siRNA accumulation | Requires specific probes; moderate sensitivity |
| sRNA Sequencing | Comprehensive siRNA profile | High cost; bioinformatics expertise needed |
| RT-qPCR | Target mRNA reduction | Most common method; requires proper controls |
| Western Blot | Target protein reduction | Antibody availability; protein half-life considerations |
| Phenotypic Scoring | Visible phenotype (e.g., photobleaching) | Quick assessment; subjective quantification |
| Histochemical Staining | Tissue-specific silencing patterns | Spatial information; may require specialized reporters |
| High-throughput Screening | Multiple siRNA combinations | Advanced platforms; specialized equipment needed |
Research Reagent Solutions
Successful implementation of dsRNA-triggered siRNA production requires specific reagents and biological materials optimized for RNAi applications. The following table summarizes key resources for establishing these methodologies.
Table 3: Essential Research Reagents for dsRNA-Triggered Silencing Studies
| Reagent Category | Specific Examples | Function and Application |
|---|---|---|
| Viral Vectors | TRV (Tobacco Rattle Virus) | Broad host range VIGS vector |
| PVX (Potato Virus X) | Alternative VIGS vector | |
| BBWV2 (Broad Bean Wilt Virus 2) | VIGS vector for challenging species | |
| Cloning Systems | Gateway-compatible vectors | Simplified cloning of target fragments |
| Golden Gate assembly systems | Modular vector construction | |
| Agrobacterium Strains | GV3101 | Standard strain for agroinfiltration |
| LBA4404 | Alternative strain for specific hosts | |
| dsRNA Production Systems | T7/T3 RNA polymerase | In vitro transcription of dsRNA |
| HT115(DE3) E. coli | Bacterial expression of dsRNA | |
| Nanocarrier Materials | Cationic polymers (e.g., PDDA) | dsRNA complexation and protection |
| Layered double hydroxide (LDH) | Clay nanosheets for dsRNA delivery | |
| Carbon dots | Enhanced cellular uptake | |
| Detection Reagents | siRNA detection probes | Northern blot or in situ hybridization |
| Antibodies for AGO proteins | Immunoprecipitation of RISC complexes |
Double-stranded RNA serves as the essential triggering molecule for siRNA production in plant RNAi pathways, forming the mechanistic foundation for powerful functional genomics tools such as Virus-Induced Gene Silencing. The molecular journey from dsRNA to functional siRNA involves a sophisticated interplay of DCL-mediated processing, RISC assembly, and systemic amplification that can be harnessed for precise gene silencing. As research advances, emerging technologies including nanocarrier-enhanced delivery, synthetic tasiRNA platforms, and high-throughput screening methods are expanding the applications and efficiency of dsRNA-triggered silencing. By understanding and optimizing the parameters that influence siRNA production and function, researchers can continue to leverage this fundamental biological pathway to address diverse questions in plant biology, crop improvement, and sustainable agriculture.
In the field of functional genomics, precise gene silencing is a cornerstone for elucidating gene function. Two key classes of small non-coding RNAs, small interfering RNA (siRNA) and microRNA (miRNA), play central roles in RNA interference (RNAi) pathways. Although both are approximately 20-25 nucleotides in length and operate through the RNA-induced silencing complex (RISC), they originate from different precursors and exhibit distinct mechanisms of target recognition and regulation [27] [28]. Understanding their differences is critical for selecting the appropriate tool for gene silencing experiments, especially in advanced applications like Virus-Induced Gene Silencing (VIGS), a powerful reverse genetics technique used to study gene function in plants [4] [5].
Fundamental Differences in Origin and Biogenesis
The pathways through which siRNAs and miRNAs are synthesized represent a primary point of differentiation, influencing their subsequent roles and specificities.
Table 1: Biogenesis Pathways of siRNA and miRNA
| Feature | siRNA (Small Interfering RNA) | miRNA (MicroRNA) |
|---|---|---|
| Origin | Exogenous or endogenous long double-stranded RNA (dsRNA) [28]. | Endogenous genome-encoded transcripts [28]. |
| Precursor | Long, perfectly base-paired double-stranded RNA [28]. | Primary miRNA (pri-miRNA) with hairpin structures, transcribed by RNA polymerase II [28] [29]. |
| Nuclear Processing | Not applicable; typically cytoplasmic. | Pri-miRNA is processed by Drosha/DGCR8 into precursor miRNA (pre-miRNA) [28]. |
| Export to Cytoplasm | Not applicable. | Pre-miRNA is exported by Exportin-5 [28]. |
| Cytoplasmic Processing | Dicer cleaves long dsRNA into double-stranded siRNAs [28]. | Dicer cleaves the pre-miRNA loop, forming a miRNA duplex [28]. |
| RISC Loading | The siRNA duplex is loaded into RISC; the guide strand is selected to target complementary mRNA for cleavage [28]. | The miRNA duplex is loaded into RISC; the guide strand is selected to target mRNA for translational repression or degradation [28]. |
Diagram 1: Biogenesis and Mechanism of siRNA and miRNA
Mechanisms of Action and Target Specificity
The functional distinction between siRNAs and miRNAs lies in their mode of interaction with messenger RNA (mRNA) targets, which dictates the outcome of silencing.
siRNA: High Specificity for mRNA Cleavage
siRNAs mediate RNA interference (RNAi) by guiding RISC to perfectly complementary mRNA sequences. This perfect match allows the catalytic component of RISC, typically an Argonaute protein, to cleave the target mRNA, leading to its rapid degradation [27] [28]. This mechanism provides high specificity, making siRNAs ideal for knocking down the expression of a single, specific gene. Consequently, siRNAs are powerful tools for functional gene studies and therapeutic development aimed at silencing particular pathogenic genes [27] [30].
miRNA: Broad Regulation through Translational Repression
In contrast, miRNAs typically bind to the 3' untranslated region (3' UTR) of target mRNAs with partial complementarity, particularly through a short "seed region" [27]. This imperfect binding leads to translational repression without significant mRNA degradation, although it can also lead to mRNA destabilization in some cases [28]. A single miRNA can regulate hundreds of different mRNAs, and a single mRNA may be targeted by multiple miRNAs, allowing miRNAs to function as master regulators of complex genetic networks involved in development, differentiation, and stress responses [27] [31].
Overlap and Experimental Observations
Notably, the distinction can sometimes blur under experimental conditions. A seminal 2003 study demonstrated that an exogenously introduced siRNA could repress a reporter gene with partially complementary binding sites in its 3' UTR, functioning much like an miRNA by causing translational repression without mRNA cleavage [32]. This highlights a level of mechanistic overlap dependent on the nature of the siRNA-mRNA interaction.
Table 2: Functional Comparison of siRNA and miRNA
| Aspect | siRNA | miRNA |
|---|---|---|
| Mechanism of Action | mRNA cleavage and degradation [27] [28]. | Translational repression and/or mRNA destabilization [27] [28]. |
| Complementarity | Perfect or near-perfect match to the target [28]. | Partial complementarity, especially in the seed region [27]. |
| Specificity | Highly specific, usually targeting a single gene [27]. | Broad, regulating multiple genes and pathways [27] [31]. |
| Primary Function | Defense against viruses and transposons; experimental gene knockdown [28] [17]. | Endogenous regulation of gene expression during development and stress [29] [31]. |
| Outcome | Potent knockdown of a specific protein by destroying its mRNA template. | Fine-tuning of protein output from a network of genes. |
siRNA as the Engine of Virus-Induced Gene Silencing (VIGS)
VIGS is a powerful technique in plant functional genomics that leverages the siRNA pathway. It uses recombinant viral vectors to deliver a fragment of a host plant gene, triggering the plant's innate antiviral RNAi defense system [4] [5].
The Molecular Mechanism of VIGS
When a modified virus containing a plant gene sequence infects the host, the viral RNA replicates, generating double-stranded RNA intermediates. The plant's Dicer-like enzymes recognize and process these dsRNAs into virus-derived siRNAs (vsiRNAs) of 21–24 nucleotides [4] [17]. These vsiRNAs are then incorporated into RISC, which they program to target and cleave not only the viral RNA but also the complementary endogenous mRNA transcripts from which the inserted fragment was derived [17] [5]. This results in the specific "silencing" or knockdown of the target plant gene, allowing researchers to observe the resulting phenotype and infer gene function.
VIGS Workflow and Key Vectors
The standard VIGS protocol involves cloning a 200-500 bp fragment of the plant gene of interest into a viral vector, such as the widely used Tobacco Rattle Virus (TRV) system [4]. The recombinant vector is then introduced into plant tissues, often via Agrobacterium tumefaciens-mediated transformation (agroinfiltration). The virus spreads systemically, and the silencing phenotype can be observed within weeks.
Diagram 2: Experimental Workflow of VIGS
Key viral vectors used in VIGS include:
- Tobacco Rattle Virus (TRV): One of the most versatile and widely used systems, especially in Solanaceae family plants like pepper and tomato, due to its efficient systemic movement and mild symptoms [4].
- Tobacco Mosaic Virus (TMV): The first virus used to develop a VIGS vector [4] [5].
- DNA Viruses (e.g., Geminiviruses): Vectors based on viruses like Cotton leaf crumple virus (CLCrV) are also employed [4].
The Scientist's Toolkit: Key Reagents and Applications
Selecting the right molecules and tools is fundamental for successful gene regulation studies. The choice between siRNA and miRNA tools depends heavily on the research objective.
Table 3: Research Reagent Solutions for RNAi Studies
| Tool / Reagent | Function / Description | Application Example |
|---|---|---|
| Pre-designed Synthetic siRNAs | Chemically synthesized siRNA duplexes designed for high specificity and potency against a single target mRNA [27]. | Studying the function of a specific gene in ribosome biogenesis or mechanisms of cancer therapy resistance [27]. |
| VIGS Viral Vectors (e.g., TRV1/TRV2) | Bipartite viral vector system used in plants to deliver target gene fragments and trigger siRNA-mediated silencing of endogenous genes [4]. | High-throughput forward genetics screens in plants like Capsicum annuum to identify genes governing fruit quality, disease resistance, or stress tolerance [4]. |
| miRNA Mimics | Synthetic small RNAs that mimic endogenous mature miRNAs, supplementing their levels to observe downstream protein down-regulation [27]. | Gain-of-function studies to investigate the role of a specific miRNA in processes like breast cancer metastasis or cellular aging [27]. |
| miRNA Inhibitors | Chemically modified antisense oligonucleotides that sequester and block the function of endogenous miRNAs [27]. | Loss-of-function studies to observe the resulting protein up-regulation and elucidate the endogenous role of a specific miRNA [27]. |
| Artificial miRNAs (amiRNAs) | Engineered miRNA precursors where the native miRNA sequence is replaced with a user-defined sequence to silence specific targets [29] [31]. | Used in plant metabolic engineering to precisely silence key enzymes in biosynthetic pathways, such as those for flavonoids [29]. |
siRNAs and miRNAs are mighty molecules in the RNAi arsenal, each with distinct origins, mechanisms, and applications. siRNAs are highly specific agents of sequence-specific mRNA degradation, making them ideal for targeted gene knockdown in both basic research (e.g., VIGS) and therapeutic development. miRNAs, in contrast, are endogenous master regulators that fine-tune complex gene networks through translational repression. The context of VIGS research powerfully illustrates the application of the siRNA pathway as a robust functional genomics tool. Understanding their differences and potential overlaps is essential for designing rigorous experiments, interpreting results accurately, and selecting the optimal strategy for gene silencing and functional analysis.
Virus-induced gene silencing (VIGS) represents a conserved RNA interference (RNAi) mechanism that provides adaptive antiviral defense across diverse eukaryotic organisms. This whitepaper examines VIGS from an evolutionary perspective, detailing the molecular machinery of small interfering RNA (siRNA) biogenesis and activity within the context of host-pathogen co-evolution. We present quantitative frameworks for predicting VIGS efficiency, experimental protocols for robust silencing, and visualization of key pathways. The content specifically addresses the role of siRNA in directing sequence-specific viral RNA degradation, highlighting its applications in functional genomics and therapeutic development for researchers and drug development professionals.
Virus-induced gene silencing (VIGS) is an evolutionarily conserved RNAi mechanism that provides adaptive antiviral defense in plants, invertebrates, and other eukaryotes. This defense system leverages small interfering RNAs (siRNAs) to mediate sequence-specific degradation of viral RNA, representing a fundamental genetic immune response against pathogenic invasion [33]. The evolutionary significance of VIGS stems from its dual role as both a natural antiviral mechanism and a powerful biotechnology tool. In natural ecosystems, VIGS operates under constant evolutionary pressure from rapidly mutating viruses, leading to an arms race between host defense strategies and viral evasion tactics [33]. The specificity and adaptability of the siRNA pathway make it particularly effective against diverse viral pathogens, allowing hosts to target essential viral genes with precision while minimizing collateral damage to cellular processes. This evolutionary perspective provides crucial insights for developing siRNA-based therapeutics that can overcome viral resistance mechanisms.
Molecular Mechanisms of siRNA Biogenesis and Activity
Core Pathway of Antiviral RNAi
The antiviral RNAi pathway is initiated when viral double-stranded RNA (dsRNA) replicates intermediates are recognized and processed by host Dicer or Dicer-like (DCL) enzymes into 21-24 nucleotide primary siRNAs [34] [33]. These primary siRNAs are loaded into Argonaute (AGO) proteins within the RNA-induced silencing complex (RISC), where they guide identification and cleavage of complementary viral RNA sequences [35] [33]. In an amplification phase, RNA-dependent RNA polymerases (RdRPs) use the cleaved viral RNA fragments as templates to synthesize secondary dsRNAs, which are subsequently processed into secondary siRNAs (22G RNAs in C. elegans) [34]. This amplification mechanism creates a robust, systemic antiviral response capable of controlling viral replication throughout infected tissues.
Key Protein Components and Their Functions
Table 1: Core Protein Components of the Antiviral RNAi Pathway
| Protein Component | Function in Antiviral Defense | Organismic Context |
|---|---|---|
| Dicer/DCL | Processes viral dsRNA into primary siRNAs | Conserved across eukaryotes [34] [33] |
| AGO/RDE-1 | Slicer activity in RISC for viral RNA cleavage | Central effector in C. elegans and plants [34] [35] |
| RdRP/RRF-1 | Amplifies silencing by generating secondary siRNA precursors | Critical in C. elegans and plants [34] |
| RDE-4/DRH-1 | dsRNA binding facilitates Dicer activity | Important for primary vsiRNA biogenesis [34] |
| RDE-3 | Adds poly(UG) tails to cleaved viral RNAs | Recruits RdRPs in C. elegans [34] |
| SGS3 | Stabilizes cleaved RNA fragments for RdRP synthesis | Plant VIGS systems [35] |
siRNA-Mediated Viral RNA Degradation
The guide strand of mature siRNAs directs RISC to complementary viral RNA targets through Watson-Crick base pairing. Once bound, the AGO protein catalyzes endonucleolytic cleavage of the target RNA between nucleotides 10 and 11 of the siRNA guide strand [33]. This cleavage event disrupts viral gene expression and replication, effectively limiting infection spread. The resulting RNA fragments are either degraded by cellular nucleases or used as templates for secondary siRNA production, further amplifying the antiviral response.
The following diagram illustrates the core siRNA biogenesis and antiviral activity pathway:
Quantitative Analysis of VIGS Efficiency Parameters
Computational Prediction of Silencing Efficacy
The Sfold program provides a robust computational framework for predicting VIGS efficiency by analyzing three key parameters that influence target accessibility and siRNA activity [22]. These parameters enable researchers to select optimal target sequences before experimental validation:
Table 2: Key Parameters for Predicting VIGS Efficiency with Sfold
| Parameter | Description | Impact on VIGS Efficiency |
|---|---|---|
| ΔGdisruption | Free energy cost for local alteration of target structure to allow siRNA binding | Lower ΔGdisruption values increase probability of target binding [22] |
| DSSE | Stability of 5'--antisense end of 4 base segments | Lower DSSE values enhance RISC assembly and guide strand selection [22] |
| AIS | Asymmetric instability score of siRNA duplex | Higher AIS values facilitate unwinding and strand loading into RISC [22] |
Experimental Validation of Computational Predictions
In Gossypium hirsutum (cotton), application of Sfold analysis to the phytoene desaturase (PDS) gene demonstrated that integrating these three parameters successfully predicted VIGS efficiency across different target sequences [22]. Target fragments with optimal ΔGdisruption, DSSE, and AIS values produced 56.7% silencing efficiency in experimental validation, significantly higher than suboptimal fragments [22]. This computational-experimental pipeline allows researchers to pre-screen target sequences and select those most likely to produce strong silencing phenotypes, saving considerable time and resources in functional genomics studies.
Advanced Experimental Protocols for VIGS
Injection of No-Apical-Bud Stem Sections (INABS) Protocol
The INABS method represents a significant advancement in VIGS delivery, offering higher efficiency and faster results than traditional agroinfiltration techniques [36]. This protocol optimizes plant tissue selection and inoculation sites for maximum transformation success:
Plant Material Preparation: Select no-apical-bud stem sections with "Y-type" asymmetric structure containing an axillary bud approximately 1-3 cm in length from tomato plants [36].
Agrobacterium Preparation: Transform Agrobacterium tumefaciens with appropriate TRV vectors (pTRV1 and pTRV2 containing target gene fragments). Grow cultures to OD₆₀₀ = 1.0, which provides optimal transformation efficiency [36].
Infiltration Process: Slowly inject 100-200 μl of agroinfiltration liquid into the bare stem of no-apical-bud stem sections using a plastic syringe and needle. Continue injection until a film of agroinfiltration liquid forms at the top of the injected stem sections, indicating complete filling [36].
Post-Inoculation Care: Maintain injected plants under controlled conditions (25°C with 16/8-h day/night cycle). Silencing phenotypes typically appear within 6-8 days post-inoculation (dpi), with full development by 10-12 dpi [36].
Efficiency Assessment: Monitor target gene expression reduction via qRT-PCR and observe visual phenotypes (e.g., bleaching for PDS silencing). The INABS method achieves 56.7% VIGS efficiency and 68.3% virus inoculation success at 8 dpi with OD₆₀₀ = 1.0 [36].
The following workflow diagram illustrates the optimized INABS protocol:
Syn-tasiR-VIGS: A Transgene-Free Approach
Recent advances have enabled VIGS implementation without transgenic components through synthetic trans-acting siRNAs (syn-tasiRNAs) expressed from viral vectors:
Minimal Precursor Design: Engineer minimal non-TAS precursors containing a 22-nt endogenous microRNA target site, 11-nt spacer, and 21-nt syn-tasiRNA sequence(s) [35].
Viral Vector Construction: Clone minimal precursors into RNA virus vectors (e.g., Tobacco rattle virus) that can be delivered by spraying infectious crude extracts onto leaves [35].
Plant Immunization: Apply viral vectors containing syn-tasiRNAs targeting essential pathogen genes to vaccinate plants against viruses, achieving complete immunization in some applications [35].
This transgene-free approach addresses regulatory concerns while maintaining high silencing efficacy through proper siRNA design and delivery.
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Research Reagent Solutions for VIGS Experiments
| Reagent/Resource | Function/Application | Specific Examples |
|---|---|---|
| VIGS Vectors | Viral vectors for inducing gene silencing | Tobacco rattle virus (TRV), Tomato yellow leaf curl virus (TYLCV) [36] [22] |
| Agrobacterium Strains | Delivery system for VIGS constructs | A. tumefaciens GV3101 [35] [36] |
| Computational Tools | Predicting target sequences and silencing efficiency | Sfold program (ΔGdisruption, DSSE, AIS parameters) [22] |
| Plant Materials | Model systems for VIGS optimization | Nicotiana benthamiana, Gossypium hirsutum, Arabidopsis thaliana [35] [36] [22] |
| Efficiency Reporters | Visual markers for successful silencing | Phytoene desaturase (PDS) [36] [22] |
| Analysis Tools | Validating silencing efficiency | qRT-PCR, Northern blotting, H₂O₂ detection assays [36] |
Evolutionary Arms Race: Viral Escape and Host Adaptation
The continuous evolutionary dynamics between host VIGS mechanisms and viral pathogens has driven sophisticated adaptation strategies on both sides. Understanding these co-evolutionary processes is essential for developing durable siRNA-based therapeutics:
Viral Escape Mechanisms
RNA viruses encode error-prone RNA-dependent RNA polymerases that lack proofreading capabilities, resulting in mutation rates up to 10⁷-fold higher than DNA viruses [33]. This genetic plasticity enables rapid evolution of siRNA resistance through:
- Point mutations in siRNA target sequences that create mismatches and reduce silencing efficiency [33]
- Nucleotide deletions or substitutions that alter local RNA secondary structure, occluding target sites from RISC binding [33]
- Homologous recombination that rapidly generates novel viral variants capable of evading RNAi [33]
Experimental evidence demonstrates that escape mutants can emerge within just 3-10 passages under siRNA selection pressure, typically through single or double nucleotide substitutions [33].
Host Counter-Adaptation Strategies
Host organisms have evolved diverse countermeasures to maintain effective antiviral RNAi despite viral escape attempts:
Multiplexed siRNA targeting: Simultaneous production of multiple siRNAs targeting different viral regions, as demonstrated in syn-tasiRNA technology, reduces the probability of complete escape through single mutations [35]
Conserved target selection: Focusing siRNA responses on essential, evolutionarily constrained viral genes limits the fitness of escape mutants [33]
Genetic polymorphism: Natural variation in RNAi pathway components (e.g., argonaute genes) within host populations creates diverse selective landscapes that impede viral adaptation [33]
VIGS represents an evolutionarily optimized antiviral defense system that leverages the specificity and amplifiability of siRNA-mediated silencing to control viral infections. The quantitative parameters governing VIGS efficiency (ΔGdisruption, DSSE, and AIS) provide researchers with predictive frameworks for experimental design, while advanced protocols like INABS and syn-tasiR-VIGS enable efficient, transgene-free implementation. The ongoing evolutionary arms race between host silencing mechanisms and viral escape variants underscores the need for sophisticated siRNA design strategies that anticipate and circumvent resistance development.
Future directions in VIGS research will likely focus on optimizing multiplexed siRNA cocktails against conserved viral targets, developing delivery systems that overcome tissue-specific barriers, and leveraging evolutionary principles to design durable RNAi-based therapeutics. As our understanding of host-virus co-evolution deepens, siRNA technology will continue to provide powerful tools for both basic research and clinical applications against viral pathogens.
VIGS Implementation: Vector Systems and Research Applications
Virus-induced gene silencing (VIGS) has emerged as an indispensable reverse genetics tool for functional genomics in plants, particularly for species resistant to stable genetic transformation. The technology leverages the plant's innate antiviral defense mechanism—post-transcriptional gene silencing (PTGS)—to knock down expression of target genes. Central to this process are small interfering RNAs (siRNAs), which serve as the sequence-specific guides that direct mRNA degradation. When a recombinant virus carrying a fragment of a plant gene infects the host, the plant's defense machinery processes the viral RNA into 21-24 nucleotide siRNAs. These siRNAs are incorporated into the RNA-induced silencing complex (RISC), which then targets and cleaves complementary endogenous mRNA transcripts, resulting in gene silencing [4] [5] [37]. The efficiency and specificity of VIGS are therefore intrinsically linked to the production and systemic movement of these siRNAs, making viral vector selection a critical determinant of experimental success.
Major VIGS Vector Systems and Their Characteristics
The choice of viral vector is paramount, as it influences the host range, silencing efficiency, duration, and capacity for foreign gene inserts. Vectors are derived from modified plant viruses whose genomes are engineered to carry fragments of host genes.
Tobacco Rattle Virus (TRV)
Tobacco Rattle Virus (TRV) is one of the most versatile and widely deployed VIGS vectors, especially within the Solanaceae family. Its bipartite, positive-sense single-stranded RNA genome requires the use of two plasmid components: TRV1 and TRV2. TRV1 encodes proteins for replication and movement (134K and 194K replicases, a 29K movement protein), while TRV2 carries the coat protein gene and serves as the insertion site for the target plant gene fragment [4] [38]. The broad host range of TRV extends beyond model plants to include horticultural crops like pepper and tomato, and even woody species such as walnut and poplar [4] [39] [38]. A key advantage is its ability to infect meristematic tissues and cause relatively mild viral symptoms, which minimizes the interference with phenotypic analysis [4] [38]. Silencing phenotypes typically manifest systemically within 2-3 weeks post-inoculation and can be maintained for several months under optimized conditions [38].
Barley Stripe Mosaic Virus (BSMV)
For monocotyledonous plants, Barley Stripe Mosaic Virus (BSMV) is a cornerstone VIGS vector. BSMV has a tripartite, single-stranded RNA genome (designated RNAα, RNAβ, and RNAγ), with the target gene fragment typically cloned into the RNAγ component [38]. Its application is predominantly reported in cereals like barley (Hordeum vulgare) and wheat (Triticum aestivum), where it has been instrumental in characterizing genes involved in abiotic stress tolerance and disease resistance [38]. A notable limitation of BSMV is the tendency to induce pronounced viral symptoms, such as leaf mosaic and chlorosis, which can complicate the interpretation of silencing phenotypes. Furthermore, its host range is generally more restricted compared to TRV [38].
Cucumber Mosaic Virus (CMV)
Cucumber Mosaic Virus (CMV) is another significant RNA virus with a tripartite, single-stranded RNA genome. It has been successfully modified into a VIGS vector for both monocot and dicot species. A notable application is in orchids, where a CMV-based system was used to silence the phytoene desaturase (PDS) gene, with a remarkably short 33-bp insertion fragment proving sufficient to induce silencing that persisted into the second year in new leaves [40]. This demonstrates CMV's potential for long-term silencing studies. However, similar to BSMV, CMV infection can cause severe viral symptoms in some host plants, which is a significant drawback for phenotypic observation [4].
Table 1: Comparative Overview of Major VIGS Vector Systems
| Vector | Genome Type | Primary Host Range | Key Advantages | Major Limitations |
|---|---|---|---|---|
| Tobacco Rattle Virus (TRV) | Bipartite ssRNA | Broad (Dicots, some Woody plants) | Efficient systemic spread, mild symptoms, meristem invasion [4] [38] | Requires two plasmid components |
| Barley Stripe Mosaic Virus (BSMV) | Tripartite ssRNA | Monocots (Barley, Wheat) | Effective in cereal crops; well-established protocol [38] | Strong viral symptoms; restricted host range [38] |
| Cucumber Mosaic Virus (CMV) | Tripartite ssRNA | Broad (Monocots and Dicots) | Can achieve very long-lasting silencing [40] | Often causes severe viral symptoms [4] |
| Geminiviruses (e.g., CLCrV) | Circular ssDNA | Dicots (Cotton, Nicotiana benthamiana) | High silencing efficiency in specific hosts [4] | Limited insert capacity; potential for integration concerns |
Molecular Workflow of VIGS and siRNA Biogenesis
The following diagram illustrates the core molecular mechanism of VIGS, highlighting the central role of siRNA biogenesis and function.
Figure 1: The Core Mechanism of VIGS and siRNA Biogenesis. The process begins with the introduction of recombinant viral RNA, leading to the generation of double-stranded RNA (dsRNA) by RNA-directed RNA polymerase (RDRP). Dicer-like (DCL) enzymes process dsRNA into siRNAs, which are loaded into RISC to guide target mRNA cleavage.
Optimized Experimental Protocols for VIGS
A successful VIGS experiment depends on meticulous optimization of several parameters. The following protocol synthesizes best practices from recent studies in various plant systems.
Vector Construction and Target Fragment Selection
- Target Gene Cloning: A 300-500 base pair fragment of the target gene is typically amplified from cDNA and cloned into a multiple cloning site of the VIGS vector (e.g., TRV2) using restriction enzymes (e.g., EcoRI and BamHI) or Gateway recombination [41]. For specialized applications like virus-induced transcriptional gene silencing (ViTGS), the insert should target the promoter region of the gene to induce DNA methylation [5] [37].
- Fragment Selection and Design: To ensure high silencing efficiency and minimize off-target effects, the fragment should be highly specific to the target gene. Bioinformatics tools like the SGN-VIGS tool (https://vigs.solgenomics.net/) can be used to select optimal fragments and check for specificity [41]. Using a shorter fragment (e.g., as short as 84 bp in cornflower) can sometimes enhance viral mobility and silencing efficiency [40].
- Control Constructs: Always include both positive and negative controls. The phytoene desaturase (PDS) gene is a standard positive control, as its silencing produces a visible photobleaching phenotype. An empty vector (e.g., TRV2:0) serves as a negative control for virus-induced symptoms [41] [39].
Agrobacterium Preparation and Inoculation
- Strain and Culture: The recombinant vector plasmids (e.g., TRV1 and TRV2-derived) are transformed into an Agrobacterium tumefaciens strain such as GV3101. A single colony is inoculated into YEP liquid medium supplemented with appropriate antibiotics (e.g., 50 mg/L kanamycin and rifampicin) and cultured at 28°C with shaking (200 rpm) until the mid-logarithmic growth phase (OD600 = 0.6–0.8) [41].
- Induction and Resuspension: Bacterial cells are pelleted and resuspended in an infiltration buffer (commonly containing 10 mM MES, 200 µM acetosyringone, 10 mM MgCl2, and 0.03% Silwet-77). The OD600 is adjusted to an optimal concentration, which varies by species—for example, OD600 = 0.8 for Atriplex canescens [41], OD600 = 1.0 for walnut [39], and OD600 = 1.2–1.8 for cornflower capitula [40]. The TRV1 and TRV2 suspensions are mixed in a 1:1 ratio and incubated in darkness for 3 hours at room temperature to induce virulence gene expression [41].
- Plant Inoculation Methods: The choice of inoculation method is critical and depends on the plant species and its developmental stage.
- Vacuum Infiltration: Germinated seeds or young seedlings are submerged in the Agrobacterium suspension and subjected to a vacuum (e.g., 0.5 kPa for 10 minutes for Atriplex canescens) [41]. This is highly effective for achieving systemic silencing.
- Syringe Infiltration: Using a needleless syringe, the suspension is infiltrated directly into the abaxial air spaces of leaves. This is common for Nicotiana benthamiana and tomato.
- Spraying and Other Methods: For larger plants or specific tissues like flowers, spraying the suspension or direct injection into buds (e.g., in cornflower) [40] or fruits can be effective.
Table 2: Key Research Reagents and Solutions for VIGS
| Reagent/Solution | Composition / Key Feature | Function in VIGS Protocol |
|---|---|---|
| pTRV1 & pTRV2 Vectors | Binary plasmids under CaMV 35S promoter | Deliver viral RNA components; TRV2 carries the target gene insert [4] |
| Agrobacterium Strain GV3101 | Disarmed helper strain | Mediates delivery of T-DNA containing viral vectors into plant cells [41] [39] |
| Infiltration Buffer | 10 mM MES, 200 µM Acetosyringone, 10 mM MgCl2, 0.03% Silwet-77 | Suspension medium that induces Agrobacterium virulence and facilitates plant cell infiltration [41] |
| Phytoene Desaturase (PDS) | Carotenoid biosynthesis gene | Visual reporter gene; silencing causes photobleaching, validating system efficiency [41] [39] |
| YEP Medium | Yeast Extract, Peptone, NaCl (with antibiotics) | Culture medium for growing transformed Agrobacterium [41] |
Post-Inoculation Procedures and Efficiency Validation
- Plant Growth Conditions: After inoculation, plants are maintained under controlled environmental conditions. Temperature is a critical factor, as it significantly impacts viral replication and spread. Most protocols use temperatures between 22-25°C [41]. Moderate humidity and a 16-hour light/8-hour dark photoperiod are standard.
- Phenotypic Monitoring and Validation: Systemic silencing phenotypes typically appear in newly emerged leaves 10-28 days post-inoculation (dpi), depending on the species and target gene [41] [42]. For the positive control PDS, photobleaching is a clear indicator.
- Molecular Confirmation: Silencing efficiency must be quantified using quantitative reverse transcription PCR (qRT-PCR) to measure the reduction in target gene transcript levels compared to control plants. A successful experiment typically shows a 40-80% reduction in mRNA abundance [41]. The presence of the virus and the generation of target-specific siRNAs can be confirmed via Northern blot analysis.
Advanced Applications: From VIGS to VIGE and Epigenetics
The principles of VIGS are now being extended into more advanced genetic technologies, opening new frontiers in plant research and breeding.
Virus-Induced Genome Editing (VIGE)
Virus-Induced Genome Editing (VIGE) represents a powerful convergence of VIGS and CRISPR/Cas9 technology. In VIGE, viral vectors are used to transiently deliver CRISPR/Cas components into plant cells, enabling targeted genome editing without the need for stable transformation [43] [42]. This approach has the potential to generate transgene-free, edited plants in a single generation, bypassing the regulatory and technical hurdles associated with traditional genetically modified organisms (GMOs). Current research focuses on overcoming challenges such as the large size of the Cas gene, which can exceed the packaging capacity of many viral vectors, and achieving efficient editing in meristematic cells to ensure heritability [43].
Virus-Induced Epigenetic Modifications
Beyond transcript degradation, VIGS can induce heritable epigenetic modifications through a process known as RNA-directed DNA methylation (RdDM). When the viral vector carries a sequence homologous to a gene's promoter (rather than the coding sequence), the resulting siRNAs can guide DNA methyltransferases to methylate the promoter region. This leads to transcriptional gene silencing (TGS), which can be stable and transgenerationally inherited [5] [37]. For instance, silencing the FWA promoter in Arabidopsis via TRV-based vectors led to DNA methylation and a heritable late-flowering phenotype [5] [37]. This application, known as virus-induced transcriptional gene silencing (ViTGS), provides a robust tool for studying epigenetic regulation and has significant potential for crop improvement.
The selection of an appropriate viral vector—be it TRV, BSMV, CMV, or another—is a foundational decision that dictates the success of a VIGS experiment. This choice must be guided by the host plant species, the target tissue, and the specific scientific question. The core mechanism, driven by the elegant specificity of siRNAs, provides a relatively fast and flexible alternative to stable transformation. As the field advances, the integration of VIGS with cutting-edge technologies like CRISPR/Cas9 (VIGE) and epigenetics (ViTGS) is pushing the boundaries of functional genomics. These developments promise to accelerate the pace of gene discovery and validation, ultimately contributing to the development of improved crop varieties with enhanced traits.
Virus-Induced Gene Silencing (VIGS) has emerged as a powerful functional genomics tool that leverages the plant's endogenous RNA interference (RNAi) machinery to suppress target gene expression. The foundation of VIGS lies in the biological mechanism of post-transcriptional gene silencing (PTGS), an antiviral defense system in plants [4]. Within this framework, small interfering RNAs (siRNAs) serve as the central effector molecules that guide sequence-specific mRNA degradation. When a recombinant viral vector containing a fragment of a plant gene infects the plant, the replication process generates double-stranded RNA (dsRNA), which is recognized and processed by Dicer-like (DCL) enzymes into 21-24 nucleotide siRNAs [4]. These siRNAs are then incorporated into the RNA-induced silencing complex (RISC), directing the cleavage of complementary endogenous mRNA transcripts, thereby leading to reduced gene expression and observable phenotypic changes that enable functional characterization [4].
The efficiency and specificity of VIGS are fundamentally determined by the careful design of the construct inserted into the viral vector. Key parameters—insert size, orientation, and specificity—directly influence the quality and diversity of siRNA populations generated, ultimately determining the effectiveness and reliability of gene silencing. This technical guide examines these core construct design principles within the context of modern VIGS research, providing evidence-based methodologies for optimizing silencing constructs, with particular emphasis on Solanaceous species like Capsicum annuum L. where stable transformation remains challenging and VIGS often represents the most viable tool for high-throughput functional screening [4].
Core Principles of VIGS Construct Design
Insert Size Optimization
Insert size significantly impacts processing efficiency by cellular DCL enzymes and subsequent systemic silencing. Both excessively short and long inserts present limitations for effective VIGS.
Table 1: Optimal Insert Size Parameters for VIGS Constructs
| Insert Size Range | Processing Efficiency | siRNA Diversity | Risks and Limitations |
|---|---|---|---|
| 200-500 bp | High | Moderate | Ideal for generating sufficient siRNAs; minimizes off-target effects |
| 500-800 bp | High | High | Optimal range; maximizes siRNA diversity and silencing robustness |
| >800 bp | Variable | High | Increased chance of recombination in viral vector; potential for non-specific silencing |
| <100 bp | Low | Low | May generate insufficient siRNAs for effective silencing |
Recent advancements have introduced minimal precursor designs for synthetic trans-acting small interfering RNAs (syn-tasiRNAs), where a precursor of only 54 nucleotides proved as efficient as canonical ~1 kb TAS precursors [24]. This minimal precursor consists of a 22-nt miRNA target site, an 11-nt spacer, and the 21-nt syn-tasiRNA, demonstrating that precisely engineered small inserts can mediate highly efficient silencing [24].
Insert Orientation and Placement
The orientation of the insert within the viral vector profoundly affects the formation of dsRNA replication intermediates, which are essential for triggering the siRNA pathway. Inserts should be cloned in an antisense or sense orientation relative to the viral promoter to ensure production of translatable dsRNA molecules during viral replication. The transcriptional start and termination signals must be considered to ensure proper expression of the insert fragment. For Tobacco Rattle Virus (TRV)-based vectors—among the most widely used VIGS systems—inserts are typically cloned between the coat protein subgenomic promoter and the terminator sequence in the TRV2 vector [4].
Strategic placement should avoid regions with high secondary structure that might interfere with viral replication or movement. Additionally, the insert should be positioned to maximize exposure to DCL enzymes, typically achieved by flanking it with viral sequences that facilitate proper processing.
Ensuring Specificity and Minimizing Off-Target Effects
Specificity remains a critical challenge in RNAi-based approaches. Several strategies can enhance target specificity:
- Computational Screening: Before construct design, potential insert sequences should be computationally screened against the entire transcriptome using tools like BLASTN to identify regions with minimal homology to non-target genes [24].
- Unique Sequence Selection: Target regions with least similarity to other genes, typically focusing on 3' UTRs that exhibit higher sequence divergence.
- Syn-tasiRNA Design: Synthetic trans-acting siRNAs can be computationally designed to achieve highly specific silencing with reduced off-target effects [24]. These syn-tasiRNAs are generated from modified TAS transcripts where endogenous tasiRNA sequences are substituted with designed syn-tasiRNAs, allowing simultaneous silencing of multiple genes with high specificity [24].
- Avoiding Functional Domains: Regions encoding highly conserved protein domains (e.g., kinase domains, DNA-binding motifs) should generally be avoided as they increase likelihood of cross-silencing related genes.
Table 2: Specificity Enhancement Strategies for VIGS Constructs
| Strategy | Methodology | Effect on Specificity |
|---|---|---|
| Sequence Alignment | BLAST against entire transcriptome | Identifies and eliminates cross-homology regions |
| UTR Targeting | Prefer 3' UTR sequences | Higher sequence divergence reduces off-targets |
| Syn-tasiRNA Design | Computational design of artificial sRNAs | Maximizes on-target, minimizes off-target effects |
| GC Content Optimization | Maintain 30-60% GC content | Prevents extreme structures affecting processing |
| Multi-target Approach | Different constructs for same gene | Confirms phenotype is target-specific |
Experimental Protocols for VIGS Construct Design and Validation
In Silico Design of Syn-tasiRNAs for VIGS
This protocol adapts the syn-tasiRNA design approach for VIGS applications [24]:
- Target Sequence Selection: Identify the target mRNA sequence (NM_001234567.1) and extract 21-nt candidate sites every 21-nt along the transcript.
- Off-Target Analysis: For each candidate sequence, perform genome-wide alignment using tools like Bowtie or BLASTN to identify potential off-targets. Discard any sequence with ≤3 mismatches to non-target transcripts.
- Efficiency Prediction: Rank remaining candidates using scoring algorithms that consider positional nucleotide preferences (A at position 1, U at position 10) for AGO loading.
- Precursor Design: For minimal syn-tasiRNA precursors, incorporate the 22-nt miRNA target site (e.g., NbmiR482a for Nicotiana benthamiana), an 11-nt spacer, and the selected 21-nt syn-tasiRNA sequence [24].
- Vector Compatibility: Add appropriate restriction sites (e.g., BsaI for Golden Gate assembly) to the 5' and 3' ends for cloning into your selected VIGS vector.
Cloning into VIGS Vectors
Materials:
- pTRV1 and pTRV2 vectors (or other viral vectors)
- Syn-tasiRNA insert designed above
- Restriction enzymes and buffers
- T4 DNA ligase or Gibson Assembly Master Mix
- Competent E. coli cells
Method:
- Digest pTRV2 vector and syn-tasiRNA insert with appropriate restriction enzymes (e.g., BsaI for Golden Gate assembly).
- Purify digested fragments using Zymo-Spin I columns or similar.
- Set up ligation reaction with T4 DNA ligase or Gibson Assembly Master Mix.
- Transform into competent E. coli cells and select on appropriate antibiotics.
- Verify constructs by colony PCR and Sanger sequencing.
Plant Inoculation and Silencing Validation
Agroinfiltration Protocol [4]:
- Transform validated pTRV1 and pTRV2-syn-tasiRNA constructs into Agrobacterium tumefaciens strain GV3101.
- Grow overnight cultures in LB medium with appropriate antibiotics.
- Resuspend bacterial pellets in infiltration medium (10 mM MES, 10 mM MgCl₂, 200 μM acetosyringone) to OD₆₀₀ = 0.3-1.0.
- Mix pTRV1 and pTRV2-syn-tasiRNA cultures in 1:1 ratio.
- Infiltrate into expanded leaves of 2-3 week old plants using needleless syringe.
- Maintain plants at 20-22°C with high humidity for 3-5 days post-infiltration, then lower to normal growth conditions.
Silencing Efficiency Validation:
- Sampling: Collect systemic leaves 2-4 weeks post-infiltration.
- RNA Extraction: Use TRIzol method to extract total RNA.
- RT-qPCR: Perform reverse transcription followed by quantitative PCR with target-specific primers to measure mRNA reduction.
- Phenotypic Assessment: Document visual phenotypes and correlate with molecular data.
Visualizing Key Molecular Pathways and Workflows
VIGS Molecular Pathway
Construct Design Workflow
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Research Reagents for VIGS Experiments
| Reagent/Vector | Function/Application | Key Features |
|---|---|---|
| pTRV1 & pTRV2 Vectors | Bipartite TRV-based VIGS system | Broad host range, efficient systemic movement, targets meristematic tissues [4] |
| pMDC32B-AtmiR173aTS-B/c | Syn-tasiRNA precursor with Arabidopsis miRNA target | Contains AtmiR137a target site for Arabidopsis applications [24] |
| pMDC32B-NbmiR482aTS-B/c | Syn-tasiRNA precursor with N. benthamiana miRNA target | Contains NbmiR482a target site for Solanaceae applications [24] |
| pLBPVXBa-M | Potato virus X-based vector | For transgene-free virus-based expression of syn-tasiRNAs [24] |
| Agrobacterium tumefaciens GV3101 | Plant transformation | Standard strain for agroinfiltration delivery of VIGS constructs [24] |
| T4 DNA Ligase | DNA fragment joining | Essential for conventional cloning of inserts into VIGS vectors [24] |
| GeneArt Gibson Assembly | DNA assembly without restriction sites | Enables seamless cloning of syn-tasiRNA precursors [24] |
The strategic design of VIGS constructs—with careful attention to insert size, orientation, and specificity—forms the foundation of successful gene silencing studies in plants. The emergence of syn-tasiRNA technology and minimal precursor designs represents a significant advancement in achieving highly specific and efficient silencing. By integrating these construct design principles with appropriate viral vectors and delivery methods, researchers can leverage VIGS as a powerful tool for functional genomics across a broad range of plant species, including those recalcitrant to stable transformation. As these methodologies continue to evolve, they will further accelerate gene discovery and functional characterization in plant biology and agricultural biotechnology.
The functional analysis of genes through Virus-Induced Gene Silencing (VIGS) relies critically on efficient delivery systems to introduce silencing triggers into plant cells. The core mechanism of VIGS utilizes the plant's own RNA interference machinery, where virus-derived small interfering RNAs (siRNAs) guide the sequence-specific degradation of complementary endogenous mRNA transcripts [4] [5]. The efficacy of this process is fundamentally dependent on the methodology used to deliver these viral vectors or RNA molecules into plant tissues. This guide examines both established and emerging delivery technologies, with a focus on their application in modern plant research and development. Agrobacterium-mediated transformation remains the most widely adopted method, but recent advances in nanoparticle-based delivery systems offer promising alternatives that could overcome species-specific limitations.
Core Mechanism: siRNA Biogenesis and Action in VIGS
The following diagram illustrates the fundamental molecular pathway of siRNA-mediated gene silencing in plants, which forms the biological basis for VIGS.
Figure 1: The core mechanism of Virus-Induced Gene Silencing (VIGS) relies on post-transcriptional gene silencing (PTGS). The process initiates when viral double-stranded RNA (dsRNA) is recognized and cleaved by host Dicer-like (DCL) enzymes into 21-24 nucleotide small interfering RNAs (siRNAs). These siRNAs are incorporated into the RNA-induced silencing complex (RISC), where an Argonaute (AGO) protein uses the siRNA as a guide to identify and cleave complementary target mRNA. This sequence-specific degradation leads to gene silencing. The process can be amplified by host RNA-dependent RNA polymerases (RDRPs), which use the cleaved RNA as a template to generate secondary siRNAs, enabling systemic spread of the silencing signal throughout the plant [4] [5].
Established Delivery Method: Agrobacterium-Mediated Transformation
Agrobacterium tumefaciens is a soil-borne bacterium naturally capable of transferring DNA into plant genomes. This property has been harnessed to deliver VIGS vectors into plant cells. The standard methodology involves using disarmed (non-pathogenic) Agrobacterium strains engineered to carry binary plasmid vectors. These vectors contain the viral genome components, including the gene of interest targeted for silencing [44] [4]. The general workflow for Agrobacterium-mediated VIGS is illustrated below.
Figure 2: General workflow for Agrobacterium-mediated VIGS. The process begins with the preparation of Agrobacterium strains harboring the engineered VIGS vectors. The bacterial culture is then prepared to optimal density and induced with acetosyringone to enhance virulence. Plants are inoculated using one of several methods, followed by a period of co-cultivation to allow for T-DNA transfer. After incubation under controlled environmental conditions, silencing efficiency is assessed through molecular and phenotypic analysis [44] [8].
Key Agrobacterium Strains and VIGS Vectors
Table 1: Common Agrobacterium Strains and Viral Vectors Used in VIGS Research
| Component | Examples | Key Characteristics & Applications | References |
|---|---|---|---|
| Agrobacterium Strains | GV3101, EHA105, C58C1, LBA4404 | Disarmed strains with varying host range and transformation efficiency; choice is often empirical and species-dependent. | [44] [45] [8] |
| RNA Virus Vectors | Tobacco Rattle Virus (TRV), Tobacco Mosaic Virus (TMV) | TRV: Broad host range (Solanaceae, Arabidopsis), efficient systemic movement, mild symptoms. TMV: One of the first VIGS vectors developed. | [44] [4] [8] |
| DNA Virus Vectors | Geminiviruses (e.g., CLCrV, ACMV) | Useful in some dicotyledonous plants; based on single-stranded DNA viruses. | [4] |
| Model Plant Reporters | Phytoene Desaturase (PDS), Magnesium Chelatase (ChlH) | Silencing causes visible photobleaching phenotypes, allowing for easy visual assessment of VIGS efficiency. | [44] [8] |
Optimized Protocol: Agrobacterium-VIGS for Sunflower
The following detailed protocol, optimized for sunflower, demonstrates a specific application of this methodology and can be adapted for other species [8].
- Vector Construction and Agrobacterium Transformation: Clone a ~200 bp fragment of the target gene (e.g., HaPDS) into the multiple cloning site of a TRV2 vector (e.g., pYL156). Transform the recombinant plasmid and the necessary helper plasmid (TRV1, e.g., pYL192) into an Agrobacterium strain (e.g., GV3101) via electroporation.
- Culture Preparation: Streak transformed Agrobacterium from a glycerol stock onto LB agar plates with appropriate antibiotics (e.g., kanamycin, gentamicin, rifampicin). Incubate at 28°C for 36-48 hours. Pick a single colony to inoculate a liquid LB culture with the same antibiotics and grow overnight at 28°C with shaking. Centrifuge the culture and resuspend the pellet in an induction medium (10 mM MgCl₂, 10 mM MES, 200 µM acetosyringone) to a final OD₆₀₀ of 1.0. Incubate this suspension for 3-4 hours at room temperature with shaking.
- Plant Inoculation (Seed-Vacuum Method):
- Prepare sunflower seeds by removing the seed coat.
- Mix the induced TRV1 and TRV2 Agrobacterium cultures in a 1:1 ratio.
- Submerge the seeds in the bacterial mixture and apply a vacuum (approximately 0.8 bar) for 5 minutes.
- Release the vacuum abruptly to ensure infiltration.
- Co-cultivate the seeds on moist filter paper in the dark for 6 hours.
- Plant Growth and Analysis: Sow the treated seeds directly in soil and grow under controlled greenhouse conditions (e.g., 22°C, 18h light/6h dark photoperiod). Monitor plants for the development of silencing phenotypes (e.g., photobleaching for PDS). Validate silencing efficiency 3-4 weeks post-inoculation using RT-qPCR to measure the downregulation of the endogenous target transcript.
Novel Approach: Nanoparticle-Mediated siRNA Delivery
A cutting-edge alternative to biological vectors is the use of synthetic nanoparticles for direct siRNA delivery. This approach bypasses the need for viral replication and the host range limitations of Agrobacterium. A recent breakthrough involves Guanidinium-containing disulfide assembled siRNA (Gu+-siRNA) nanoparticles [46].
These nanoparticles are synthesized by electrostatically complexing negatively charged siRNA with a positively charged guanidinium-functionalized disulfide molecule (GDM). At an optimal N/P ratio ([Gu+]/[PO₄⁻]) of 15:1, this self-assembly forms monodisperse, spherical nanoparticles approximately 200 nm in diameter. A key feature is the disulfide bond, which is stable extracellularly but cleaves in the reductive environment of the cell interior, facilitating siRNA release [46].
Experimental Protocol: Gu+-siRNA NPs in Arabidopsis
- Nanoparticle Preparation: Mix GDM adjuvant material with siRNA (e.g., targeting STM, WER) in nuclease-free water at an N/P ratio of 15:1. Vortex and incubate at room temperature for 30 minutes to allow for self-assembly into Gu+-siRNA NPs. Characterize the resulting nanoparticles for size and stability using Dynamic Light Scattering (DLS) and Transmission Electron Microscopy (TEM).
- Plant Treatment (Root Immersion): Grow Arabidopsis thaliana or rice seedlings on 1/2 MS medium for 7-10 days. Immerse only the root tips of the seedlings in a solution of Gu+-siRNA NPs (concentration ~1.5 mM), ensuring the aerial parts are not in direct contact. Maintain the plants in this configuration for 1-2 hours.
- Analysis of Systemic Delivery: After the incubation period, gently rinse the roots and analyze the plants. Use confocal microscopy to detect fluorescently labeled FITC-siRNA in root tips, root growth zones, and distal leaves to confirm long-distance transport. Harvest leaf tissue from various nodes and perform RT-qPCR to quantify the knockdown of the target genes. Assess the resulting phenotypic consequences, such as altered development or enhanced salt tolerance [46].
Comparative Analysis of Delivery Methodologies
Table 2: Quantitative Comparison of Agrobacterium vs. Nanoparticle Delivery Methods
| Parameter | Agrobacterium-Mediated VIGS | Nanoparticle-Mediated Delivery |
|---|---|---|
| Underlying Mechanism | Biological T-DNA transfer and viral replication. | Chemical/Physical complexation and cellular uptake. |
| Key Reagents | Agrobacterium strains, TRV vectors, antibiotics, acetosyringone. | Guanidinium (Gu+) molecules, synthetic siRNA, nuclease-free water. |
| Transformation Efficiency | Varies by species and protocol: e.g., ~65% in C. tenuiflora (Co-cultivation), 71-91% in sunflower (seed-vacuum) [44] [8]. | High efficiency demonstrated in protoplasts and whole plants; species-independent [46]. |
| Silencing Onset & Duration | Systemic; phenotypes observed from 3-4 weeks post-infiltration; can be sustained [44] [8]. | Rapid; systemic silencing detected in distal leaves within days; sustained effect shown [46]. |
| Host Range | Can be limited by plant genotype and susceptibility to the viral vector [4] [8]. | Broad and species-independent; effective in both dicots (Arabidopsis) and monocots (rice) [46]. |
| Key Advantages | Established protocol, high efficiency in amenable species, systemic spread via virus. | Bypasses genotype dependency, no viral genome or foreign DNA integration, high biocompatibility. |
| Major Limitations | Genotype-dependent efficiency, potential for viral symptomology, regulatory concerns for GMOs. | Technology is nascent, optimization required for different tissues/targets, cost of synthetic siRNA. |
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Reagent Solutions for VIGS Delivery Research
| Reagent / Material | Function in Experimental Workflow | Specific Examples & Notes |
|---|---|---|
| Agrobacterium tumefaciens | Biological vector for delivering T-DNA containing the VIGS construct. | GV3101: Commonly used for Solanaceae and Arabidopsis. EHA105: Used in kiwifruit and other crops. Strain choice impacts efficiency [45] [8]. |
| TRV-Based Vectors | Plasmid system carrying the viral genome and the insert to be silenced. | pYL192 (TRV1) & pYL156 (TRV2): A widely adopted bipartite system. The gene fragment is cloned into TRV2 [8]. |
| Acetosyringone | A phenolic compound that induces the Agrobacterium vir genes, enhancing T-DNA transfer. | Used in culture induction and infiltration media; typical concentration of 150-200 µM [47] [8]. |
| Guanidinium (Gu+) Molecules | Cationic adjuvant that self-assembles with siRNA to form stable, protective nanoparticles. | Enables endocytosis-independent cellular uptake and protects siRNA from RNase degradation [46]. |
| Selection Antibiotics | Maintain selective pressure for plasmids in bacterial and plant cultures. | Kanamycin: Selects for TRV vectors. Rifampicin & Gentamicin: Select for specific Agrobacterium strains [45] [8]. |
| Reporter Genes | Visual markers for rapid assessment of transformation or silencing efficiency. | GUS (β-glucuronidase): Histochemical staining. GFP (Green Fluorescent Protein): Non-destructive, in vivo monitoring [44] [47] [45]. |
High-throughput functional genomics is indispensable for translating genomic sequences into biological understanding, with Virus-Induced Gene Silencing (VIGS) emerging as a powerful reverse genetics tool. VIGS enables rapid, transient gene knockdown by harnessing the plant's innate RNA interference (RNAi) machinery, facilitating large-scale gene function screening without stable transformation [4] [5]. This process is primarily mediated by small interfering RNAs (siRNAs), which guide sequence-specific silencing of target genes [4] [48]. The integration of VIGS with high-throughput methodologies is dramatically accelerating the pace of gene discovery in plant biology and beyond.
VIGS and the Central Role of siRNA
The Molecular Mechanism of VIGS
VIGS operates through the plant's post-transcriptional gene silencing (PTGS) pathway, an antiviral defense mechanism. The process initiates when a recombinant viral vector, carrying a fragment of a host gene, is introduced into the plant via agroinfiltration or other methods [4]. Inside the plant cell, the viral genome replicates, generating double-stranded RNA (dsRNA) intermediates [5]. These dsRNA molecules are recognized and cleaved by the host enzyme Dicer-like (DCL) into 21-24 nucleotide small interfering RNAs (siRNAs) [4] [5]. These siRNAs are then loaded into the RNA-induced silencing complex (RISC), where they serve as guides for the sequence-specific identification and cleavage of complementary messenger RNA (mRNA) transcripts, leading to gene silencing [4] [48] [5]. A key feature of this process is the amplification of silencing signals by host RNA-directed RNA polymerases (RDRPs), which generate secondary siRNAs, enabling systemic spread and persistence of the silencing effect throughout the plant [5].
siRNA as the Effector Molecule
The siRNA is the central effector in the VIGS pathway, determining the specificity and efficiency of gene silencing. The 21-24 nucleotide siRNAs generated from viral dsRNA direct the RISC complex to degrade homologous endogenous mRNAs [5]. Furthermore, these siRNAs can move systemically through the plant's vascular system, enabling silencing in tissues distant from the initial infection site [46]. In some cases, siRNA can also lead to epigenetic modifications through RNA-directed DNA methylation (RdDM), resulting in stable, heritable gene silencing [5].
Diagram: siRNA-mediated silencing pathways in VIGS. PTGS occurs in the cytoplasm, leading to mRNA degradation, while nuclear siRNA can induce TGS via DNA methylation [4] [5].
Advanced VIGS Technologies and Optimization
Viral Vector Systems for High-Throughput Applications
The choice of viral vector is critical for successful high-throughput VIGS. Different vectors offer distinct advantages depending on the host plant and experimental goals.
Table 1: Key Viral Vectors Used in High-Throughput VIGS
| Vector Name | Virus Type | Host Range | Key Features | Primary Applications |
|---|---|---|---|---|
| Tobacco Rattle Virus (TRV) | RNA virus | Broad (Solanaceae, Arabidopsis) | Mild symptoms, efficient systemic movement, targets meristematic tissues [4] | Functional genomics in model plants and crops [4] [48] |
| Bean Bean Wilt Virus 2 (BBWV2) | RNA virus | Broad | — | Used in Solanaceae functional studies [4] |
| Cucumber Mosaic Virus (CMV) | RNA virus | Very broad | — | Wide host range applications [4] |
| Geminiviruses (CLCrV, ACMV) | DNA virus | Variable | — | — |
| Cotton Leaf Crumple Virus (CLCrV) | DNA virus | Dicots | — | — |
Enhancing VIGS Efficiency through Suppressor Protein Engineering
A significant advancement in VIGS technology is the engineering of viral suppressors of RNA silencing (VSRs) to enhance silencing efficiency. Viruses naturally encode VSRs to counteract host RNAi defenses. Recent research has successfully decoupled the dual functions of the Cucumber Mosaic Virus 2b (C2b) protein. A truncated variant, C2bN43, was engineered to retain its ability to suppress systemic silencing (promoting viral spread) while losing its local suppression activity, thereby potentiating stronger gene silencing in systemically infected tissues [48]. This innovation significantly improves VIGS efficacy in challenging systems like pepper plants, particularly in reproductive organs such as anthers [48].
Nanoparticle-Mediated siRNA Delivery
For species resistant to traditional viral vectors, nanoparticle-based siRNA delivery represents a breakthrough. A recently developed guanidinium-containing disulfide assembled siRNA (Gu+-siRNA) nanoparticle system demonstrates remarkable species independence and efficient systemic movement [46]. These nanoparticles protect siRNA from degradation, facilitate cellular uptake via an endocytosis-independent pathway, and utilize the plant's vascular system for long-distance transport, enabling effective gene silencing in both monocot and dicot plants, including rice and Arabidopsis [46]. This technology bypasses the limitations of biolistics and VIGS, offering a highly flexible and efficient alternative for high-throughput functional genomics.
Experimental Protocols for High-Throughput VIGS
Standard TRV-Based VIGS Protocol
The following protocol is adapted for high-throughput applications using the widely adopted Tobacco Rattle Virus (TRV) system [4].
Week 1: Vector Preparation and Agrobacterium Transformation
- Clone Target Fragment: Insert a 300-500 bp PCR fragment of the target gene (e.g., CaPDS or CaAN2) into the multiple cloning site of the TRV2 vector [4] [48].
- Transform Agrobacterium: Introduce the constructed TRV1 and recombinant TRV2 plasmids into an appropriate Agrobacterium tumefaciens strain (e.g., GV3101) via electroporation or freeze-thaw transformation [4].
- Select Positive Colonies: Plate transformed agrobacteria on selective media containing appropriate antibiotics (e.g., kanamycin, rifampicin) and incubate at 28°C for 2 days [4].
Week 2: Agroinoculum Preparation and Plant Infiltration
- Starter Cultures: Inoculate single colonies of TRV1 and TRV2 agrobacteria into liquid LB media with antibiotics and incubate at 28°C with shaking for ~24 hours [4].
- Induction Cultures: Subculture the bacteria into induction media (e.g., LB with acetosyringone and MES buffer) and grow to an optical density (OD600) of 0.4-2.0 [4] [48].
- Agroinoculum Mix: Combine the TRV1 and TRV2 cultures in a 1:1 ratio, resuspend in infiltration buffer (10 mM MgCl₂, 10 mM MES, 200 μM acetosyringone), and incubate at room temperature for 3-6 hours [4].
- Plant Infiltration: Inoculate plants at the 2-4 leaf stage using a needleless syringe for leaf infiltration or vacuum infiltration for whole seedlings [4]. For pepper, silencing is often performed at the cotyledon stage [48].
Week 3-6: Phenotypic Analysis and Validation
- Monitor Phenotype: Observe plants for the development of silencing phenotypes (e.g., photo-bleaching for PDS silencing) 2-4 weeks post-infiltration [4] [48].
- Molecular Validation: Quantify silencing efficiency using quantitative RT-PCR (qRT-PCR) to measure target gene transcript levels. Use the 2−ΔΔCt method for analysis, with a housekeeping gene (e.g., GAPDH) for normalization [48].
- Documentation: Photograph clear phenotypic changes, such as altered pigmentation in anthers or leaves [48].
Protocol for Enhanced VIGS Using Engineered Suppressors
To implement the optimized TRV-C2bN43 system [48]:
- Clone the C2bN43 truncated suppressor fragment downstream of a subgenomic RNA promoter (e.g., from Pea Early Browning Virus) into the pTRV2-lic vector.
- Follow the standard protocol above, using the engineered pTRV2-C2bN43 vector instead of the standard TRV2.
- This system is particularly recommended for silencing genes in reproductive tissues or in plant species where conventional VIGS efficiency is low.
Critical Factors for Optimization
Successful high-throughput VIGS requires careful optimization of several parameters [4]:
- Plant Growth Conditions: Maintain plants at 20-25°C post-inoculation, as temperature significantly impacts silencing efficiency. Ensure appropriate humidity and photoperiod (e.g., 16h light/8h dark) [4] [48].
- Agroinoculum Concentration: Test a range of OD600 (e.g., 0.5 to 2.0) to find the optimal concentration that maximizes silencing while minimizing tissue damage.
- Plant Developmental Stage: Inoculate at the cotyledon or early true-leaf stage for robust systemic silencing.
The Scientist's Toolkit: Essential Research Reagents
Table 2: Key Reagents for High-Throughput VIGS Experiments
| Reagent / Material | Function / Purpose | Examples / Specifications |
|---|---|---|
| Viral Vectors | Delivery of target gene fragments into the host plant to initiate silencing. | TRV1 and TRV2 plasmids [4]; Engineered pTRV2-C2bN43 for enhanced silencing [48] |
| Agrobacterium Strain | Biological vector for delivering viral plasmids into plant cells. | Agrobacterium tumefaciens GV3101 [4] |
| Induction Agents | Activate virulence genes of Agrobacterium for efficient T-DNA transfer. | Acetosyringone (200 μM) in infiltration buffer [4] |
| Infiltration Buffer | Suspension medium for agrobacteria during inoculation. | 10 mM MgCl₂, 10 mM MES, pH 5.6 [4] |
| Selection Antibiotics | Maintain plasmid integrity in bacterial and plant cultures. | Kanamycin, Rifampicin, others as per vector and strain [4] |
| siRNA Nanoparticles | Species-independent delivery of siRNA for silencing in recalcitrant plants. | Gu+-siRNA NPs (Thiolated, ~200 nm diameter) [46] |
| qRT-PCR Reagents | Molecular validation and quantification of target gene silencing. | SYBR Green Master Mix, gene-specific primers, reverse transcriptase [48] |
Application in Gene Discovery and Breeding
High-throughput VIGS has proven instrumental in identifying genes governing agronomically vital traits. In pepper (Capsicum annuum), a species notoriously recalcitrant to stable transformation, VIGS has enabled large-scale functional screening [4] [48]. Key successes include the discovery of genes involved in:
- Fruit Quality: Color, biochemical composition, and pungency (capsaicinoid biosynthesis) [4].
- Biotic Stress Resistance: Defense against bacteria, oomycetes, and insects [4].
- Abiotic Stress Tolerance: Responses to temperature, salt, and osmotic stress [4] [46].
- Plant Architecture and Development: Regulation of growth habits and developmental timing [4].
- Reproductive Traits: Identification of transcription factors like CaAN2, which regulates anthocyanin biosynthesis in anthers, using the optimized TRV-C2bN43 system [48].
Furthermore, VIGS can induce heritable epigenetic modifications via RNA-directed DNA methylation (RdDM), leading to stable, transgenerational silencing of target genes. This application opens new avenues for epigenetic breeding and the development of non-transgenic plants with desired traits [5].
High-throughput functional genomics, spearheaded by advanced VIGS technologies, is fundamentally accelerating the pace of gene discovery. The continuous refinement of viral vectors, the innovative engineering of silencing suppressors, and the emergence of nanoparticle-based delivery systems are collectively enhancing the efficiency, scope, and applicability of this approach. As these tools are integrated with multi-omics technologies and breeding programs, they hold immense potential for rapid crop improvement, deeper understanding of gene regulatory networks, and the development of sustainable agricultural solutions. The central role of siRNA as the guiding molecule in these processes ensures that VIGS will remain a cornerstone of functional genomics for the foreseeable future.
Virus-Induced Gene Silencing (VIGS) is a powerful functional genomics tool that leverages the plant's innate RNA interference (RNAi) machinery to suppress target gene expression. The process is initiated by introducing a recombinant viral vector carrying a fragment of a host plant gene. Upon infection, the plant's defense system recognizes the viral RNA and processes it, triggering a sequence-specific silencing mechanism that targets both the viral RNA and the corresponding endogenous host mRNA for degradation. This technology provides a rapid, cost-effective alternative to stable transformation for characterizing gene function, particularly in species where genetic transformation remains challenging [4].
The core mechanism of VIGS relies on post-transcriptional gene silencing (PTGS). Double-stranded RNA (dsRNA) replication intermediates from the virus, or dsRNA formed from host-encoded transcripts, are cleaved by Dicer-like (DCL) enzymes. This cleavage generates 21- to 24-nucleotide small interfering RNAs (siRNAs). These siRNAs are then incorporated into the RNA-induced silencing complex (RISC), where they serve as guides for the sequence-specific cleavage and degradation of complementary messenger RNA (mRNA) transcripts. A key component of RISC is the Argonaute (AGO) protein, which executes the slicing activity [4] [49]. The siRNAs, fundamental to this process, can also act as mobile silencing signals, enabling the systemic spread of the silencing effect throughout the plant. Furthermore, the system can be amplified by host RNA-dependent RNA polymerases (RDRs), which use the cleaved RNA fragments as templates to generate secondary dsRNA, leading to the production of secondary siRNAs and a more robust, sustained silencing response [49].
VIGS Workflow and Signaling Pathways
The typical VIGS workflow involves the design of a viral vector, most commonly the bipartite Tobacco Rattle Virus (TRV), to carry a fragment of the target host gene. The TRV1 plasmid encodes viral replication and movement proteins, while the TRV2 plasmid carries the coat protein and the host gene insert. Recombinant Agrobacterium tumefaciens strains harboring these plasmids are cultured and co-infiltrated into plant leaves. The Agrobacterium transfers the T-DNA containing the viral genome into the plant cell, where viral replication and systemic movement commence, spreading the silencing signal [4] [50].
The following diagram illustrates the core molecular pathway of siRNA biogenesis and action within the plant cell during VIGS, culminating in the degradation of target viral and host mRNAs.
Figure 1: The Core siRNA Pathway in Virus-Induced Gene Silencing
Application of VIGS in Engineering Disease Resistance
VIGS has been extensively used to identify and validate plant genes involved in disease resistance pathways, enabling the development of crops with enhanced immunity. Functional screens in crops like pepper (Capsicum annuum L.) have successfully identified genes conferring resistance to bacterial, oomycete, and fungal pathogens [4]. The technology allows for the high-throughput knockdown of candidate resistance (R) genes and other immune components to assess their role in pathogen recognition and defense activation.
A key application is the dissection of the plant immune system, which comprises Pattern-Triggered Immunity (PTI) and Effector-Triggered Immunity (ETI). VIGS can be used to silence genes involved in the recognition of Pathogen-Associated Molecular Patterns (PAMPs), such as receptor-like kinases, or downstream signaling components like MAPK cascades. For instance, the role of callose deposition at plasmodesmata as a physical barrier to pathogen spread has been elucidated using VIGS. Research on viroid infection has shown that a virulent strain can suppress callose synthase genes via RNA silencing, a process that can be studied and potentially reversed using VIGS-based approaches [49]. By knocking down specific genes, researchers can determine their contribution to defense responses such as the production of reactive oxygen species, activation of pathogenesis-related (PR) genes, and the hypersensitive response (HR).
Table 1: VIGS-Idenfied Genes Involved in Disease Resistance Pathways
| Gene Category | Target Gene Example | Pathogen Type | Silencing Effect on Plant Phenotype |
|---|---|---|---|
| Immune Receptors | Resistance (R) genes | Bacteria, Oomycetes | Increased susceptibility; loss of effector-triggered immunity (ETI) [49] |
| Pattern Recognition | Receptor-like kinases (RLKs) | Fungi, Bacteria | Compromised PAMP recognition; reduced pattern-triggered immunity (PTI) [49] |
| Defense Signaling | MAPK cascade components | Multiple | Attenuated defense gene activation and callose deposition [49] |
| Structural Defense | Callose synthase genes | Viroids, Viruses | Enhanced pathogen spread due to impaired plasmodesmata sealing [49] |
| Antiviral RNAi | DCLs, AGOs, RDRs | Viruses, Viroids | Increased viral/viroid accumulation and symptom severity [49] |
Application of VIGS in Enhancing Abiotic Stress Tolerance
Beyond biotic stress, VIGS is a pivotal tool for unraveling the genetic networks that underpin plant responses to abiotic stresses such as drought, salinity, and extreme temperatures. By silencing genes suspected to be involved in stress perception, signaling, or tolerance mechanisms, researchers can rapidly assess their functional importance. In pepper, VIGS has been deployed to identify genes governing tolerance to osmotic, salt, and temperature stress [4].
Recent advances in siRNA delivery have expanded the potential of VIGS for abiotic stress research. For example, the development of guanidinium-containing disulfide nanoparticle (Gu+-siRNA NPs) systems enables efficient, species-independent delivery of siRNAs. This technology has been successfully used to silence key genes like EIL1 and EIL2 in Arabidopsis and rice, resulting in enhanced salt tolerance [46]. This nanoparticle-based approach facilitates long-distance silencing, allowing for whole-plant phenotypic analysis in response to abiotic stress conditions. The ability to systemically silence genes in this way provides a more complete picture of their role in complex physiological adaptations to environmental challenges.
Table 2: VIGS Applications in Abiotic Stress Tolerance Research
| Abiotic Stress | Target Gene/Family | Experimental Outcome of Silencing | Key Findings |
|---|---|---|---|
| Salt Stress | EIL1, EIL2 | Enhanced salt tolerance in Arabidopsis and rice [46] | Identified negative regulators of salt adaptation. |
| Salt Stress | Ion transporters, SOS pathway genes | Altered ion homeostasis and plant survival [4] | Validated function of key salinity response genes. |
| Temperature Stress | Heat-shock factors (HSFs), Chaperones | Modified thermotolerance in pepper [4] | Uncovered genes critical for protein stability under heat stress. |
| Drought/Osmotic Stress | Osmoprotectant biosynthesis genes | Altered osmotic adjustment and wilting response [4] | Confirmed role of specific metabolites in water retention. |
Essential Protocols for VIGS Experiments
The reliability of VIGS data hinges on rigorous and optimized experimental protocols. Below is a detailed methodology for conducting a TRV-based VIGS experiment in solanaceous plants like Nicotiana benthamiana and tomato.
Agrobacterium Preparation and Plant Inoculation
Day 1: Streak Agrobacterium tumefaciens strains (e.g., GV3101) harboring the pTRV1 and pTRV2 plasmids (including pTRV2-PDS control and pTRV2-target gene) onto LB agar plates containing appropriate antibiotics (50 μg/mL kanamycin, 100 μg/mL rifampicin). Incubate plates at 30°C for 48 hours [50].
Day 3: Inoculate a 2-3 mL liquid culture of LB media with the same antibiotics for each strain. Incubate by shaking at 30°C and 200 r.p.m. for 16-18 hours (primary culture) [50].
Day 4: Dilute the primary culture 1:25 into a secondary liquid Induction Media (IM) containing antibiotics and 200 μM acetosyringone. Incubate by shaking at 30°C and 200 r.p.m. for 20-24 hours [50].
Day 5:
- Harvest the bacterial cells by centrifugation at 3000 x g for 10 minutes.
- Resuspend the pellet in an infiltration buffer (10 mM MgCl₂, 10 mM MES, pH 5.5) to an optical density (OD₆₀₀) of 0.3.
- Add acetosyringone to the pTRV1 culture to a final concentration of 400 μM.
- Mix the pTRV1 and pTRV2 (or pTRV2-target gene) cultures in a 1:1 ratio. The final OD₆₀₀ for each should be 0.15, and the final acetosyringone concentration 200 μM.
- Incubate the mixture at room temperature for 3-4 hours.
- For infiltration, use a needleless syringe to inject the bacterial suspension into the leaves of young plants. For tomato, infiltrate both cotyledons; for N. benthamiana, infiltrate the two largest true leaves [50].
Critical Factors for Optimization
The efficiency of VIGS is influenced by several factors that must be carefully controlled:
- Plant Developmental Stage: N. benthamiana plants should be ~2.5 weeks old with cotyledons and the first 2-4 true leaves. Tomato plants should be used 7-8 days post-emergence, before true leaves appear [50].
- Insert Design: The fragment inserted into pTRV2 should be 200-1000 bp in length, avoid homopolymeric regions (e.g., polyA tails), and have sufficient homology to the target gene [50].
- Environmental Conditions: After infiltration, plants should be maintained at 20-22°C with a 16-hour day length and ~50% relative humidity for optimal viral spread and silencing [4] [50].
- Controls and Validation: Always include a positive control (e.g., TRV2-PDS, which causes photobleaching). Silencing efficiency must be quantified by measuring target gene transcript abundance using RT-PCR or qRT-PCR, ensuring one primer binds outside the region targeted for silencing [50].
The Scientist's Toolkit: Key Research Reagents
Successful implementation of VIGS relies on a suite of specialized reagents and biological materials. The following table details essential components for a standard TRV-based VIGS experiment.
Table 3: Essential Reagents for TRV-based VIGS Experiments
| Reagent / Material | Function and Description | Example / Specification |
|---|---|---|
| TRV Viral Vectors | Bipartite system for delivering silencing trigger; TRV1 for replication/movement, TRV2 for coat protein and target gene insert [4] [50]. | pTRV1, pTRV2 (Gateway-compatible versions available) |
| Agrobacterium tumefaciens | Plant pathogen used as a biological vector to deliver T-DNA containing the TRV constructs into plant cells [50]. | Strain GV3101 (for tomato, N. benthamiana) |
| Antibiotics | Selection for bacterial cultures to maintain plasmid integrity. | Kanamycin (50 µg/mL), Rifampicin (100 µg/mL) [50] |
| Acetosyringone | Phenolic compound that induces the Agrobacterium vir genes essential for T-DNA transfer [50]. | 200 µM final concentration in infiltration mix |
| Infiltration Buffer | Resuspension medium for Agrobacterium cells, providing optimal pH and ionic conditions for infiltration. | 10 mM MgCl₂, 10 mM MES, pH 5.5 [50] |
| Positive Control Silencing Construct | Validates the entire VIGS protocol is working by producing a clear, visible phenotype. | TRV2-PDS (Phytoene desaturase) [50] |
Advanced Delivery Systems and Future Perspectives
While traditional Agrobacterium-mediated VIGS is powerful, new delivery technologies are emerging to overcome limitations like species dependency and variable efficiency. A promising advancement is the use of nanoparticles for siRNA delivery. For instance, guanidinium-containing disulfide nanoparticles (Gu+-siRNA NPs) can protect siRNAs from degradation and facilitate their efficient uptake and long-distance movement in plants. This system has demonstrated high biocompatibility, low toxicity, and effective gene silencing in both monocot and dicot species, representing a species-independent alternative for systemic gene silencing [46].
The future of VIGS in crop improvement lies in its integration with other technologies. Combining VIGS with high-throughput phenotyping and multi-omics analyses (transcriptomics, proteomics) will accelerate the functional annotation of genes controlling complex traits like disease resistance and stress tolerance [4]. Furthermore, VIGS can serve as a validation tool for genes identified through genome-wide association studies (GWAS) or as a preliminary screen prior to the deployment of more permanent genome editing techniques like CRISPR/Cas9. As delivery methods improve and our understanding of RNAi machinery deepens, VIGS will remain an indispensable component of the plant biologist's toolkit for rapid, transient gene functional analysis and targeted crop improvement.
The validation of therapeutic targets is a critical bottleneck in the drug discovery pipeline, requiring robust, scalable, and physiologically relevant models. Within this landscape, small interfering RNA (siRNA) has emerged as a transformative technology for sequence-specific gene silencing. siRNA harnesses the endogenous RNA interference (RNAI) pathway, a conserved mechanism in eukaryotes that mediates gene silencing through the degradation of complementary messenger RNA (mRNA) transcripts [24] [51]. The core of this mechanism involves the loading of a 21-25 nucleotide siRNA guide strand into the RNA-induced silencing complex (RISC), which then directs the cleavage of the target mRNA [52] [53].
Virus-Induced Gene Silencing (VIGS) represents a powerful application of this mechanism in plant systems. VIGS utilizes recombinant viral vectors to deliver gene fragments into plant cells, triggering the plant's post-transcriptional gene silencing (PTGS) machinery and leading to systemic suppression of endogenous genes [4] [5]. This technology has become an indispensable reverse genetics tool for functional genomics, allowing for rapid characterization of gene function in a wide array of plant species, from model organisms like Nicotiana benthamiana to crops such as pepper (Capsicum annuum L.) and cotton (Gossypium hirsutum) [4] [54]. This technical guide delineates the molecular principles and methodologies of siRNA-mediated silencing, with a focus on VIGS in plant models, and explores its direct implications and translational applications for validating targets in biomedical research.
Molecular Mechanisms of siRNA and VIGS
The Core RNAi Pathway
The RNAi pathway is initiated when long double-stranded RNA (dsRNA) is processed by a Dicer or Dicer-like (DCL) enzyme into short 21-24 nucleotide siRNA duplexes [5] [51]. One strand of this duplex, the guide strand, is then incorporated into the Argonaute (AGO) protein within the RISC. The siRNA-loaded RISC scans cellular mRNAs and catalyzes the cleavage of those sequences perfectly complementary to the siRNA guide strand [24] [52]. This sequence-specific degradation leads to a reduction in the corresponding protein levels, effectively silencing the gene.
The VIGS Workflow
In VIGS, this native cellular defense mechanism is co-opted for functional genomics. The experimental workflow involves engineering a viral vector to carry a fragment (typically 100-300 bp) of the plant gene to be silenced [4] [8]. This recombinant vector is then introduced into the plant, often via Agrobacterium tumefaciens-mediated transformation (agroinfiltration) [4] [54]. As the virus replicates and moves systemically through the plant, it produces dsRNA replicative intermediates. The plant's DCL enzymes recognize and process these dsRNAs into virus-derived siRNAs (vsiRNAs). A subset of these vsiRNAs are perfectly complementary to the endogenous target gene's mRNA, leading to its degradation by RISC and the manifestation of a loss-of-function phenotype that can be studied to deduce gene function [4] [5]. The following diagram illustrates this integrated process.
Key Viral Vectors for VIGS
The success of VIGS is heavily dependent on the choice of viral vector. Different vectors offer distinct advantages and have specific host ranges. The table below summarizes the commonly used vectors in VIGS research.
Table 1: Key Viral Vectors Used in Virus-Induced Gene Silencing
| Viral Vector | Virus Type | Key Features | Primary Applications |
|---|---|---|---|
| Tobacco Rattle Virus (TRV) | RNA virus | Bipartite genome (TRV1, TRV2); broad host range; efficient systemic movement; mild symptoms [4]. | One of the most versatile systems, especially for Solanaceae family plants (e.g., pepper, tomato) [4] [54]. |
| Potato Virus X (PVX) | RNA virus | ~6.4 kb single-stranded RNA genome; used for expressing syn-tasiRNA precursors [24]. | A versatile vector for transient gene expression and silencing, particularly in Nicotiana benthamiana [24]. |
| Geminiviruses(e.g., CLCrV, ACMV) | DNA virus | Single-stranded DNA genome; can infect meristematic tissues [4]. | Useful for targeting genes in developing tissues and for studying DNA virus-host interactions [4]. |
Technical Guide: Experimental Protocols for VIGS
A Generalized VIGS Protocol Using TRV Vectors
The following protocol is adapted from established methods for TRV-based VIGS in various plant species, including cotton and sunflower [8] [54].
Principle: The bipartite TRV system is used. TRV1 encodes proteins for replication and movement, while TRV2 carries the coat protein and the insert from the target plant gene. Agrobacterium strains harboring these two plasmids are mixed and infiltrated into plant tissues, leading to viral infection and subsequent silencing of the target gene.
Materials:
- Agrobacterium tumefaciens strain GV3101
- TRV1 plasmid (e.g., pYL192; Addgene #148968)
- TRV2 plasmid (e.g., pYL156; Addgene #148969) with and without the target gene insert
- Antibiotics: Kanamycin, Gentamicin, Rifampicin
- Induction buffer: 10 mM MES, 10 mM MgCl₂, 200 µM acetosyringone
- Needleless syringe or vacuum infiltration apparatus
Procedure:
- Vector Construction: Clone a 100-300 bp fragment of the target gene into the multiple cloning site of the TRV2 vector using standard molecular biology techniques (e.g., restriction enzyme digestion and ligation, Gateway cloning, or Gibson Assembly) [8].
- Agrobacterium Preparation:
- Transform the recombinant TRV2 and the helper TRV1 plasmids separately into Agrobacterium strain GV3101.
- Plate on LB agar with appropriate antibiotics (e.g., 50 µg/mL kanamycin, 25 µg/mL gentamicin) and incubate at 28°C for 2 days [54].
- Inoculate single colonies into liquid LB with antibiotics and shake overnight at 28°C.
- Sub-culture the bacteria into fresh, antibiotic-supplemented LB with 10 mM MES and 20 µM acetosyringone. Grow until OD₆₀₀ reaches 0.8-1.2.
- Agroinfiltration:
- Harvest bacterial cells by centrifugation and resuspend in induction buffer to a final OD₆₀₀ of 1.5. Incubate at room temperature for 3-4 hours.
- Mix the TRV1 and TRV2 cultures in a 1:1 ratio.
- For cotyledon infiltration: Use a needle to lightly wound the abaxial side of the cotyledons of 7-14 day-old seedlings. Gently press a needleless syringe containing the agrobacterial mixture against the leaf and infiltrate [54].
- For seed vacuum infiltration: Peel the seed coats and subject the seeds to vacuum infiltration with the agrobacterial mixture, followed by a co-cultivation period (e.g., 6 hours) [8].
- Plant Incubation: Maintain infiltrated plants under controlled environmental conditions (e.g., 22-25°C, 16/8h light/dark cycle) with high humidity for 1-2 days post-infiltration before returning to standard growth conditions.
- Phenotypic and Molecular Analysis:
- Silencing phenotypes typically become visible 2-4 weeks post-infiltration.
- Validate gene knockdown using reverse-transcription quantitative PCR (RT-qPCR). It is critical to use stable reference genes (e.g., GhACT7 and GhPP2A1 in cotton under biotic stress) for accurate normalization [54].
Advanced Silencing Tools: Synthetic tasiRNAs
A recent innovation is the use of synthetic trans-acting small interfering RNAs (syn-tasiRNAs). These are highly specific 21-nt siRNAs designed to be processed from a minimal, modified TAS precursor RNA.
Protocol Overview [24]:
- Computational Design: Design syn-tasiRNA sequences with high specificity to the target mRNA and minimal off-target potential using bioinformatic tools.
- Cloning: Clone the designed syn-tasiRNA sequence into a minimal precursor backbone (e.g., containing a 22-nt miRNA target site, an 11-nt spacer, and the 21-nt syn-tasiRNA) using Golden Gate or Gibson Assembly. This minimal precursor (~54 nt) can be inserted into standard plant expression vectors or viral vectors like PVX.
- Delivery: Deliver the construct to plants via:
- Agrobacterium-mediated transient expression.
- A transgene-free method: Infect plants with recombinant PVX expressing the minimal precursor, then homogenize the systemically infected leaves and spray the crude extract onto new plants to induce whole-plant gene silencing [24].
The Scientist's Toolkit: Essential Research Reagents
Successful implementation of VIGS and siRNA-based research relies on a core set of reagents and biological materials. The following table details these essential components.
Table 2: Key Research Reagents for VIGS and siRNA Studies
| Reagent / Material | Function / Role | Examples & Notes |
|---|---|---|
| Viral Vectors | Delivers the target gene fragment into plant cells to initiate silencing. | TRV (pYL192, pYL156), PVX-based vectors. Selection depends on host plant [4] [8]. |
| Agrobacterium Strain | Mediates the delivery of the viral vector DNA into the plant cell. | GV3101 is a commonly used disarmed strain for agroinfiltration [8] [54]. |
| Antibiotics | Selective maintenance of plasmids in bacterial and plant cultures. | Kanamycin (for TRV vectors), Gentamicin, Rifampicin [24] [54]. |
| Induction Compounds | Activates the Agrobacterium Vir genes for efficient T-DNA transfer. | Acetosyringone, MES buffer [54]. |
| Stable Reference Genes | Critical for accurate normalization in RT-qPCR validation of silencing. | GhACT7 & GhPP2A1 (in cotton-aphid studies); must be validated for specific system [54]. |
| Chemical Modification Tools | Enhances stability and efficacy of therapeutic siRNAs; reduces off-target effects. | 2'-O-methyl (2'-OMe), 2'-fluoro (2'-F) ribose modifications [52]. |
Translational Applications: From Plant Models to Biomedical Research
The principles of siRNA and VIGS have direct translational applications in biomedical research and drug development.
Validating Drug Targets in Plants
Plant VIGS serves as a high-throughput platform for preliminary target validation. It has been successfully used to identify genes governing complex traits such as:
- Disease Resistance: Silencing susceptibility genes to enhance resistance against bacteria, oomycetes, and insects [4] [51].
- Abiotic Stress Tolerance: Identifying genes involved in responses to temperature, salt, and osmotic stress [4] [5].
- Metabolic Engineering: Characterizing genes in specialized metabolic pathways, such as capsaicinoid biosynthesis in pepper [4].
The insights gained from these plant studies can inform the selection of homologous targets in mammalian systems for therapeutic intervention.
siRNA as a Direct Therapeutic Modality
Chemically modified siRNAs have matured into a validated drug class. As of 2024, six siRNA therapeutics have gained regulatory approval, primarily for treating metabolic and genetic diseases in the liver [53].
Table 3: Clinical Landscape of siRNA Therapeutics (Data up to 2024)
| Area | Number of Clinical Trials | Key Approved Drugs | Primary Targets | Key Challenges |
|---|---|---|---|---|
| Non-Oncology | ~90% of 424 global trials | Patisiran, Inclisiran, Givosiran | PCSK9, TTR, ALAS1, HAO1 [53]. | Delivery to extra-hepatic tissues. |
| Oncology | ~10% of trials, later initiation | None approved as of 2024 | CSF2, PTGS2, TGFB1 [53]. | Complex tumor biology, inefficient targeted delivery to extra-hepatic tissues [52] [53]. |
Key factors impacting the efficacy of therapeutic siRNAs, as identified through systematic analyses, include:
- Chemical Modification Pattern: The level of 2'-O-methyl content significantly impacts efficacy and stability in vivo [52].
- Target mRNA Context: Native mRNA features such as exon usage, polyadenylation site selection, and ribosomal occupancy can greatly influence siRNA performance, underscoring the need for testing in a native context [52].
- Delivery Systems: Nanocarriers (e.g., liposomes, polymeric nanoparticles) are revolutionizing siRNA delivery by protecting dsRNA from degradation and enhancing cellular uptake, a strategy also being explored in plants via Spray-Induced Gene Silencing (SIGS) [51].
siRNA-mediated technologies, exemplified by VIGS in plants, provide a robust and versatile framework for therapeutic target validation across biological kingdoms. The experimental paradigms established in plant models—ranging from vector design and delivery to mechanistic analysis of silencing—offer invaluable blueprints for developing RNAi-based therapies in humans. Future progress will be driven by advances in the design of highly specific silencing triggers like syn-tasiRNAs [24], the development of sophisticated delivery vehicles such as nanocarriers [51], and a deeper understanding of the factors governing siRNA efficacy in different cellular contexts [52] [53]. The integration of these insights from basic plant science into biomedical research promises to accelerate the development of novel, precise therapeutic interventions.
Optimizing Silencing Efficiency: Challenges and Solutions
The discovery of RNA interference (RNAi) has revolutionized functional genomics and therapeutic development, offering unprecedented potential for sequence-specific gene silencing. Short interfering RNAs (siRNAs), the key effectors of the RNAi pathway, are double-stranded RNA molecules typically 19-23 nucleotides in length that guide the RNA-induced silencing complex (RISC) to complementary mRNA targets for degradation [55]. While Virus-Induced Gene Silencing (VIGS) and other RNAi-based approaches have become powerful tools for studying gene function, their application is significantly constrained by off-target effects—the unintended silencing of genes with partial complementarity to the siRNA [56]. These effects can obscure experimental results, lead to misinterpretation of phenotypic data, and pose substantial safety risks in therapeutic contexts. This technical guide examines the molecular basis of off-target effects and presents integrated computational, chemical, and experimental strategies to mitigate them, with particular emphasis on their relevance to VIGS research.
Molecular Mechanisms of Off-Target Effects
Off-target silencing occurs through multiple mechanisms, primarily mediated by sequence complementarity between the siRNA and non-target transcripts. The seed region (nucleotides 2-8 from the 5' end of the guide strand) plays a particularly crucial role, as partial complementarity in this region can lead to miRNA-like translational repression or mRNA degradation [57] [52].
Computational analysis reveals the alarming prevalence of potential off-targets. Studies examining 25 plant species found that approximately 50% to 70% of gene transcripts have potential off-targets when used for post-transcriptional gene silencing (PTGS), with experimental validation confirming that up to 50% of predicted off-target genes were actually silenced [56]. In Arabidopsis, the average number of predicted off-targets per trigger sequence was 3.9, with an average off-target region length of 41.2 nucleotides [56].
The diagram below illustrates the primary molecular mechanisms through which siRNA off-target effects occur:
The thermodynamic properties of siRNAs significantly influence off-target potential. siRNAs with lower thermodynamic stability in the seed region exhibit reduced off-target effects while maintaining on-target potency [58] [59]. This occurs because the RISC loading complex more efficiently incorporates guide strands with lower 5' stability, and the seed region's accessibility affects non-perfect matching to off-target transcripts.
Computational Design Strategies for Specificity
Algorithmic Design Principles
Rational siRNA design incorporates established rules that maximize target specificity while minimizing off-target potential. The following table summarizes key design rules and their molecular basis:
Table 1: Established siRNA Design Rules for Minimizing Off-Target Effects
| Design Rule | Molecular Basis | Specificity Contribution |
|---|---|---|
| Ui-Tei Rules [60] | 5' terminus of antisense strand must contain A/U; 5' terminus of sense strand must include G/C; A/U content in 5' terminal 7 bp | Promotes correct strand selection; reduces off-target incorporation |
| Amarzguioui Rules [60] | 5' terminal 7 base pairs should have at least 4 A/U residues; GC stretch no longer than 9 nucleotides | Optimizes thermodynamic asymmetry for RISC loading |
| Reynolds Rules [60] | A/U differential of duplex end should be >0; specific nucleotide preferences at positions 3, 10, 13, 19 | Enhances specificity through position-dependent nucleotide effects |
| Seed-Target Duplex Tm [58] [60] | Melting temperature of seed-target duplex <21.5°C | Reduces miRNA-like off-target effects through seed region binding |
| Off-Target Filtering [56] [52] | ≥2 mismatches to any non-targeted transcripts; exclude sequences with CCCC or GGGG stretches | Prevents unintended targeting of partially complementary sequences |
Bioinformatics Tools for Off-Target Prediction
Several computational tools have been developed specifically to identify potential off-targets during PTGS construct design:
siRNA Scan: A web-based computational tool that identifies potential off-targets by searching for identical or reverse complementary regions of ≥21 nt between a trigger sequence and all other genes in the genome [56]. The tool provides an integrated sequence similarity search environment for both plant and animal species.
siDirect 2.0: Implements Ui-Tei, Amarzguioui, and Reynolds algorithms while keeping the seed-target duplex Tm below 21.5°C as a default parameter [58] [60]. It selects siRNA sequences that have at least two mismatches to any other non-targeted transcripts.
RNAxs Webserver: Utilizes predefined parameters including accessibility threshold values for seed region (0.01157) and full siRNA (0.001002), self-folding energy (0.9022), and sequence/energy asymmetry (0.5 and 0.4655 respectively) for functional siRNA design [60].
The following workflow illustrates the integrated computational design process for specificity-enhanced siRNAs:
Chemical Modification Strategies for Enhanced Specificity
Chemical modifications represent a powerful approach to enhance siRNA specificity while addressing other pharmacological challenges. The strategic incorporation of modified nucleotides can significantly reduce off-target effects by modulating siRNA structure, thermodynamics, and interaction with the RNAi machinery.
Ribose Modifications
Ribose modifications at the 2'-position of the nucleotide sugar represent the most extensively studied and applied chemical alterations for improving siRNA properties:
Table 2: Ribose Modifications for Reducing siRNA Off-Target Effects
| Modification | Impact on Specificity | Structural Basis | Optimal Positioning |
|---|---|---|---|
| 2'-O-methyl (2'-OMe) [61] [62] | Reduces immune stimulation; improves nuclease resistance; decreases off-target silencing | Maintains C3'-endo sugar pucker; enhances thermal stability of A-form duplex | Seed region (positions 2-8 of guide strand); passenger strand |
| 2'-fluoro (2'-F) [61] [62] | Increases binding affinity to target; reduces seed-mediated off-target effects | Strengthens Watson-Crick base pairing; improves duplex thermostability | Distributed pattern throughout duplex |
| Locked Nucleic Acid (LNA) [62] [57] | Significantly increases specificity and potency; reduces seed-based off-targeting | Ribose locked in C3'-endo conformation through methylene bridge; extremely high binding affinity | Selective placement at 3-5 positions, avoiding seed region |
| Unlocked Nucleic Acid (UNA) [62] | Reduces thermodynamic stability; decreases seed-mediated off-target effects | Flexible open-chain ribose structure; decreases melting temperature | Strategic placement in seed region to modulate stability |
| 2'-O-methoxyethyl (2'-MOE) [61] | Enhances nuclease resistance; reduces immunostimulatory potential | Increased steric bulk and hydration; maintains RNA-like conformation | 3' and 5' termini; passenger strand |
Backbone and Nucleobase Modifications
Beyond ribose modifications, alterations to the phosphate backbone and nucleobases provide additional avenues for optimizing siRNA specificity:
Phosphorothioate (PS) linkages: Replacement of non-bridging oxygen with sulfur in the phosphate backbone increases nuclease resistance and improves pharmacokinetics, allowing for reduced dosage and consequently lower off-target potential [62] [55]. Optimal placement typically involves 1-3 modifications at the 3' and 5' termini.
5'-(E)-vinylphosphonate (5'-E-VP): This metabolically stable 5'-phosphate modification increases binding affinity to AGO2 while enhancing metabolic stability, leading to improved on-target activity at lower concentrations [57].
Nucleobase modifications: Strategic modifications to nucleobases can fine-tune siRNA specificity. For example, the introduction of 6-mCEPh-purine at the 5'-end of the guide strand creates additional interactions with the AGO2 MID domain (specifically with Tyr529), enhancing on-target activity and specificity [57]. Similarly, 8-Br AMP modification facilitates specific hydrophobic interactions with AGO2, improving binding affinity and selectivity [57].
Modification Patterns and Strategic Placement
The pattern of chemical modifications throughout the siRNA duplex significantly impacts both specificity and potency. Research indicates that fully modified siRNAs (with 2'-OMe or 2'-F modifications at all positions) can dramatically improve metabolic stability but may compromise RISC loading and activity if not properly designed [52]. Current optimal strategies employ:
- Asymmetric modification patterns: Heavier modification of the passenger strand compared to the guide strand to facilitate RISC unloading [55]
- Seed region modulation: Strategic placement of destabilizing modifications (e.g., UNA) or specificity-enhancing modifications (e.g., LNA) in the seed region to reduce off-target effects [62]
- Terminal stabilization: Modification of 3' and 5' termini with nuclease-resistant analogs (e.g., PS linkages, 2'-OMe) to prolong half-life [61]
Experimental Validation and Screening Protocols
In silico Validation Workflow
Before experimental validation, comprehensive computational assessment is essential:
Step 1: Off-target prediction analysis
- Use siRNA Scan with genome/transcriptome databases to identify potential off-targets [56]
- Perform BLAST analysis against relevant genomic databases (e.g., NCBI RefSeq)
- Calculate seed complementarity to all transcripts using position-specific scoring matrices
Step 2: Molecular docking studies
- Model siRNA-AGO2 interactions using crystal structures (e.g., PDB ID: 3LUD) [57]
- Assess binding affinity and orientation of modified nucleotides in the MID domain
- Calculate free energy of binding for candidate siRNAs
Step 3: Thermodynamic profiling
- Calculate melting temperature (Tm) and free energy (ΔG) for seed-target duplexes
- Ensure seed-target duplex Tm remains below 21.5°C [58] [60]
- Analyze internal stability profile across the siRNA duplex
Experimental Validation Protocols
Protocol 1: Transcriptome-wide off-target assessment
- Transfert target cells with candidate siRNAs using appropriate reagents (e.g., Lipofectamine RNAiMAX, Interferin) [62]
- Isolate total RNA 24-48 hours post-transfection using TRIzol reagent
- Perform RNA sequencing (RNA-seq) with minimum 30 million reads per sample
- Analyze differential expression using DESeq2 or similar tools (threshold: fold change >2, adjusted p-value <0.05)
- Validate potential off-targets using quantitative RT-PCR with SYBR Green chemistry [56]
Protocol 2: Specificity index calculation
- Determine IC50 values for on-target silencing using dose-response curves (typically 0.1-100 nM range)
- Identify top 10 potential off-targets through computational prediction
- Measure off-target silencing at concentration yielding 80% on-target silencing
- Calculate specificity index as: SI = (On-target IC50) / (Average off-target IC50)
- Acceptable SI value: >10-fold specificity for on-target versus off-target effects [52]
Table 3: Key Research Reagents and Tools for siRNA Specificity Enhancement
| Tool/Reagent | Function | Application Notes |
|---|---|---|
| siRNA Scan [56] | Computational prediction of potential off-target genes | Web-based tool; supports 25+ plant and animal species |
| siRNAmod Database [62] | Repository of 4894 chemically modified siRNAs with efficacy data | Contains 128 unique chemical modifications; enables design optimization |
| 2'-OMe Phosphoramidites [61] [52] | Solid-phase synthesis of modified siRNAs | Most common modification (17% of reported modifications); natural analog |
| 2'-F Phosphoramidites [61] [52] | Enhanced stability and binding affinity | Second most common ribose modification (7% of reported modifications) |
| LNA Phosphoramidites [62] | Extreme thermal stabilization and specificity enhancement | Use selectively (3-5 positions per strand) to avoid over-stabilization |
| Lipofectamine RNAiMAX [62] | Transfection reagent for siRNA delivery | Common choice for in vitro screening; minimal immune activation |
| Dual-Luciferase Reporter System [52] | High-throughput assessment of on/off-target effects | Enables rapid screening of modification patterns |
The strategic integration of computational design and chemical modification represents a powerful approach to overcome off-target effects in siRNA applications, including VIGS research. By combining rational in silico design employing established rules and prediction tools with strategic incorporation of specificity-enhancing chemical modifications, researchers can significantly improve the reliability of gene silencing data and the safety profile of therapeutic siRNAs. The continued development of modified nucleotide chemistries and refinement of computational prediction algorithms will further enhance our ability to design highly specific siRNAs, ultimately expanding the utility of RNAi technologies in both basic research and clinical applications.
Within the framework of investigating the role of small interfering RNA (siRNA) in Virus-Induced Gene Silencing (VIGS) research, achieving efficient systemic spreading of the silencing signal is a paramount objective. The efficacy of VIGS is fundamentally governed by the delivery and movement of the viral vector throughout the plant, which in turn dictates the distribution and potency of siRNA molecules that mediate gene silencing [4]. This technical guide delves into two pivotal strategic fronts for enhancing this systemic spreading: the engineering of the viral vectors themselves and the deployment of viral suppressor proteins (VSRs). The interplay between vector mobility and the host's RNA interference (RNAi) machinery determines the success of functional genomics studies, making the optimization of systemic spreading a critical endeavor for researchers and drug development professionals aiming to harness RNAi for therapeutic and biotechnological applications [4] [8].
siRNA and the Core Mechanism of VIGS
Virus-Induced Gene Silencing is a powerful technique that co-opts the plant's innate post-transcriptional gene silencing (PTGS) antiviral defense mechanism to target endogenous plant genes [4]. The process is initiated when a recombinant viral vector, carrying a fragment of a host gene of interest, is introduced into the plant. Upon infection, the viral RNA replicates, generating double-stranded RNA (dsRNA) intermediates.
These dsRNA molecules are recognized and processed by the host enzyme Dicer-like (DCL) into a pool of 21- to 24-nucleotide small interfering RNAs (siRNAs) [4]. These siRNAs are the key mobile silencing signals. They are loaded into an RNA-induced silencing complex (RISC), which uses the siRNA guide strand to identify and cleave complementary mRNA sequences—both viral and endogenous—leading to targeted gene knockdown [4] [24]. The systemic nature of VIGS relies on the movement of these siRNAs between cells, a process facilitated by viral movement proteins and the plant's own symplastic transport systems [63].
The following diagram illustrates the core siRNA pathway in VIGS, from viral infection to systemic gene silencing.
Engineering Viral Vectors for Enhanced Movement
The design of the viral vector is a primary determinant of its ability to spread systemically and induce robust silencing. Key engineering strategies focus on modifying components critical for viral movement and packaging.
Optimization of Viral Movement Proteins (MPs)
Movement Proteins are essential for the cell-to-cell and long-distance movement of viruses through plasmodesmata and the phloem. Engineering the MP can directly enhance the spread of the VIGS vector and the associated siRNAs.
A significant advancement involves the modification of MPs from the 30K family. A 2024 study demonstrated that small heterologous sequences (e.g., 18-54 bp fragments of a target gene) can be inserted in-frame directly into the coding sequence of the MP of viruses like Alfalfa Mosaic Virus (AMV), Cucumber Mosaic Virus (CMV), and Tobacco Mosaic Virus (TMV) without abolishing its function [63]. This strategy not only creates a new VIGS vector but also establishes a correlation between the size of the inserted fragment and the efficiency of gene silencing, allowing for the calibration of silencing levels. Smaller inserts (21-39 nt) achieved around 45% silencing, while fragments of 45 nt or larger achieved 75-90% silencing efficiency [63].
Furthermore, the study revealed that high efficiency of viral encapsidation, mediated by the Coat Protein (CP), can sometimes reduce silencing efficacy, possibly by protecting the viral RNA from DCL processing. Therefore, vectors designed with reduced CP activity might, in some cases, promote better siRNA amplification and spread [63].
Selection and Design of Viral Backbones
The choice of viral backbone is crucial and must be matched to the host plant. The Tobacco Rattle Virus (TRV) system is one of the most widely used due to its broad host range, efficient systemic movement, and ability to target meristematic tissues, making it a versatile tool, especially in Solanaceae plants like pepper and tobacco [4] [8].
For the vector to function, specific structural elements are required. The TRV system, for instance, is bipartite, requiring two vectors:
- TRV1: Encodes replicase proteins and a movement protein for virus replication and spread.
- TRV2: Contains the coat protein gene and a multiple cloning site (MCS) for inserting the target gene fragment, playing the key role in initiating silencing [4].
The design of the insert itself is also critical. Inserts are typically between 100-500 base pairs in length and must be carefully selected to ensure high specificity and minimize off-target effects [4]. Using bioinformatic tools to predict effective siRNA target sites within the gene of interest is a standard practice to improve silencing efficiency [8].
Table 1: Key Viral Vectors and Their Engineering Strategies for Enhanced Spreading
| Viral Vector | Genome Type | Key Engineering Strategy | Impact on Systemic Spreading & Silencing |
|---|---|---|---|
| Tobacco Rattle Virus (TRV) | RNA | Bipartite system (TRV1/TRV2); broad host range optimization; subgenomic promoters for insert expression. | Efficient systemic movement, including to meristems; mild symptoms; highly versatile [4]. |
| Alfalfa Mosaic Virus (AMV) | RNA | In-frame insertion of small inserts (18-54 bp) into the 30K-family Movement Protein. | Allows calibration of silencing levels; insert size directly correlates with silencing efficiency (45-90%) [63]. |
| Cucumber Mosaic Virus (CMV) | RNA | Insertion in MP (30K family) or replacement of 2b protein sequence (a VSR). | Modifies viral movement and suppression dynamics, potentially enhancing spread and siRNA accumulation [4] [63]. |
| Potato Virus X (PVX) | RNA | Delivery of minimal synthetic trans-acting siRNA (syn-tasiRNA) precursors. | Enables highly specific, multiplexed silencing from a compact vector, suitable for topical application [24]. |
| Geminiviruses (e.g., CLCrV) | DNA | Use of satellite virus-based systems to carry target gene inserts. | Useful for plants resistant to RNA viruses; can provide an alternative route for siRNA production [4]. |
Harnessing Viral Suppressors of RNAi (VSRs)
Plants have evolved a sophisticated RNAi defense against viruses, and in turn, viruses have encoded Viral Suppressors of RNA silencing (VSRs) to counteract this defense. In VIGS, these VSRs can be strategically deployed to transiently inhibit the host's silencing machinery, thereby allowing the viral vector to replicate and spread more effectively before being degraded, ultimately leading to a stronger and more widespread silencing signal [4].
Mechanism of Action of Key VSRs
Different VSRs act at distinct points in the RNAi pathway:
- P19 (from Tombusviruses): Binds and sequesters siRNA duplexes, preventing their loading into the RISC complex. This is one of the most potent and commonly used VSRs in VIGS research, particularly in Nicotiana benthamiana [4].
- HC-Pro (from Potyviruses): Inhibits siRNA amplification by preventing the activity of RNA-dependent RNA polymerases (RDRs) and may also hinder systemic signaling.
- 2b (from Cucumoviruses): Binds to and inhibits Argonaute (AGO) proteins, the catalytic components of the RISC complex, thus directly interfering with the silencing effector step [4].
The diagram below illustrates how these suppressors interfere with the siRNA pathway to enhance VIGS.
Application and Optimization of VSRs
The efficacy of a given VSR can vary significantly between plant species, necessitating empirical testing [4]. Co-expressing a well-characterized VSR like P19 or C2b with the VIGS vector in Agrobacterium-mediated infiltration can dramatically enhance silencing efficiency in susceptible hosts. However, the concentration and timing of VSR expression are critical, as overly potent or prolonged suppression can lead to severe viral symptoms and plant toxicity, confounding phenotypic analysis [4]. The goal is to achieve a transient and balanced suppression that boosts siRNA accumulation and spread without compromising plant health.
Table 2: Viral Suppressors of RNAi (VSRs) and Their Applications in VIGS
| VSR | Viral Origin | Mode of Action | Effect on VIGS | Considerations |
|---|---|---|---|---|
| P19 | Tombusviruses | Binds and sequesters siRNA duplexes. | Potently enhances siRNA accumulation and silencing intensity. | Can cause severe viral symptoms; use requires optimization [4]. |
| HC-Pro | Potyviruses | Inhibits siRNA amplification (RDR activity). | Enhances initial viral replication and spread. | Broad-spectrum suppressor; can alter miRNA pathways [4]. |
| 2b / C2b | Cucumoviruses | Binds to and inhibits Argonaute (AGO) proteins. | Blocks the effector step of silencing, allowing viral accumulation. | Efficacy is highly host-species dependent [4]. |
Experimental Protocols for Assessing Systemic Spreading
To evaluate the success of vector engineering and VSR strategies, robust experimental protocols are required. Below is a detailed methodology for establishing a VIGS system and quantifying systemic silencing.
Protocol: Agrobacterium-Mediated VIGS with Vacuum Infiltration
This protocol, adapted from a 2024 study on sunflowers, provides a highly efficient method for difficult-to-transform species and can be optimized for others [8].
I. Vector Construction and Agrobacterium Preparation
- Clone Target Fragment: Amplify a ~200 bp fragment of the target gene (e.g., Phytoene Desaturase [PDS] for a visible photo-bleaching phenotype). Use bioinformatic tools (e.g., pssRNAit) to select a fragment with high predicted siRNA density [8].
- Gateway Cloning: Clone the fragment into a VIGS vector (e.g., pTRV2) using restriction enzymes (e.g., XbaI/BamHI) or Gateway recombination.
- Transform Agrobacterium: Introduce the recombinant pTRV2 and the helper pTRV1 plasmids into Agrobacterium tumefaciens strain GV3101 via electroporation.
- Prepare Agrobacterium Culture:
- Streak transformed Agrobacterium from glycerol stock onto LB agar plates with appropriate antibiotics (e.g., kanamycin, rifampicin). Incubate at 28°C for 48 hours.
- Pick a single colony and inoculate a 5 mL liquid LB culture with antibiotics. Shake overnight at 28°C.
- Pellet cells and resuspend in infiltration medium (10 mM MgCl₂, 10 mM MES, 200 µM acetosyringone). Adjust the optical density at 600 nm (OD₆₀₀) to 0.8-1.0.
- Incubate the suspension at room temperature for 3-6 hours before infiltration.
II. Plant Material Preparation and Vacuum Infiltration
- Plant Material: Use sunflower seeds with seed coats partially removed to improve infiltration [8].
- Mixing Agrobacterial Suspensions: Combine the pTRV1 and pTRV2-target suspensions in a 1:1 ratio.
- Vacuum Infiltration:
- Submerge the seeds in the combined Agrobacterium suspension in a beaker.
- Place the beaker in a vacuum desiccator. Apply a vacuum of 80-85 kPa for 3 minutes.
- Slowly release the vacuum to ensure the suspension penetrates the seeds.
- Co-cultivation: Transfer the infiltrated seeds to a moist paper towel or potting soil and incubate in the dark at room temperature for ~6 hours.
- Plant Growth: Sow the seeds directly into soil and grow under standard greenhouse conditions (e.g., 22°C, 18-h light/6-h dark photoperiod).
III. Analysis of Silencing Efficiency
- Phenotypic Monitoring: Observe plants for the development of systemic silencing symptoms (e.g., photo-bleaching for PDS), typically appearing 2-4 weeks post-infiltration. Document the spread and intensity of the phenotype over time using time-lapse photography [8].
- Molecular Confirmation:
- RT-qPCR: Quantify the mRNA levels of the target gene in silenced (e.g., bleached) and non-silenced (green) tissues. Successful silencing should show a >70% reduction in target mRNA [8].
- siRNA Detection: Perform northern blotting or high-throughput sequencing to detect the accumulation of siRNAs specific to the target gene in systemic tissues.
- Viral Spread Tracking: Use RT-PCR with primers specific to the viral vector (e.g., TRV) to confirm the presence of the virus in different plant parts, including upper, non-infiltrated leaves [8].
Protocol: Co-expression of VSRs to Enhance Silencing
This protocol supplements the standard VIGS procedure to boost silencing efficiency using the P19 suppressor.
- Prepare VSR Strain: Transform the P19 expression vector into Agrobacterium strain GV3101.
- Culture Preparation: Grow separate cultures for pTRV1, pTRV2-target, and P19 as described in Section 5.1.
- Mixed Infiltration: Combine the three Agrobacterium suspensions in a final ratio of 1:1:0.5 (TRV1:TRV2-target:P19). The lower concentration of P19 helps avoid excessive toxicity.
- Infiltrate Plants: Use syringe infiltration or vacuum infiltration as appropriate for the plant species.
- Monitor and Analyze: Closely monitor plants for accelerated or intensified silencing phenotypes and potential toxic effects. Analyze siRNA and target mRNA levels as described above to quantitatively assess the enhancement provided by P19.
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Reagents for VIGS Vector Engineering and Silencing Enhancement
| Reagent / Tool | Function / Description | Example Use Case |
|---|---|---|
| TRV-based VIGS Vectors (pTRV1/pTRV2) | Standard bipartite vector system for inducing gene silencing in a wide host range. | Functional analysis of disease resistance genes in pepper (Capsicum annuum) [4]. |
| Agrobacterium tumefaciens GV3101 | Standard disarmed strain for delivering T-DNA containing viral vectors into plant cells. | Used in vacuum infiltration and syringe infiltration protocols for VIGS [8]. |
| VSR Expression Plasmids (e.g., P19) | Plasmid vectors for expressing viral suppressor proteins to enhance silencing. | Co-infiltration with TRV vectors to boost siRNA accumulation in N. benthamiana [4]. |
| pssRNAit / siRNA Design Tools | Bioinformatics software for predicting effective siRNA target sites within a gene. | Selecting a 200 bp fragment with high siRNA density for cloning into pTRV2 [8]. |
| Minimal syn-tasiRNA Precursors | Short (54 nt) synthetic DNA sequences engineered to produce highly specific artificial siRNAs. | Cloning into PVX vectors for high-specificity, transgene-free silencing via spray application [24]. |
| Acetosyringone | A phenolic compound that induces the Agrobacterium Vir genes, essential for T-DNA transfer. | Added to the Agrobacterium infiltration medium to maximize transformation efficiency. |
Genotype-Dependent Responses and Species-Specific Adaptation
Virus-Induced Gene Silencing (VIGS) has emerged as a powerful reverse genetics tool for characterizing gene function across diverse plant species. This technology leverages the plant's endogenous RNA interference machinery, wherein virus-derived small interfering RNAs guide sequence-specific silencing of target genes. However, the efficacy of VIGS is profoundly influenced by genotype-dependent responses and species-specific adaptations, presenting significant challenges for its broad application. This technical review examines the molecular basis of these variations, detailing how host genetics, viral vector selection, and environmental factors interact to determine silencing efficiency. We present comprehensive experimental data quantifying these effects across species, provide optimized protocols for challenging systems, and outline a standardized framework for evaluating genotype-specific VIGS responsiveness. Understanding these determinants is crucial for advancing functional genomics in non-model species and harnessing VIGS for crop improvement.
Virus-Induced Gene Silencing is a sophisticated plant defense mechanism that has been co-opted as a powerful functional genomics tool. At its core, VIGS operates through the post-transcriptional gene silencing pathway, an evolutionarily conserved RNA-based immune response that recognizes and degrades viral RNA [4] [5]. When recombinant viral vectors carrying host-derived sequences infect plants, the replication process generates double-stranded RNA intermediates that are recognized by the host's enzymatic machinery.
The central effector molecules in this process are small interfering RNAs, which are generated through the cleavage of double-stranded RNA by Dicer-like enzymes [5]. These 21-24 nucleotide siRNAs are then incorporated into the RNA-induced silencing complex, where they serve as guides for sequence-specific identification and cleavage of complementary endogenous mRNA targets [4] [5]. This targeted degradation leads to suppression of gene expression and produces observable phenotypic changes that enable functional characterization.
The efficiency of this siRNA-mediated silencing cascade varies considerably across plant genotypes and species, influenced by multiple factors including the host's RNAi machinery components, viral movement patterns, and environmental conditions. This review systematically analyzes these determinants of VIGS efficiency, with particular emphasis on their implications for research and crop development.
Molecular Mechanisms of siRNA Biogenesis and Action in VIGS
The molecular pathway of VIGS represents a sophisticated interplay between viral components and the plant's RNAi machinery. The process begins when a recombinant viral vector delivering a fragment of a plant gene of interest is introduced into plant tissues, typically via Agrobacterium-mediated transformation (agroinfiltration) or direct inoculation. Following viral replication and movement, the key steps of silencing are initiated [4] [5].
Figure 1. siRNA-Mediated Gene Silencing Pathway in VIGS. This diagram illustrates the core molecular mechanism of Virus-Induced Gene Silencing, from viral infection to systemic silencing. The process involves viral replication generating double-stranded RNA, which is processed into siRNAs that guide the RNA-induced silencing complex to target complementary mRNAs for degradation.
The initial trigger for silencing is the formation of double-stranded RNA molecules during viral replication. These dsRNA structures are recognized by host Dicer-like enzymes, which process them into small interfering RNAs of characteristic lengths (21-24 nucleotides) [5]. This step is crucial for determining the specificity and efficiency of silencing, as the siRNAs generated will guide the silencing machinery to complementary sequences.
The siRNA duplexes are then loaded into the RNA-induced silencing complex, with the Argonaute protein serving as the catalytic component. The RISC complex uses the siRNA as a guide to identify and cleave complementary mRNA targets, resulting in post-transcriptional gene silencing [5]. Importantly, the silencing signal amplifies and spreads systemically through the plant, mediated by the movement of siRNAs between cells and through the vasculature, ultimately leading to observable phenotypic changes in tissues distant from the initial infection site [4].
Genotype-Dependent Efficiency of VIGS
Empirical Evidence of Genotype-Dependent Responses
The efficiency of VIGS varies significantly across plant genotypes, influenced by genetic factors that affect viral susceptibility, siRNA biogenesis, and systemic signaling. Quantitative studies across multiple species have demonstrated this genotype-dependent responsiveness.
In sunflower, a comprehensive evaluation of six different genotypes revealed substantial variation in VIGS efficiency using the same TRV-based vector and inoculation protocol. The genotype 'Smart SM-64B' showed the highest infection rate (91%), while other commercial cultivars exhibited lower susceptibility, with infection percentages ranging from 62% to 91% [8]. Notably, the extent of silencing phenotype spread also varied independently of infection rates, with 'Smart SM-64B' showing more limited spreading of photo-bleached areas compared to other genotypes despite its high infection percentage [8].
Similar genotype-dependent responses have been documented in other crops. In soybean, cassava, citrus, and wheat, researchers have observed significant variation in VIGS efficiency across different cultivars [8]. These findings highlight the critical importance of genotype selection and optimization when establishing VIGS protocols for new species or varieties.
Table 1: Genotype-Dependent VIGS Efficiency in Sunflower
| Genotype | Infection Percentage | Silencing Phenotype Spread | Key Observations |
|---|---|---|---|
| Smart SM-64B | 91% | Limited | High infection but restricted phenotype spread |
| ZS | 77% | Moderate | Balanced efficiency and spread |
| Buzuluk | 68% | Extensive | Good systemic movement of silencing signal |
| Kubanski Semechki | 62% | Moderate | Lower infection rate but reasonable spread |
| Lakomka | 65% | Extensive | Good phenotype manifestation despite moderate infection |
| Shelkunshik | 70% | Moderate | Consistent silencing across infected plants |
Molecular Determinants of Genotypic Variation
The genetic factors underlying these genotype-dependent responses operate at multiple levels within the VIGS pathway. Key determinants include:
Viral replication efficiency: Natural variation in genes involved in viral recognition and defense can significantly impact the establishment and spread of the VIGS vector [4].
RNAi machinery components: Polymorphisms in genes encoding Dicer-like proteins, Argonaute proteins, and RNA-dependent RNA polymerases can alter siRNA biogenesis, amplification, and function [4] [5].
Systemic silencing signal movement: The capacity for cell-to-cell and long-distance movement of silencing signals varies between genotypes, affecting the extent and uniformity of silencing phenotypes [4] [8].
Viral suppressor of RNA silencing interactions: Some viruses encode proteins that suppress the RNAi response, and the efficacy of these suppressors can vary across plant genotypes, indirectly affecting VIGS efficiency [4].
Understanding these molecular determinants enables researchers to select appropriate genotypes for VIGS studies or implement strategies to overcome genotypic limitations, such as using viral suppressors of RNA silencing to enhance silencing efficiency in recalcitrant genotypes [4].
Species-Specific Adaptations and Vector Selection
Viral Vector Selection for Different Plant Species
The choice of viral vector is a critical determinant of VIGS success and must be tailored to the specific host species. Different viral vectors exhibit distinct host ranges, tissue tropism, and movement patterns that significantly impact silencing efficiency.
Table 2: Viral Vectors for VIGS Across Plant Species
| Viral Vector | Virus Type | Host Range | Key Features | Optimal Species |
|---|---|---|---|---|
| Tobacco Rattle Virus | RNA virus | Broad | Efficient systemic movement; mild symptoms | Solanaceae (pepper, tomato, tobacco) [4] |
| Barley Stripe Mosaic Virus | RNA virus | Moderate | Effective in monocots | Barley, wheat [64] |
| Cotton Leaf Crumple Virus | DNA virus | Limited | Geminivirus-based; good for meristematic tissues | Cotton, legumes [4] |
| Cucumber Mosaic Virus | RNA virus | Broad | Rapid systemic infection; can induce severe symptoms | Various dicots [4] |
| Apple Latent Spherical Virus | RNA virus | Very broad | Mild symptoms; wide host range | Diverse species including legumes |
The TRV-based vector system has emerged as one of the most versatile and widely used systems for VIGS, particularly for plants in the Solanaceae family [4]. Its bipartite genome organization requires two vectors: TRV1, encoding replicase and movement proteins, and TRV2, containing the coat protein gene and a multiple cloning site for inserting target sequences [4]. This system combines efficient systemic movement with relatively mild viral symptoms, facilitating phenotypic analysis.
For monocot species, BSMV-based vectors have proven effective, enabling functional genomics studies in cereal crops like barley and wheat [64]. Geminivirus-based vectors (e.g., CLCrV, ACMV) offer advantages for certain applications, including potential access to meristematic tissues and different patterns of systemic movement [4].
Case Studies: Species-Specific Protocol Optimization
Sunflower VIGS Optimization
Sunflower presents particular challenges for VIGS due to its recalcitrance to transformation and variable genotype responses. Recent methodological advances have identified optimal parameters for efficient VIGS in this species:
Delivery method: Seed vacuum infiltration followed by 6 hours of co-cultivation proved most effective, achieving up to 77% infection rate and strong silencing of the target HaPDS gene (normalized relative expression = 0.01) [8].
Developmental stage: Silencing establishment in younger tissues showed more active spreading of photo-bleached spots compared to mature tissues [8].
Viral movement analysis: TRV was detected in leaves at the highest node (up to node 9) in infected sunflower plants, demonstrating extensive viral spreading throughout the plant using the optimized protocol [8].
This protocol successfully circumvented the need for in vitro recovery or surface sterilization steps that had limited previous approaches, significantly streamlining the VIGS procedure for sunflower [8].
Barley VIGS for Cell Wall Biosynthesis Genes
In barley, BSMV-based VIGS has been successfully employed to study genes involved in cell wall biosynthesis, including cellulose synthase (CesA) genes. This approach revealed coordinated regulation of multiple cell wall biosynthetic genes, demonstrating the power of VIGS for analyzing metabolic pathways [64].
Quantitative real-time RT-PCR protocols were optimized for accurate assessment of silencing efficiency in barley, incorporating appropriate reference genes (EF-1α, actin, ubiquitin) to normalize target gene transcript levels [65]. This methodological refinement was essential for detecting the subtle but biologically significant changes in gene expression resulting from VIGS.
Experimental Protocols for Assessing VIGS Efficiency
Quantitative Assessment of Silencing Efficiency
Accurate evaluation of VIGS efficiency requires robust molecular methods to quantify target gene repression. Real-time reverse transcription PCR has emerged as the gold standard for this application due to its sensitivity, accuracy, and broad dynamic range [65].
Protocol: Real-Time RT-PCR for VIGS Efficiency quantification
RNA Isolation and Quality Control:
- Extract high-quality total RNA from silenced and control tissues using standard methods (e.g., TRIzol or column-based kits).
- Assess RNA integrity by gel electrophoresis or microfluidic analysis.
- Treat samples with DNase I to remove contaminating genomic DNA.
- Verify DNA removal by performing real-time PCR "no cDNA control" reactions; samples with Ct values >32 require re-treatment [65].
Reference Gene Selection and Validation:
- Test multiple candidate reference genes for stability under experimental conditions.
- Common reference genes include elongation factor 1α (EF-1), actin, and ubiquitin [65].
- In tomato and N. benthamiana, EF-1α and ubiquitin have shown superior stability compared to actin during VIGS experiments [65].
- Normalize target gene expression to stable reference genes using the 2^(-ΔΔCt) method [65].
Primer Design and Validation:
- Design primers that efficiently amplify specific products without primer-dimers or non-specific amplification.
- Validate primer efficiency using standard curves with serial dilutions of cDNA.
- Consider designing primers targeting different regions of the transcript to assess potential asymmetric silencing effects [65].
Data Interpretation:
- Compare target gene expression in VIGS-treated plants to empty vector controls.
- Include biological and technical replicates to enable statistical analysis (typically 3-6 biological replicates per treatment).
- Correlative molecular and phenotypic analyses strengthen functional interpretations.
Standardized Framework for Evaluating Genotype-Specific VIGS Responsiveness
To systematically evaluate VIGS efficiency across genotypes, we propose the following standardized framework:
Phenotypic Scoring System:
- Develop a quantitative scale for silencing-associated phenotypes (e.g., 0-5 for photo-bleaching intensity and distribution).
- Document the timing of phenotype appearance and persistence.
- Assess phenotype stability under different environmental conditions.
Molecular Confirmation:
- Quantify target gene expression reduction using real-time RT-PCR.
- Detect virus-derived siRNAs by northern blotting or small RNA sequencing.
- Verify systemic spread of silencing signal through grafting experiments or local vs. distal tissue analysis.
Environmental Optimization:
- Test silencing efficiency under different temperature regimes (typically 18-25°C).
- Evaluate photoperiod effects (often 16h light/8h dark cycles optimal).
- Assess humidity influences on agroinfiltration efficiency and viral spread.
The Scientist's Toolkit: Essential Research Reagents
Table 3: Essential Research Reagents for VIGS Experiments
| Reagent/Category | Specific Examples | Function and Application |
|---|---|---|
| Viral Vectors | TRV (pYL192/TRV1, pYL156/TRV2), BSMV, CLCrV | Delivery of target gene fragments to trigger silencing; selection depends on host species [4] [8] |
| Agrobacterium Strains | GV3101, AGL1 | Delivery of viral vectors into plant cells via agroinfiltration [8] |
| Selection Antibiotics | Kanamycin, Gentamicin, Rifampicin | Selection of recombinant Agrobacterium strains carrying VIGS constructs [8] |
| Reference Genes | EF-1α, Ubiquitin, Actin | Normalization of target gene expression in real-time RT-PCR analysis [65] |
| Visual Marker Genes | Phytoene Desaturase (PDS), ChlH | Positive control for VIGS through visible photo-bleaching phenotype [4] [8] |
| Infiltration Buffers | Acetosyringone-containing buffers | Enhancement of Agrobacterium virulence gene induction for improved transformation efficiency |
| RNA Isolation & qPCR Kits | DNase treatment reagents, SYBR Green master mixes | Accurate quantification of silencing efficiency through real-time RT-PCR [65] |
The successful application of Virus-Induced Gene Silencing as a functional genomics tool requires careful consideration of genotype-dependent responses and species-specific adaptations. The efficiency of siRNA-mediated silencing is governed by complex interactions between host genetic factors, viral vector properties, and environmental conditions. The experimental frameworks and optimized protocols presented here provide a roadmap for addressing these challenges across diverse plant systems.
As VIGS technology continues to evolve, integrating multi-omics approaches with high-throughput silencing screens will further illuminate the genetic networks underlying these variable responses. This knowledge will expand the utility of VIGS for functional gene characterization in non-model species and accelerate crop improvement through rapid identification of genes controlling agronomically valuable traits. The ongoing refinement of species-specific protocols and systematic evaluation of genotype-dependent factors will ensure that VIGS remains at the forefront of plant functional genomics.
Virus-Induced Gene Silencing (VIGS) has emerged as a powerful reverse genetics tool for analyzing gene function in plants by exploiting the endogenous RNA interference (RNAi) machinery. This process is mediated by small interfering RNAs (siRNAs), which guide sequence-specific degradation of complementary target mRNAs through the RNA-induced silencing complex (RISC) [5] [48]. The efficiency of VIGS is not merely a function of vector design or inoculation technique but is profoundly influenced by environmental conditions that modulate plant physiology, viral replication, and the systemic spread of silencing signals [4] [8]. This technical guide examines the critical role of temperature, humidity, and photoperiod in optimizing VIGS efficacy, providing researchers with evidence-based protocols to maximize silencing efficiency in functional genomics studies, particularly within the context of siRNA biogenesis and movement.
The siRNA Mechanistic Foundation of VIGS
The efficacy of VIGS is fundamentally rooted in the complex lifecycle of siRNA-mediated silencing. Understanding this mechanism is prerequisite to appreciating how environmental parameters can be leveraged to enhance silencing efficiency.
Figure 1. The siRNA Pathway in Virus-Induced Gene Silencing. This diagram outlines the core mechanistic steps from viral double-stranded RNA (dsRNA) generation to target mRNA degradation, highlighting key amplification and systemic movement steps.
The process initiates when recombinant viral vectors introduce target gene fragments into plant cells. During viral replication, double-stranded RNA (dsRNA) intermediates are formed, which are recognized as foreign molecules by the plant's innate antiviral defense system [5]. Dicer-like (DCL) enzymes then process these long dsRNAs into 21–24 nucleotide small interfering RNA (siRNA) duplexes [5] [48]. These siRNAs are loaded into an Argonaute (AGO) protein within the RNA-induced silencing complex (RISC), which uses the siRNA as a guide for sequence-specific recognition and endonucleolytic cleavage of complementary endogenous mRNA targets, leading to post-transcriptional gene silencing (PTGS) [5] [48]. A critical feature for the observed phenotypic changes is the systemic spread of the silencing signal. This signal, which is likely a combination of siRNAs and possibly amplified secondary siRNAs produced by host RNA-directed RNA polymerases (RDRPs), moves through the phloem to distal tissues, establishing silencing throughout the plant [5] [48]. The efficiency of every step in this pathway—from DCL activity and RISC assembly to the phloem-mediated mobility of siRNAs—is susceptible to modulation by environmental conditions.
Quantitative Effects of Environmental Parameters on VIGS Efficiency
Environmental factors influence physiological processes such as plant defense responses, viral replication rates, and vascular transport, thereby directly impacting the establishment and spread of gene silencing. The table below summarizes key experimental data and optimal ranges for major environmental parameters.
Table 1. Optimization of Environmental Parameters for VIGS Efficiency
| Environmental Factor | Optimal Range | Experimental Findings & Impact on Silencing | Plant Species Studied |
|---|---|---|---|
| Temperature | 16–20 °C | 20°C/18°C (day/night) induced stronger silencing than 23°C/18°C or 26°C/18°C; lower temperatures (16-18°C) promote silencing efficiency and reduce viral symptom severity. [66] | Petunia [66] |
| Lower temperatures promote silencing efficiency and reduce viral symptom severity. [66] | Potato [66] | ||
| Photoperiod | 16–18 hours light / 6–8 hours dark | An 18-h light/6-h dark photoperiod was used in an optimized VIGS protocol, supporting efficient viral spread and silencing manifestation. [8] | Sunflower [8] |
| Humidity | Moderate to High (~45-69%) | An average relative humidity of ~69% was maintained in growth chambers for effective VIGS. Lower humidity (~45%) has also been successfully used, but very low humidity can stress plants and impair silencing. [66] [8] | Petunia [66] |
Experimental Protocols for Environmental Optimization
Protocol 1: Temperature Optimization Assay
This protocol is designed to systematically evaluate the effect of temperature on VIGS efficacy, using Phytoene Desaturase (PDS) as a visual reporter gene.
- Plant Material & Growth: Select a uniform set of plant seeds (e.g., Nicotiana benthamiana, petunia, or tomato). Sow seeds in a standardized soil mix and grow them under controlled, identical conditions until they reach the 3-4 leaf stage, which is ideal for inoculation [66].
- VIGS Inoculation: Inoculate all plants using a consistent method (e.g., agroinfiltration, root wounding-immersion [67]). Use a TRV2-PDS construct to induce photobleaching.
- Temperature Treatments: Post-inoculation, randomly divide plants into different growth chambers with the following day/night temperature regimes:
- Treatment A (Low): 16°C / 14°C
- Treatment B (Optimal): 20°C / 18°C
- Treatment C (Sub-optimal): 23°C / 20°C
- Treatment D (High): 26°C / 22°C Maintain all other environmental factors (photoperiod, light intensity, humidity) constant across chambers.
- Data Collection and Analysis:
- Phenotypic Scoring: Record the onset and progression of photobleaching in newly emerged leaves daily for 3-4 weeks post-inoculation. Calculate the percentage of plants showing silencing and the area of leaf tissue silenced.
- Molecular Validation: At the peak of silencing (e.g., 14-21 days post-inoculation), harvest silenced and non-silenced leaf tissue from each treatment. Quantify the knockdown of PDS mRNA using RT-qPCR with species-specific primers. Additionally, detect the presence and level of the TRV virus using PCR or western blot to correlate environmental conditions with viral accumulation [66].
Protocol 2: Integrated Environment Setup for High-Throughput VIGS
This protocol establishes a standardized environment for robust, high-efficiency VIGS suitable for large-scale functional genomics screens.
- Pre-Inoculation Environment: Grow plants to the desired inoculation stage (3-5 weeks after sowing) under a long-day photoperiod (16-h light / 8-h dark) using metal halide or LED lights providing approximately 150 μmol m⁻² s⁻¹ photosynthetic photon flux density (PPFD) [66]. Maintain temperature at 22-25°C and relative humidity at ~60% for vigorous growth.
- Post-Inoculation Environment: Immediately after agroinoculation, transfer plants to a growth chamber with a lower temperature regime. Based on empirical data, a day/night setting of 20°C/18°C is highly effective for promoting silencing efficiency in species like petunia [66].
- Humidity and Light Control: Maintain moderate relative humidity (65-70%) to prevent excessive water stress while minimizing fungal growth risks [66]. Continue the 16-h light / 8-h dark photoperiod to support plant health and metabolic activity necessary for systemic silencing signal propagation.
- Monitoring and Maintenance: Monitor plants daily for the development of viral symptoms or silencing phenotypes. Fertilize as needed with a balanced nutrient solution (e.g., 250 mg L⁻¹ N) to avoid nutrient stress that can confound phenotypic analysis [66].
The Scientist's Toolkit: Essential Research Reagents
Table 2. Key Reagent Solutions for VIGS Optimization
| Reagent / Tool | Function in VIGS Protocol | Specification / Example |
|---|---|---|
| TRV Vectors (pTRV1, pTRV2) | Standard bipartite viral vector system for inducing silencing; TRV2 carries the target gene fragment. | pYL192 (TRV1), pYL156 (TRV2) [8] |
| Control Insert Vectors | Negative control to distinguish viral symptoms from silencing phenotypes; contains a non-plant insert. | pTRV2-sGFP (fragment of GFP gene) [66] |
| Visual Reporter Constructs | Allows visual tracking of silencing efficiency and spread through pigment loss. | TRV2-PDS (photobleaching), TRV2-CHS (white patches on petals) [66] |
| Engineered Silencing Suppressors | Enhances VIGS efficacy by promoting systemic spread of the virus while minimizing local suppression. | TRV2-C2bN43 (truncated CMV 2b protein) [48] |
| Syn-tasiRNA Minimal Precursors | Provides high-specificity silencing with minimal off-target effects; expressible from viral vectors. | 54-nt precursor with miRNA target site and syn-tasiRNA sequence [24] |
| Agroinfiltration Medium | Preparation of Agrobacterium suspension for plant inoculation. | 10 mM MgCl₂, 10 mM MES (pH 5.6), 150 μM acetosyringone [67] |
The precise modulation of temperature, photoperiod, and humidity is not merely a matter of improving plant health but is a fundamental strategy for maximizing the efficacy of siRNA-mediated gene silencing in VIGS. By understanding the mechanistic basis of how these conditions influence DCL activity, RISC complex stability, and the systemic mobility of siRNAs, researchers can transform VIGS from a potentially variable technique into a robust, high-throughput platform for gene functional analysis. The protocols and data presented herein provide a concrete foundation for standardizing and optimizing VIGS conditions across plant species, thereby accelerating discovery in functional genomics and crop biotechnology.
Virus-Induced Gene Silencing (VIGS) represents a powerful reverse genetics tool that leverages the plant's innate RNA interference (RNAi) machinery to study gene function. The foundation of this technology rests on the pivotal role of small interfering RNAs (siRNAs), which are processed from double-stranded RNA precursors and guide the sequence-specific degradation of complementary messenger RNA (mRNA) targets [4] [5]. The core mechanism involves the incorporation of siRNAs into the RNA-induced silencing complex (RISC), which subsequently cleaves or translationally represses the target mRNA, leading to post-transcriptional gene silencing (PTGS) [11] [68]. This siRNA-mediated silencing pathway forms the biological basis for VIGS, wherein recombinant viral vectors are engineered to carry fragments of host genes, triggering the production of siRNAs that direct the silencing of the corresponding endogenous plant genes [4].
The efficacy of VIGS is critically dependent on the efficient delivery of these viral vectors into plant cells. Agrobacterium tumefaciens, a natural genetic engineer, has emerged as the most prevalent delivery vehicle, facilitating the transfer of T-DNA containing the viral genome into the plant nucleus [4] [69]. Among the various inoculation methods, vacuum infiltration and co-cultivation have proven to be particularly effective, especially for challenging plant species and tissues. This technical guide details the refinement of these two key delivery protocols, framing them within the essential context of siRNA function in VIGS research, and provides actionable methodologies for researchers aiming to implement these techniques in their functional genomics workflows.
Core Principles of siRNA in VIGS Workflows
The application of VIGS for functional genomics is intrinsically linked to the biogenesis and activity of siRNAs. The entire process, from vector delivery to observable phenotypic change, is orchestrated by these small RNA molecules. Figure 1 below illustrates the integrated experimental workflow of a VIGS experiment, highlighting the central role of siRNA-mediated silencing and the points at which vacuum infiltration and co-cultivation are applied.
Figure 1. Integrated VIGS Workflow with Key Delivery Techniques. This diagram outlines the complete sequence of steps in a VIGS experiment, from vector construction to phenotypic analysis. The critical delivery step (4), where vacuum infiltration and co-cultivation are applied, is highlighted, leading to the core siRNA-mediated silencing steps (6-7).
The molecular machinery triggered after successful vector delivery is summarized in Table 1, which details the key reagents and their functions in the siRNA/VIGS process.
Table 1: Research Reagent Solutions for siRNA/VIGS Studies
| Reagent / Component | Function in siRNA/VIGS Workflow |
|---|---|
| Viral Vector (e.g., TRV, BBWV2) | Delivers the target plant gene fragment into the host; the viral RNA serves as a template for the formation of dsRNA, the precursor to siRNAs [4]. |
| Agrobacterium tumefaciens | A biological vehicle that efficiently transfers T-DNA containing the viral vector into the plant cell nucleus [69] [47]. |
| siRNA / shRNA | The effector molecules; synthetic siRNAs or vector-derived shRNAs are processed into functional siRNAs that guide the RISC complex to the target mRNA [11] [70]. |
| Dicer-like (DCL) Enzyme | An endonuclease that cleaves long dsRNA (of viral origin) into 21-24 nucleotide siRNA duplexes [4] [5]. |
| RNA-induced Silencing Complex (RISC) | The core effector complex, containing Argonaute (AGO) protein, which uses the siRNA as a guide to identify and cleave complementary mRNA targets [4] [11]. |
| Acetosyringone | A phenolic compound that induces the virulence genes of Agrobacterium, enhancing the efficiency of T-DNA transfer [69] [47]. |
| Infiltration Buffer Additives (Cysteine, Tween 20) | Reduces oxidative stress and enhances surfactant activity, respectively, improving the survival of plant tissues and the penetration of the agrobacterial suspension during infiltration [69]. |
Vacuum Infiltration Protocol
Vacuum infiltration is a physical method that forces the Agrobacterium suspension into the intercellular spaces of plant tissues by applying a negative pressure environment. This technique is highly effective for whole-plant level silencing, especially when applied to germinated seeds or young seedlings, enabling gene function studies during early developmental stages [69].
Step-by-Step Methodology
- Agrobacterial Preparation: Inoculate Agrobacterium strains containing the necessary VIGS vectors (e.g., pTRV1 and pTRV2-target) in appropriate antibiotic-containing media. Grow the cultures to the mid-log phase (OD₆₀₀ ≈ 1.0) [47]. Centrifuge the bacterial cells and resuspend them in an infiltration buffer. A proven, effective buffer consists of 10 mM MES, 10 mM MgCl₂, and 150 μM acetosyringone, with critical additives being 2 mM L-cysteine and 0.01% Tween 20 to reduce tissue browning and improve wettability, respectively [69].
- Plant Material Preparation: For seed infiltration, surface-sterilize seeds and allow them to germinate until the radicle just emerges. This stage is critical for successful uptake [69]. For seedling infiltration, use young, healthy seedlings.
- Vacuum Application: Submerge the plant material (germinated seeds or seedlings) in the Agrobacterium suspension inside a sealed container. Apply a vacuum of 0.5 to 1.0 bar (approximately 15-20 inches of Hg) for a specific duration. Research in wheat and maize has shown that multiple short treatments are more effective than a single long one; a protocol of six negative-pressure applications of 30 seconds each has been successfully used [69].
- Co-cultivation and Recovery: After infiltration, briefly blot the plant material dry and transfer it to co-cultivation media (solid medium for seeds/seedlings) or soil (for older plants). Maintain the plants in the dark at room temperature for 2-3 days to allow for T-DNA transfer and the initiation of infection.
- Plant Growth and Phenotyping: Transfer the plants to standard growth conditions. Gene silencing phenotypes, such as photo-bleaching from PDS gene silencing, typically become visible within 2-4 weeks post-infiltration, depending on the species and target gene [69].
Key Optimization Parameters
The efficiency of vacuum infiltration is highly dependent on several physical and biological parameters. Table 2 synthesizes quantitative data from optimization studies to guide researchers.
Table 2: Key Optimization Parameters for Vacuum Infiltration
| Parameter | Optimal Range | Impact on Efficiency | Citation |
|---|---|---|---|
| Bacterial Density (OD₆₀₀) | 0.8 - 1.2 | Critical; OD~600~ = 1.0 identified as a robust optimum for embryo-derived seedlings, balancing delivery and plant cell stress. | [47] |
| Acetosyringone Concentration | 150 - 200 μM | Essential for inducing Agrobacterium virulence genes; 200 μM significantly enhances transformation efficiency. | [69] [47] |
| Vacuum Duration & Cycles | 5-6 cycles of 30 sec | Multiple short cycles are more effective than a single prolonged application for achieving whole-plant silencing in monocots. | [69] |
| Plant Developmental Stage | Germinated seeds | Highest efficiency achieved at 35 days after germination for peony seedlings; germinated seeds allow silencing from the earliest developmental stages. | [69] [47] |
| Infiltration Solution Additives | 2 mM Cysteine, 0.01% Tween 20 | Markedly improves plant tissue survival and agrobacterial penetration, leading to more consistent silencing. | [69] |
Co-cultivation Techniques
Co-cultivation is a biological delivery method that involves bringing plant tissues and Agrobacterium into intimate contact for a period sufficient for T-DNA transfer. Unlike vacuum infiltration, it relies on the natural tropism and virulence mechanism of the bacterium.
Step-by-Step Methodology
- Preparation of Explants and Agrobacterium: Young, metabolically active tissues such as leaf discs, germinated seeds, or in vitro seedling segments are ideal explants. Prepare the Agrobacterium suspension as for vacuum infiltration, resuspending to an OD₆₀₀ of 0.5 - 1.0 in a buffer containing acetosyringone [47].
- Inoculation: Immerse the explants in the Agrobacterium suspension for 15 minutes to 2 hours with gentle agitation. The study on Paeonia ostii identified a 2-hour infection duration as optimal when combined with other parameters [47].
- Co-cultivation Phase: After inoculation, blot the explants to remove excess bacteria and transfer them to solid co-cultivation media. This media is typically a standard plant tissue culture medium (e.g., MS or WPM) supplemented with 200 μM acetosyringone to maintain Agrobacterium virulence. Co-cultivate in the dark at 22-24°C for 2-3 days [47].
- Decontamination and Recovery: Following co-cultivation, wash the explants thoroughly with sterile water, often containing antibiotics like carbenicillin or timentin to eliminate the Agrobacterium. Transfer the explants to fresh regeneration or rooting media to recover and allow for the development of silenced plants.
Key Optimization Parameters
Table 3 outlines the critical factors for optimizing co-cultivation-based VIGS delivery.
Table 3: Key Optimization Parameters for Co-cultivation
| Parameter | Optimal Range | Impact on Efficiency | Citation |
|---|---|---|---|
| Co-cultivation Duration | 2 - 3 days | Allows sufficient time for T-DNA transfer without bacterial overgrowth that can kill the explant. | [47] |
| Acetosyringone in Co-cultivation Media | 200 μM | Maintains the induction of virulence genes during the T-DNA transfer process, crucial for high efficiency. | [47] |
| Bacterial Strain & Density | OD₆₀₀ = 0.5 - 1.0 | Lower densities than vacuum infiltration can be used; prevents overgrowth and "hypersensitive" plant responses. | [47] |
| Explant Type & Physiology | Germinated seeds, in vitro seedlings | Juvenile tissues with active cell division are more competent for T-DNA integration and/or transient expression. | [69] [47] |
| Temperature | 22-24°C | A slightly lower temperature than standard culture conditions can improve T-DNA transfer and explant viability. | [47] |
Technical Comparison and Selection Guide
Choosing between vacuum infiltration and co-cultivation depends on the research objectives, plant species, and available resources. Figure 2 provides a decision-making framework to guide this selection, based on key experimental criteria.
Figure 2. VIGS Delivery Technique Selection Guide. This flowchart assists researchers in selecting the most appropriate delivery method (Vacuum Infiltration or Co-cultivation) based on their specific experimental system and goals. The dashed line indicates a common and effective path for seed/seedling silencing.
- Vacuum Infiltration is superior for achieving systemic, whole-plant silencing. It is highly effective for germinated seeds, allowing the study of gene functions in early development and yielding a high proportion of silenced plants [69]. It is also amenable to higher-throughput workflows once the equipment is established. Its main limitation is its variable effectiveness across species, particularly those with dense or waxy leaf structures that are difficult to infiltrate.
- Co-cultivation is a versatile technique that is less dependent on plant architecture and can be applied to a wider range of explants, including those from recalcitrant species. It is the method of choice for stable transformation and is ideal when working with precious or limited plant material, as the conditions are more controlled [47]. Its primary drawbacks are the longer timeline and the more stringent requirement for sterile tissue culture techniques.
The refinement of vacuum infiltration and co-cultivation protocols has significantly advanced the application of siRNA-based VIGS in plant functional genomics. The strategic optimization of parameters such as agrobacterial density, acetosyringone concentration, and the use of protective additives in infiltration buffers directly enhances the efficiency of siRNA delivery and, consequently, the robustness of target gene silencing. As VIGS continues to evolve, integrating with multi-omics technologies and breeding programs, these optimized delivery protocols will remain foundational. They enable researchers to reliably connect genotype to phenotype, accelerating the discovery of gene functions underlying agronomically important traits in crops. By providing a detailed, comparative guide to these techniques, this document equips researchers with the knowledge to implement and further refine these critical methods in their own investigations.
Virus-induced gene silencing (VIGS) represents a foundational RNA interference (RNAi) mechanism that has revolutionized plant functional genomics. This natural antiviral defense pathway utilizes post-transcriptional gene silencing to target invasive viral transcripts, establishing a conserved biological framework that has informed therapeutic siRNA development [5]. The molecular machinery of VIGS—including Dicer-mediated cleavage of double-stranded RNA into 21-24 nucleotide siRNA duplexes and RISC-mediated target mRNA cleavage—provides the fundamental blueprint for modern siRNA therapeutic design [5]. Within this context, enhancing siRNA stability and durability through chemical modifications and advanced formulations has become paramount for translating RNAi from a research tool into a robust therapeutic modality.
The journey from VIGS discovery to approved siRNA therapeutics has spanned decades, with the first siRNA therapeutic (patisiran) receiving regulatory approval in 2018 for hereditary transthyretin-mediated amyloidosis [71]. Since then, additional siRNA drugs have emerged, including givosiran for acute hepatic porphyria, lumasiran for primary hyperoxaluria type 1, and inclisiran for hypercholesterolemia management [71]. These advances established siRNA as a precise therapeutic approach across metabolic, cardiovascular, and genetic disorders, with over 260 siRNA drug candidates currently in preclinical or clinical development [71]. However, unmodified siRNAs face substantial challenges, including rapid nuclease degradation, inefficient cellular uptake, and potential immunogenicity, which have driven innovations in chemical stabilization and delivery technologies [71] [72].
Fundamental Challenges in Unmodified siRNA Applications
Unmodified siRNAs administered without chemical modifications or specialized delivery systems encounter multiple extracellular and intracellular barriers that limit their therapeutic potential. The phosphodiester bonds of native siRNA are highly susceptible to degradation by ubiquitous ribonucleases in biological fluids, drastically reducing their effective half-life [71]. Furthermore, their inherent negative charge and hydrophilic nature prevent efficient crossing of cell membranes, achieving inadequate intracellular concentrations for sustained gene silencing [71].
Without chemical modifications, unmodified siRNAs also carry an increased risk of eliciting innate immune responses through pattern recognition receptors, potentially triggering off-target inflammatory pathways that compromise safety and efficacy [71] [72]. These limitations are particularly relevant in VIGS research, where viral vectors must maintain siRNA integrity long enough to establish effective gene silencing. The transient nature of early VIGS systems highlighted the need for enhanced stability, mirroring the challenges faced in therapeutic applications [5].
Chemical Modification Strategies for Enhanced siRNA Stability
Chemical modifications represent the foremost approach for addressing the inherent limitations of unmodified siRNAs. These modifications are strategically designed to enhance nuclease resistance, improve binding affinity, reduce immunogenicity, and optimize pharmacokinetic properties while preserving RNAi activity [71] [72]. The selection and combination of chemical modifications must balance stability enhancement with maintained functionality through the RNAi pathway.
Ribose Sugar Modifications
Modifications at the 2'-position of the ribose sugar are among the most frequently employed and well-tolerated changes to siRNA structure:
- 2'-O-methyl (2'-OMe): Improves nuclease resistance and reduces immune stimulation by mimicking naturally occurring mammalian RNA modifications [71] [73]
- 2'-fluoro (2'-F): Significantly enhances resistance to enzymatic degradation while maintaining high RNAi activity [71]
- 2'-O-methoxyethyl (2'-MOE): Further enhances stability and reduces toxicity compared to 2'-OMe [71]
- Locked Nucleic Acid (LNA): Incorporates a methylene bridge between the 2'-oxygen and 4'-carbon, creating a rigid bicyclic structure that confers exceptionally high binding affinity and nuclease resistance [72]
Table 1: Common Ribose Modifications and Their Properties
| Modification | Nuclease Resistance | Binding Affinity | Immunogenicity | Toxicity Profile |
|---|---|---|---|---|
| 2'-O-methyl | Moderate improvement | Moderate increase | Reduced | Low |
| 2'-fluoro | Significant improvement | Maintained | Reduced | Low |
| 2'-O-methoxyethyl | High improvement | Increased | Reduced | Low |
| LNA | Very high improvement | Very high increase | Variable | Moderate at high usage |
Phosphate Backbone Modifications
The phosphate backbone of siRNAs is critical for molecular stability and cellular uptake, with several key modifications employed:
- Phosphorothioate (PS): Replacement of a non-bridging oxygen with sulfur significantly increases resistance to nuclease activity and enhances protein binding, improving pharmacokinetics [71] [72]
- Phosphorodithioate: Both non-bridging oxygens replaced by sulfur, offering even greater protection against nucleases [71]
- Boranophosphate: Incorporation of a borane moiety alters backbone properties and enhances stability [73]
- Methylphosphonate: Renders the backbone neutral, reducing immunogenicity without compromising cellular delivery [71]
Terminal and Conjugate Modifications
Strategic modifications at siRNA termini further enhance pharmacological properties:
- 5'--(E)-vinylphosphonate: Serves as a non-hydrolyzable phosphate analog that enhances tissue accumulation and efficacy in vivo [74] [73]
- GalNAc conjugation: Enables hepatocyte-specific delivery through targeting of the asialoglycoprotein receptor, with demonstrated clinical success [71] [52]
- Cholesterol conjugation: Improves association with serum lipoproteins and enhances uptake in liver and other tissues [73]
- 3'-amino groups: Facilitate improved cell membrane interaction and uptake [71]
Systematic analysis of modification patterns reveals that the level of 2'-O-methyl content significantly impacts efficacy, while structural features like symmetric versus asymmetric configurations show less pronounced effects [52]. Molecular modeling studies further indicate that modifications at specific positions, particularly in the seed region of the guide strand (e.g., position g2), play an outsized role in determining biological activity [74].
Diagram 1: Chemical Modification Strategies for Enhanced siRNA Stability (Short Title: siRNA Modification Pathways)
Advanced Delivery Systems for siRNA Formulation
While chemical modifications address molecular stability, advanced delivery systems are crucial for overcoming biological barriers to cellular uptake and intracellular trafficking. The development of efficient delivery platforms represents perhaps the most significant advancement in enabling therapeutic siRNA applications.
Lipid-Based Nanoparticles
Lipid-based vectors remain the most clinically advanced siRNA delivery platform:
- Lipid Nanoparticles (LNPs): Combination of ionizable lipids, helper lipids, PEGylated lipids, and cholesterol that form stable vesicles for siRNA encapsulation; demonstrated in FDA-approved patisiran [71]
- Cationic Liposomes: Positively charged lipids that complex with negatively charged siRNA, enhancing cellular uptake though sometimes limited by toxicity [71]
- Solid Lipid Nanoparticles (SLNs): Offer controlled release profiles and scalable production processes [71]
- Cholesterol-Enriched Exosomes: Engineered exosomal platforms that enable siRNA to bypass endosomal entrapment via direct membrane fusion, significantly improving cytosolic delivery [75]
Targeted Conjugates and Alternative Platforms
- GalNAc-siRNA Conjugates: Triantennary N-acetylgalactosamine motifs that target the asialoglycoprotein receptor highly expressed on hepatocytes, enabling efficient liver-specific delivery with demonstrated clinical success [52] [72]
- Polymer-Based Vectors: Cationic polymers (PEI, PLL, chitosan) that electrostatically complex with siRNA, enhancing cellular uptake and facilitating endosomal escape [71]
- Inorganic Nanoparticles: Gold nanoparticles, mesoporous silica, and magnetic nanoparticles that offer tunable properties for controlled siRNA release [71]
Table 2: siRNA Delivery Systems and Applications
| Delivery System | Key Components | Advantages | Clinical Status |
|---|---|---|---|
| Lipid Nanoparticles (LNPs) | Ionizable lipids, PEG-lipids, cholesterol | Proven clinical success, good payload protection | Approved (patisiran) |
| GalNAc Conjugates | Triantennary GalNAc, stabilized siRNA | Excellent hepatocyte targeting, simple formulation | Multiple approved drugs |
| Polymer-Based Nanoparticles | PEI, PLL, PAMAM dendrimers | Tunable properties, enhanced endosomal escape | Preclinical/Clinical development |
| Exosomal Delivery | Engineered exosomes, cholesterol enrichment | Biocompatible, membrane fusion mechanism | Preclinical development |
Analysis of publication trends reveals that 63% of siRNA research mentions non-viral delivery systems, while 37% utilize viral vectors [71]. Lipid-based vectors show the most prominent increase in publications in recent years, while siRNA conjugates also demonstrate a subtle increase [71].
Experimental Protocols for Evaluating siRNA Stability and Efficacy
Robust experimental methodologies are essential for evaluating the impact of chemical modifications and delivery systems on siRNA stability and silencing activity. The following protocols represent state-of-the-art approaches in the field.
siRNA Design and Modification Strategy
Target Sequence Selection: Identify 20-nucleotide target sequences within mRNA transcripts of interest, excluding sequences with ≥60% G-C content or CCCC/GGGG stretches due to synthetic limitations and potential negative impacts on silencing [52]
Off-Target Filtering: Exclude siRNAs with guide strand positions 2-17 showing significant homology to other human genes to minimize off-target effects [52]
Modification Pattern Design: Implement position-specific modification patterns incorporating 2'-O-methyl, 2'-fluoro, and phosphorothioate modifications, focusing on:
Synthesis: Conduct oligonucleotide synthesis via phosphoramidite solid-phase synthesis using modified 2'-F, 2'-OMe phosphoramidites with standard protecting groups [52]
Stability and Thermodynamic Profiling
Serum Stability Assay:
- Incubate 5 μM siRNA in 50% fetal bovine serum at 37°C
- Withdraw aliquots at 0, 1, 2, 4, 8, 12, and 24 hours
- Analyze integrity by denaturing polyacrylamide gel electrophoresis or HPLC
- Calculate half-life based on degradation kinetics [73]
Thermal Melting Analysis:
- Prepare 5 μM siRNA duplex in appropriate buffer (e.g., 10 mM sodium phosphate, 100 mM NaCl, pH 7.0)
- Measure UV absorption at 260 nm while increasing temperature from 20°C to 95°C at 1°C/minute
- Determine melting temperature (Tm) from first derivative of melting curve [74]
Molecular Dynamics Simulations:
- Perform all-atomic molecular dynamics simulations on parent and chemically modified siRNA duplexes
- Analyze structural, dynamic, and energetic properties over microsecond timescales
- Calculate interaction energies and sugar stereochemical flexibility, particularly in seed region [74]
In Vitro Silencing Efficacy Assessment
Cell Culture and Transfection:
- Maintain appropriate cell lines (e.g., MCF7, BT474, HCT116) under standard conditions
- Plate cells at 50-60% confluence in 24-well plates 24 hours before transfection
- Transfect with siRNA using recommended transfection reagents (e.g., RNAiMAX, Lipofectamine 2000)
- Use concentration range from 0.1 nM to 100 nM to establish dose-response curves [52] [76]
mRNA Quantification:
- Harvest cells 24-72 hours post-transfection
- Extract total RNA using commercial kits
- Perform quantitative RT-PCR or QuantiGene assays to measure target mRNA levels
- Normalize to appropriate housekeeping genes [52]
Protein Analysis:
- Harvest cells 48-96 hours post-transfection for protein analysis
- Perform Western blotting or ELISA to quantify target protein reduction
- Calculate IC50 values from dose-response data [74]
Diagram 2: Experimental Workflow for siRNA Stability and Efficacy Assessment (Short Title: siRNA Evaluation Workflow)
The Scientist's Toolkit: Essential Reagents and Methodologies
Table 3: Key Research Reagent Solutions for siRNA Studies
| Reagent/Method | Function | Key Considerations |
|---|---|---|
| Phosphoramidite Chemistry | Solid-phase siRNA synthesis | Enables incorporation of 2'-OMe, 2'-F, and other modified nucleotides |
| HPLC Purification | Purification of synthesized siRNA | Critical for obtaining highly pure, functional siRNA compounds |
| RNAiMAX Transfection Reagent | In vitro siRNA delivery | Efficient cellular uptake with low toxicity; suitable for dose-response studies |
| QuantiGene Assay | mRNA quantification | Direct mRNA measurement without reverse transcription; avoids PCR biases |
| Molecular Dynamics Software | Structural and energetic analysis | Provides insights into modification effects on duplex stability and RISC compatibility |
| Lipid Nanoparticle Formulation | In vivo siRNA delivery | Protects siRNA from degradation; enables tissue-specific delivery |
| GalNAc Conjugation Chemistry | Hepatocyte-targeted delivery | Facilitates receptor-mediated uptake in liver; simple formulation |
The field of siRNA therapeutics continues to evolve rapidly, with several emerging trends shaping future research directions. The ongoing transition from empirical, systemic delivery to precision, tissue-directed platforms represents the defining technological shift in the RNAi drug delivery market [77]. Innovations in ligand targeting, cleavable linkers, and biodegradable LNP backbones are progressively improving cell specificity while reducing systemic exposure [77].
Key future directions include:
Expansion Beyond Hepatic Targets: While current successes predominantly focus on liver applications, extensive research aims to enable efficient siRNA delivery to CNS, lung, muscle, and tumoral microenvironments [77] [75]
Oral Delivery Platforms: Recent advances in engineered exosomes and protective formulations demonstrate promising results for oral siRNA administration, potentially revolutionizing treatment paradigms for chronic diseases [75]
Integration with Emerging Technologies: Combination of siRNA with gene editing (CRISPR/Cas) and other oligonucleotide modalities creates opportunities for synergistic therapeutic approaches [77]
Computational Design and AI Integration: Machine learning approaches and molecular modeling are increasingly informing siRNA design, modification patterns, and target selection, potentially accelerating optimized therapeutic development [74] [78]
Sustainable Manufacturing: Greener oligonucleotide synthesis methods, solvent recovery, and reduced energy intensity in production are emerging as important considerations for scalable manufacturing [77]
In the context of VIGS research, advances in siRNA stability and delivery echo nature's solutions to viral persistence in plants, where RNA-directed DNA methylation and transgenerational epigenetic inheritance of silencing states provide long-lasting protection [5]. The molecular machinery underlying these phenomena—including Dicer-like enzymes, ARGONAUTE proteins, and RNA-dependent RNA polymerases—represents the evolutionary foundation upon which modern siRNA therapeutics are built [5].
As chemical modification strategies and delivery platforms continue to mature, siRNA therapeutics are poised to expand their impact across increasingly diverse therapeutic areas. With sustained innovation addressing challenges of extrahepatic delivery, manufacturing scalability, and long-term safety profiles, siRNA technology stands to fully realize its potential as a transformative class of medicines, ultimately fulfilling the promise suggested by nature's own antiviral defense mechanisms observed in VIGS.
Validation Frameworks and Technology Comparison
In the study of gene function, particularly within the framework of siRNA-based Virus-Induced Gene Silencing (VIGS), robust phenotypic and molecular validation is paramount. VIGS is an RNA-mediated reverse genetics technology that uses recombinant viral vectors to trigger post-transcriptional gene silencing (PTGS) of endogenous plant genes by exploiting the plant's own antiviral defense mechanisms [4] [5]. This process relies on the generation of 21–24 nucleotide small interfering RNAs (siRNAs) that guide the sequence-specific degradation of complementary target mRNAs [5]. The successful application of this technology, whether for functional genomics or crop improvement, hinges on a multi-tiered validation strategy that connects the observed phenotype with molecular confirmation of gene knockdown at both the RNA and protein levels.
This guide provides an in-depth technical framework for researchers requiring comprehensive validation methodologies, from the initial confirmation of reduced transcript levels via Reverse Transcription-quantitative PCR (RT-qPCR) to the final demonstration of diminished target protein expression. Within the context of a broader thesis on siRNA in VIGS research, this workflow is not merely a series of technical steps but a critical chain of evidence that confirms the specificity and efficacy of the silencing trigger, ensuring that observed phenotypic changes are a direct consequence of the intended gene silencing event.
The Molecular Basis of VIGS and siRNA Action
Understanding the validation workflow requires a foundational knowledge of the VIGS mechanism. Figure 1 illustrates the key steps from vector inoculation to phenotypic readout.
The process begins with the design of a viral vector (e.g., Tobacco Rattle Virus - TRV) carrying a fragment of the plant's endogenous gene to be silenced [4]. Upon delivery into the plant, typically via Agrobacterium-mediated infiltration, the virus replicates and moves systemically. The plant's defense machinery recognizes the viral double-stranded RNA (dsRNA), and Dicer-like enzymes process it into small interfering RNAs (siRNAs) [5]. These siRNAs are then incorporated into an RNA-Induced Silencing Complex (RISC), which guides the complex to complementary endogenous mRNA transcripts, leading to their cleavage and degradation—a process known as Post-Transcriptional Gene Silencing (PTGS) [4] [5]. The ultimate consequence is a reduction in the corresponding protein levels, leading to a measurable phenotype, which must be rigorously validated.
Tier 1 Validation: Transcript-Level Analysis by RT-qPCR
The first and most common line of molecular evidence is the quantification of target mRNA reduction using RT-qPCR. This technique provides sensitive and specific quantification of transcript abundance.
Detailed RT-qPCR Workflow
The entire process, from RNA to quantitative data, is outlined in Figure 2 and described in detail below.
RNA Isolation and Quality Control
The accuracy of RT-qPCR is critically dependent on high-quality RNA.
- Isolation: Use kits designed for your sample type (e.g., PureLink RNA Mini Kit for fresh tissue, MagMAX-96 for high-throughput, RNAqueous-Micro for micro-samples) [79]. The isolation method should be chosen based on the need to capture large RNAs only (mRNA, rRNA) or both large and small RNAs (including miRNAs) [79].
- Genomic DNA Removal: A mandatory DNase treatment step is required to prevent genomic DNA amplification and false positives [80] [79].
- Quality Control: Fluorometric quantification using assays like Qubit or Quant-iT is superior to UV absorbance for specific RNA quantification, as it is less affected by contaminants like free nucleotides or DNA [79]. RNA integrity should also be checked, for example via agarose gel electrophoresis.
Reverse Transcription (RT) and Primer Design
This step converts RNA into a more stable cDNA template.
- RT Reagents: The reaction requires reverse transcriptase, dNTPs, MgCl₂, RNase inhibitors, and primers [80].
- Primer Choice: The choice of primer for cDNA synthesis depends on the target RNA.
- Gene-specific primers: Provide the highest sensitivity and specificity for targeted genes [80].
- Oligo(dT) primers: Bind to the poly-A tail of mRNAs, suitable for most protein-coding transcripts [80].
- Random primers: Reverse-transcribe a broad population of RNAs, including non-polyadenylated transcripts, but may produce less full-length cDNA for long mRNAs [80].
- Thermal Cycling: The RT reaction typically involves denaturing RNA secondary structures (65–70°C for 5–10 min), primer annealing, cDNA synthesis (37–50°C for 30–60 min), and enzyme inactivation (70–85°C) [80].
Quantitative PCR (qPCR) and Data Analysis
- qPCR Reagents: The reaction mixture includes DNA polymerase, sequence-specific primers, dNTPs, MgCl₂, and a fluorescent detection system (e.g., SYBR Green dye or TaqMan probes) [80].
- Primer Design for qPCR:
- Specificity: Design primers to span exon-exon junctions to prevent amplification of genomic DNA [80]. Use tools like NCBI BLAST to confirm specificity.
- Amplicon Length: Keep the product length between 70–200 base pairs for optimal amplification efficiency [80].
- Primer Properties: Optimal primers are 18–25 nucleotides long with a GC content of 40–60% to ensure stable binding [80]. Use tools like OligoAnalyzer to calculate melting temperature (Tm) and check for secondary structures with Primer3PLUS.
- Thermal Cycling: The process involves initial denaturation (95°C), followed by 30–40 cycles of denaturation (95°C), primer annealing (55–65°C), and extension (72°C), with fluorescence measured at each cycle [80].
- Data Analysis: The Cycle Threshold (Ct) is the key output, representing the cycle number at which the fluorescence crosses a defined threshold. A lower Ct value indicates a higher starting quantity of the target transcript [80].
- Absolute Quantification: Determines the exact copy number of the target RNA by comparing Ct values to a standard curve of known concentrations [80].
- Relative Quantification (RQ): More commonly used, this method calculates the expression level of the target gene relative to one or more stable reference genes (e.g., actin, GAPDH) in treated versus control samples, often using the 2^(-ΔΔCt) method [80].
Research Reagent Solutions for RT-qPCR
Table 1: Essential reagents and kits for the RT-qPCR workflow.
| Process Step | Reagent / Kit | Key Function | Technical Notes |
|---|---|---|---|
| RNA Isolation | PureLink RNA Mini Kit [79] | Rapid silica-column based RNA purification. | Isolates large RNA only; 20 min procedure. |
| MagMAX-96 Total RNA Isolation Kit [79] | High-throughput purification using magnetic beads. | Suitable for up to 100,000 cells; <45 min. | |
| RNAqueous-Micro Kit [79] | Purification of RNA from precious, micro-sized samples. | Isolates small & large RNAs; 15 min. | |
| RNA Quantification | Qubit RNA Assays [79] | Fluorometric, specific RNA quantification. | More accurate than UV absorbance for low-concentration or contaminated samples. |
| Reverse Transcription | SuperScript IV VILO Master Mix [79] | Converts RNA to cDNA with high efficiency and reduced bias. | Includes reverse transcriptase and optimized buffer in a single tube. |
| qPCR | SYBR Green dye [80] | Binds dsDNA; emits fluorescence during amplification. | Cost-effective; requires optimization of primer specificity to avoid off-target signals. |
| TaqMan probes [80] | Sequence-specific hydrolysis probes for highly specific detection. | Higher specificity than intercalating dyes; more expensive. |
Tier 2 Validation: Protein-Level Analysis
Confirming a reduction in the corresponding protein level is crucial, as transcript knockdown does not always lead to a proportional decrease in protein abundance. Table 2 summarizes common protein analysis techniques used in VIGS validation.
Key Protein Analysis Techniques
Table 2: A comparison of primary methods for protein-level validation.
| Method | Principle | Application in VIGS/siRNA Research | Advantages | Limitations |
|---|---|---|---|---|
| Enzyme-Linked Immunosorbent Assay (ELISA) | Uses antibodies to detect and quantify a specific protein in a sample. | Directly measure levels of the target protein (e.g., Cry1C in transgenic rice) [81]. | High specificity and sensitivity; quantitative; high-throughput. | Requires a specific, high-quality antibody. |
| Western Blotting | Proteins separated by gel electrophoresis, transferred to a membrane, and detected with specific antibodies. | Confirm silencing of a specific protein and estimate its molecular weight. | Semi-quantitative; provides information on protein size and integrity. | Less quantitative than ELISA; requires optimization. |
| Parallel Reaction Monitoring (PRM) | A targeted mass spectrometry approach that monitors specific peptides from the protein of interest. | Perform targeted proteomic quantification to verify protein reduction and study unintended effects [81]. | High specificity and multiplexing capability; absolute quantification possible; does not require an antibody. | Requires specialized, expensive equipment and expertise. |
In the context of validating VIGS, a study on Bt transgenic rice provides an excellent example. Researchers used ELISA to quantitatively measure the levels of Cry1C and Cry2A proteins in transgenic lines, confirming the expression of the introduced genes at the protein level [81]. Furthermore, to investigate unintended effects at the proteome level, they employed high-throughput proteomics, identifying differentially expressed proteins (DEPs) between transgenic and control plants, and used PRM analysis for targeted verification of specific proteins [81].
Correlating Molecular Data with Phenotypic Outcomes
The final step in the validation chain is to rigorously correlate molecular data with the observed phenotype. In VIGS studies, this often involves a visible or measurable trait.
- Example 1 (Visible Phenotype): Silencing the Phytoene Desaturase (PDS) gene, involved in carotenoid biosynthesis, results in a characteristic photo-bleaching phenotype. Successful validation would show a strong correlation between reduced PDS mRNA levels (via RT-qPCR), reduced PDS protein (e.g., via Western Blot), and the appearance of white or bleached leaves [4].
- Example 2 (Quantitative Phenotype): In a study on transgenic rice, the introduction of CRY1C and CRY2A genes led to significantly higher resistance to stem borers. This phenotypic data (reduced deadheart stems and whitehead panicles) was directly correlated with higher CRY gene expression (via qRT-PCR) and the presence of Cry proteins (via ELISA) [81].
A comprehensive phenotypic and molecular validation strategy, integrating transcript analysis (RT-qPCR) with protein-level confirmation (ELISA, Western Blot, PRM), is the cornerstone of rigorous VIGS and siRNA research. This multi-faceted approach moves beyond simple correlation to establish a causal link between the siRNA-triggered silencing event, the depletion of the target transcript and protein, and the resulting physiological phenotype. As VIGS technology evolves, including the development of non-transgenic delivery methods using minimal siRNA precursors [24] and its application for inducing heritable epigenetic modifications [5], the need for robust, reproducible validation workflows becomes ever more critical. The protocols and frameworks detailed in this guide provide a reliable path for researchers to confidently characterize gene function and advance both fundamental science and applied crop improvement.
Within modern plant functional genomics, researchers require tools that can bridge the gap between high-throughput gene discovery and detailed functional validation. This technical guide provides a comparative analysis of two pivotal technologies: Virus-Induced Gene Silencing (VIGS), an RNA-silencing based approach, and CRISPR/Cas9-mediated stable transformation. Framed within the context of a broader thesis on the role of small interfering RNA (siRNA) in plant research, this document delineates the operational paradigms, efficiencies, and practical applications of each method. The core of VIGS technology leverages the innate RNA interference (RNAi) machinery of plants, where siRNAs serve as the central effector molecules guiding sequence-specific mRNA degradation [42]. In contrast, CRISPR/Cas9 operates at the DNA level, creating permanent genomic modifications. This in-depth analysis, designed for researchers and drug development professionals, summarizes quantitative data in structured tables, provides detailed experimental protocols, and visualizes key signaling pathways and workflows to serve as a comprehensive technical resource.
Virus-Induced Gene Silencing (VIGS)
2.1.1 Core Mechanism and the Role of siRNA VIGS is a powerful reverse genetics technique that leverages the plant's innate RNA-mediated defense system against viruses. The process initiates when a recombinant viral vector, carrying a fragment of a host target gene, is introduced into the plant. During viral replication, double-stranded RNA (dsRNA) intermediates are formed, which are recognized and cleaved by the plant's Dicer or Dicer-like (DCL) nucleases into 21-24 nucleotide small interfering RNAs (siRNAs) [42]. These siRNAs are then loaded into an RNA-induced silencing complex (RISC). The single-stranded siRNA within RISC guides the complex to complementary mRNA sequences, leading to the cleavage and degradation of the target transcript, thereby resulting in gene silencing [42]. This post-transcriptional gene silencing (PTGS) mechanism allows for the rapid, transient downregulation of endogenous genes without stable genetic modification.
2.1.2 Key VIGS Vectors The Tobacco Rattle Virus (TRV) is the most widely adopted VIGS vector, demonstrated in over 25 plant species across families including Solanaceae, Brassicaceae, and Malvaceae [82]. Other vectors include Potato Virus X (PVX), Barley Stripe Mosaic Virus (BSMV), and Tomato Yellow Leaf Curl Virus (TYLCV), each with differing host ranges and silencing efficiencies [42].
CRISPR/Cas9 and Stable Transformation
2.2.1 Core Genome-Editing Mechanism The CRISPR/Cas9 system functions as a precise DNA-editing tool. It consists of two key components: the Cas9 endonuclease and a single-guide RNA (sgRNA). The sgRNA directs the Cas9 protein to a specific genomic locus complementary to its sequence, where Cas9 induces a double-strand break (DSB) [83]. The cell subsequently repairs this break through one of two primary pathways:
- Non-Homologous End Joining (NHEJ): An error-prone repair pathway that often results in small insertions or deletions (indels) at the cut site, leading to gene knockouts [83].
- Homology-Directed Repair (HDR): A precise repair pathway that can be leveraged to introduce specific point mutations or insertions using a donor DNA template [83].
2.2.2 Stable Transformation Delivery Methods Stable integration of CRISPR/Cas9 components into the plant genome is typically achieved through:
- Agrobacterium-mediated transformation: Utilizes Agrobacterium tumefaciens to transfer T-DNA containing Cas9 and sgRNA expression cassettes into the plant genome. This is the most common method but can lead to complex integration patterns [83] [84].
- Biolistic transformation: Uses a gene gun to physically bombard cells with gold or tungsten particles coated with the CRISPR/Cas9 DNA [83].
A key advancement is the development of transgene-free editing systems, such as the delivery of pre-assembled Cas9-gRNA ribonucleoprotein (RNP) complexes into protoplasts or the use of viral vectors for Virus-Induced Genome Editing (VIGE), which circumvent the need for stable T-DNA integration [83] [43].
Table 1: Core Technology Comparison
| Feature | VIGS | CRISPR/Cas9 & Stable Transformation |
|---|---|---|
| Molecular Level | mRNA (Post-transcriptional) | DNA (Genomic) |
| Core Effector Molecule | siRNA [42] | Cas9 Nuclease & sgRNA [83] |
| Primary Outcome | Transient gene knockdown | Permanent gene knockout, knock-in, or base editing |
| Key Repair/Effector Pathway | RNAi / PTGS [42] | NHEJ / HDR [83] |
| Typical Delivery | Viral vector infection [82] | Agrobacterium transformation or biolistics [83] |
| Heritability | Somatic, not heritable | Stable and heritable mutations [84] |
| Theoretical Basis | Plant antiviral defense [42] | Prokaryotic adaptive immune system [83] |
Comparative Technical Specifications
For research planning and experimental design, a direct comparison of the practical attributes of VIGS and CRISPR/Cas9 is essential. The following table summarizes key parameters, including speed, efficiency, and applicability.
Table 2: Practical Application and Performance Comparison
| Parameter | VIGS | CRISPR/Cas9 & Stable Transformation |
|---|---|---|
| Experimental Timeline | 1-3 weeks for silencing [82] | Several months to over a year for stable line generation [85] [84] |
| Editing/Knockdown Efficiency | Highly variable; can be optimized (e.g., 40.3% with TRV, improved by heat treatment) [85] | Variable; 5-17% reported in wheat, can be very high in model systems [84] |
| Mutational Spectrum | Transcript downregulation | Indels (NHEJ), precise edits (HDR), base substitutions (Base Editors) [83] |
| Off-Target Effects | Potential for off-target genes with high sequence similarity | Potential for off-target DNA cleavage; mitigated by high-fidelity Cas variants [83] |
| Host Range | Broad, depends on viral vector host range (e.g., TRV: >25 species) [82] | Technically universal, but limited by transformability/regeneration of host species |
| Handling of Essential Genes | Suitable for studying lethal knockouts due to transient nature [42] | Knockout of essential genes may block regeneration of stable lines |
| Multiplexing Capacity | Possible to silence gene families using conserved sequences [42] | High; multiple sgRNAs can be expressed from a single construct [83] |
| Regulatory Status (GMO) | Often considered a transient assay | Stable integrants are regulated as GMOs; transgene-free edits may be deregulated [43] |
The siRNA Pathway in VIGS: A Detailed Workflow
The following diagram illustrates the step-by-step mechanism of VIGS, highlighting the central role of siRNA generation and activity.
Figure 1: The siRNA-Mediated VIGS Workflow. The core process involves viral delivery and replication, formation of a dsRNA intermediate, DCL-mediated processing into siRNAs, and RISC-mediated cleavage of complementary target mRNA [42].
Experimental Protocols
Protocol: TRV-Based VIGS in Solanaceous Plants
This protocol is adapted from studies optimizing VIGS in tomatoes and other solanaceous crops [85].
1. Vector Construction:
- Vector Choice: Use the pTRV1 and pTRV2 binary vectors. pTRV2 carries the gene fragment for silencing.
- Target Gene Fragment Selection: Clone a ~200-500 bp fragment of the target gene (e.g., Phytoene desaturase, PDS as a visual marker) into the multiple cloning site of the pTRV2 vector.
- Controls: Always include a positive control (e.g., TRV2-PDS) and an empty vector control (TRV2-EV).
2. Agrobacterium Preparation:
- Transform the pTRV1 and recombinant pTRV2 vectors separately into Agrobacterium tumefaciens strain GV3101.
- Inoculate single colonies of each Agrobacterium culture in 5 mL of LB medium with appropriate antibiotics (e.g., kanamycin, rifampicin). Grow overnight at 28°C with shaking.
- Sub-culture 1 mL of the overnight culture into 50 mL of fresh LB with antibiotics and 10 mM MES buffer. Grow until OD600 reaches 1.0-2.0.
- Pellet the cells by centrifugation (3,500-5,000 g for 10-15 min). Resuspend the pellet in an induction buffer (10 mM MgCl2, 10 mM MES, 200 µM acetosyringone). Adjust the final OD600 to 1.5-2.0.
- Incubate the resuspended cultures at room temperature for 3-4 hours in the dark.
3. Plant Inoculation:
- Mix the pTRV1 and pTRV2 Agrobacterium suspensions in a 1:1 ratio.
- For tomato seedlings, use a 1 mL needleless syringe to infiltrate the mixture into the abaxial side of cotyledons or the first true leaves of 8-14 day-old seedlings. Alternatively, vacuum infiltration of whole seedlings can be performed.
- Keep inoculated plants in low-light conditions for 24 hours before transferring to standard growth chambers.
4. Optimization and Phenotyping:
- Heat Treatment: To increase silencing efficiency, subject inoculated plants to a 37°C heat treatment for 24 hours at 2-5 days post-inoculation (dpi) [85].
- Phenotype Monitoring: Observe for silencing phenotypes (e.g., photobleaching for PDS) starting from 7-10 dpi. Systemic silencing in new leaves is typically strongest between 2-4 weeks post-inoculation.
5. Validation:
- Harvest tissue from silenced and control areas.
- Extract total RNA and perform reverse transcription quantitative PCR (RT-qPCR) to quantify the downregulation of the target transcript.
- For protein-level validation, western blotting can be used if a specific antibody is available.
Protocol: Stable CRISPR/Cas9 Transformation in Wheat
This protocol summarizes the successful generation of heritable edits in hexaploid wheat using Agrobacterium-mediated transformation [84].
1. Vector Design and Construction:
- Cas9 Expression: Use a plant codon-optimized Cas9 gene driven by a strong constitutive promoter (e.g., maize Ubiquitin promoter).
- sgRNA Design: Design sgRNAs with high on-target activity and minimal off-target potential. For polyploid species like wheat, design guides that target all homeologs (tri-genome targeting) or specific homeologs using unique sequences.
- sgRNA Expression: Clone the sgRNA sequence under a Pol III promoter (e.g., Arabidopsis U6-26 or rice U3 promoter) in a binary vector.
- Selection Marker: Include a plant-selectable marker (e.g., nptII for kanamycin resistance) for screening transformants.
2. Plant Transformation:
- Explant Preparation: Use immature embryos from the wheat cultivar 'Fielder' as the target tissue.
- Agrobacterium Co-cultivation: Infect the immature embryos with Agrobacterium strain AGL1 or EHA105 carrying the CRISPR/Cas9 binary vector. Co-cultivate for 2-3 days on solid co-cultivation media.
- Selection and Regeneration: Transfer embryos to selection media containing kanamycin to inhibit the growth of non-transformed tissue. Induce callus formation and subsequent shoot regeneration on media with appropriate plant growth regulators (e.g., auxins and cytokinins).
- Rooting and Acclimatization: Transfer regenerated shoots to rooting media and subsequently transfer established plantlets to soil.
3. Molecular Analysis of T0 Plants:
- Genomic DNA Extraction: Extract DNA from leaf tissue of putative transgenic plants.
- PCR Screening: Confirm the presence of the T-DNA using PCR with primers specific to the Cas9 or selection marker gene.
- Editing Efficiency Analysis: Amplify the target genomic region from PCR-positive plants. Use Sanger sequencing followed by decomposition with tools like TIDE or ICE, or next-generation sequencing to determine the spectrum and frequency of induced mutations.
4. Segregation and Generation of Transgene-Free Plants:
- Seed Harvest: Collect T1 seeds from fertile T0 plants.
- Segregation Analysis: Grow T1 plants and screen by PCR to identify lines that have the desired gene edit but have lost the T-DNA (Cas9/sgRNA transgene). This typically follows a Mendelian ratio, allowing for the identification of transgene-free, edited homozygous lines in the T1 or T2 generation [84].
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Reagents for VIGS and CRISPR/Cas9 Research
| Reagent / Solution | Function / Application | Example Specifics / Notes |
|---|---|---|
| TRV VIGS Vectors (pTRV1/pTRV2) | Delivery of silencing constructs into plant cells via Agrobacterium [85]. | pTRV1 encodes viral replication proteins. pTRV2 carries the target gene insert. |
| Agrobacterium tumefaciens (GV3101) | Biological delivery vehicle for stable transformation and VIGS [85] [84]. | Disarmed strain; requires virulence (vir) genes for T-DNA transfer. |
| Acetosyringone | A phenolic compound that induces the Agrobacterium vir genes, crucial for efficient T-DNA transfer [85]. | Added to the Agrobacterium resuspension/infiltration buffer. |
| Binary T-DNA Vectors (e.g., pHSE401) | Carry expression cassettes for Cas9 and sgRNAs for plant transformation [86] [84]. | Contains left and right T-DNA borders, plant selection marker, and bacterial origin of replication. |
| Plant Codon-Optimized Cas9 | The effector nuclease that creates double-strand breaks in the target DNA [83] [84]. | Often driven by constitutive promoters like CaMV 35S or maize Ubi. |
| sgRNA Scaffold & Pol III Promoters | Provides the targeting specificity and structural framework for Cas9 binding. Promoters (e.g., AtU6, OsU3) drive high-level sgRNA expression [84]. | The sgRNA is a fusion of crRNA (targeting) and tracrRNA (scaffold). |
| Selection Agents (e.g., Kanamycin) | Allows for the selective growth of transformed plant cells or tissues [87] [84]. | Concentration must be optimized for each plant species (e.g., 50 mg/L for wheat embryos [84]). |
| PCR/RE Assay or T7 Endonuclease I | Methods for detecting CRISPR/Cas9-induced mutations [83]. | Detect mismatches in heteroduplex DNA formed by wild-type and mutant alleles. |
| PEG6000 (for drought stress studies) | Used to simulate osmotic/drought stress in functional studies of edited or silenced plants [87]. | Applied to growth media or soil to create a controlled water-deficit environment. |
VIGS and CRISPR/Cas9 with stable transformation represent complementary pillars of modern plant functional genomics. The selection between these technologies is not a matter of superiority but of strategic alignment with experimental goals. VIGS, with its foundation in the siRNA pathway, offers an unparalleled rapid, transient, and flexible system for high-throughput functional screening and studying essential genes in a wide range of species. CRISPR/Cas9 stable transformation provides the definitive tool for establishing causal relationships through the creation of stable, heritable genetic variants, which is indispensable for both basic research and applied crop breeding. As the field advances, the convergence of these technologies—exemplified by Virus-Induced Genome Editing (VIGE)—promises to further accelerate the pace of discovery. By leveraging the deep mechanistic understanding of siRNA biology and the precision of DNA editing, researchers are now equipped to systematically deconstruct complex traits and engineer resilient crops for future challenges.
Within Virus-Induced Gene Silencing (VIGS) research, small interfering RNA (siRNA) is a cornerstone molecule that directs post-transcriptional gene silencing. This whitepaper provides a technical analysis of the core characteristics of siRNA-mediated silencing in the VIGS context: its rapid onset, sequence-specific targeting, and transient effect. We detail how these properties make siRNA an indispensable tool for high-throughput functional genomics and initial gene validation, while also presenting limitations for studying essential genes or requiring long-term silencing. Supported by current data and experimental protocols, this review equips researchers with a framework to strategically leverage siRNA in VIGS for applications ranging from basic gene discovery to crop improvement and drug target identification.
Virus-Induced Gene Silencing (VIGS) is a powerful functional genomics tool that leverages the plant's innate RNA interference (RNAi) machinery to suppress target gene expression. The process is initiated by a recombinant viral vector carrying a fragment of a host gene, which is subsequently processed by the plant into primary small interfering RNAs (siRNAs). These 21–24 nucleotide siRNAs serve as the sequence-specific guide molecules that direct the RNA-induced silencing complex (RISC) to cleave complementary messenger RNA (mRNA), leading to gene knockdown [4]. The core attributes of siRNA—its rapid induction, high specificity, and transient nature—directly define the utility and limitations of the VIGS platform. For research scientists, a nuanced understanding of these characteristics is critical for experimental design, from initial gene screening to the validation of phenotypic effects in both plant and animal systems.
Core Characteristics: A Technical Examination
Speed: Rapid Induction of Gene Silencing
The speed of siRNA accumulation and action is a primary advantage of VIGS over stable genetic transformation. Silencing phenotypes can manifest within 1-2 weeks post-inoculation with the viral vector, enabling high-throughput functional screening [4].
- Mechanistic Basis: The speed is derived from the amplification of the silencing signal. The initial siRNAs derived from the viral RNA can trigger the production of secondary, or transitive siRNAs, by host RNA-dependent RNA polymerases (RDRs). This amplification creates a robust and rapid systemic silencing response [4] [88].
- Key Factor: The size of the siRNA precursor insert in the viral vector is a critical determinant for the speed and efficiency of silencing. Recent advances show that virus-delivered short RNA inserts (vsRNAi) as short as 24–32 nucleotides can effectively trigger silencing, simplifying vector engineering and potentially accelerating the process [89].
Table 1: Key Factors Influencing the Speed of siRNA-Mediated VIGS
| Factor | Impact on Speed | Experimental Consideration |
|---|---|---|
| Viral Vector Selection | Different vectors have varying replication and movement rates. | Tobacco Rattle Virus (TRV) is widely used for its efficient systemic movement [4]. |
| Insert Size | Shorter, optimized inserts may lead to faster processing. | vsRNAi (24-32 nt) can induce robust silencing comparable to longer ~300 nt fragments [89]. |
| Plant Growth Conditions | Temperature and light intensity affect viral replication and RNAi machinery. | Optimal temperature control (often ~21°C) is crucial for maximizing silencing efficiency [4]. |
| Inoculation Method | Influences the initial infection efficiency. | Agroinfiltration is a common and effective delivery method for the VIGS construct [4]. |
Specificity: Precision and Off-Target Risks
Specificity refers to the ability of siRNA to silence only the intended target gene. This is achieved through perfect or near-perfect sequence complementarity between the siRNA guide strand and the target mRNA [90].
- Molecular Mechanism: The Argonaute (AGO) protein within RISC performs "slicer" activity, cleaving the target mRNA at a precise site defined by the siRNA [11]. This mechanism allows for the discrimination of even single-nucleotide polymorphisms when the siRNA is well-designed.
- Limitation - Off-Target Effects: A significant challenge is the potential for siRNAs to silence non-target genes with partial sequence complementarity, particularly in short "seed" regions. This can lead to misinterpretation of phenotypic data [11]. In VIGS, the use of long inserts (200-400 nt) traditionally increases the risk of generating siRNAs that target multiple homologous genes.
- Strategies for Enhancement:
- Bioinformatic Design: Careful selection of target sequences with low homology to other genes in the genome is essential.
- vsRNAi Approach: Using defined, short RNA inserts (e.g., 32-nt) minimizes the pool of siRNAs generated, thereby reducing off-target potential and simplifying specificity validation [89].
- Chemical Modifications: In therapeutic settings, synthetic siRNAs are chemically modified (e.g., 2'-O-methyl) to enhance stability and specificity [91] [92]. These strategies are increasingly relevant for topical application of dsRNA/siRNA in plants [88].
Diagram 1: siRNA Specificity and Off-Target Pathways. The pathway leading to specific mRNA cleavage is triggered by high complementarity, while off-target effects often arise from binding to targets with partial complementarity, primarily through the seed region.
Transient Nature: A Double-Edged Sword
The transient, non-integrated nature of VIGS is one of its most defining features. Silencing is not permanent, typically lasting for several weeks to months, but eventually being diluted as the virus is cleared and plant cells divide [4] [46].
- Advantages of Transiency:
- Study of Essential Genes: Enables the study of genes whose permanent knockout would be lethal. The temporary knockdown allows observation of phenotypes that might be impossible to study with stable mutants [11].
- Functional Redundancy: In polyploid species or genes with functional redundancy, simultaneous transient silencing of multiple homologs can reveal their collective function [89].
- Reversibility: The phenotype can be reversed, allowing the same plant to be used for multiple experimental manipulations or to confirm that the observed effect is due to the silencing of the target gene.
- Limitations of Transiency:
- Temporal Constraints: Not suitable for studying gene functions required at late developmental stages or those with long-term phenotypic consequences.
- Incomplete Penetrance: Silencing efficiency can vary between tissues and over time, leading to mosaic patterns and potential misinterpretation [4].
- Instability of Signal: The silencing effect can be diluted in rapidly dividing cells, making it less effective in meristematic tissues [4].
Table 2: Advantages and Limitations of Transient siRNA Silencing in VIGS
| Aspect | Advantages | Limitations |
|---|---|---|
| Experimental Flexibility | Enables high-throughput screening; no stable transformation required. | Silencing duration is limited and variable, complicating long-term studies. |
| Phenotypic Analysis | Allows study of lethal genes; phenotype is reversible. | Tissue-specific variation can result in incomplete or mosaic silencing. |
| Therapeutic Application | Reduced risk of permanent genomic alterations. | Requires re-application for chronic conditions; delivery efficiency is a recurring challenge. |
| Breeding & Agriculture | Rapid assessment of gene function for trait evaluation; transgene-free. | Not heritable, limiting its direct use in developing stable crop lines. |
Advanced Protocols and Methodologies
Protocol: Establishing VIGS with TRV Vectors inNicotiana benthamiana
This protocol outlines the standard procedure for implementing VIGS using the widely adopted Tobacco Rattle Virus (TRV) system.
Vector Selection and Clone Design:
- Use the bipartite TRV system: TRV1 (encoding replication and movement proteins) and TRV2 (encoding coat protein and the target gene insert) [4].
- Clone a 300-500 bp fragment of the target gene into the TRV2 vector via restriction digestion and ligation. Alternatively, for vsRNAi, clone a synthesized 32-nt oligonucleotide with 100% identity to the target into the TRV2 vector [89].
Agrobacterium Transformation and Culture:
- Transform the recombinant TRV2 and the helper TRV1 plasmids into Agrobacterium tumefaciens strain GV3101.
- Culture individual colonies in LB broth with appropriate antibiotics. Induce the bacteria with acetosyringone (200 µM) to activate virulence genes.
Plant Inoculation:
- Grow N. benthamiana plants to the 4-6 leaf stage.
- Mix the TRV1 and TRV2 Agrobacterium cultures in a 1:1 ratio.
- Use a needleless syringe to infiltrate the mixture into the abaxial side of fully expanded leaves. The optical density at 600 nm (OD₆₀₀) is typically adjusted to 0.5-1.0 for optimal results [4].
Phenotype Monitoring and Validation:
- Maintain plants under controlled conditions (e.g., 21°C, 16/8h light/dark) to optimize silencing.
- Observe systemic (non-infiltrated) leaves for phenotypic changes 10-21 days post-infiltration.
- Validate silencing efficiency by RT-qPCR to measure target mRNA knockdown and/or by western blot to assess protein level reduction.
Protocol: Topical Application of 22-nt siRNAs for Enhanced Silencing
This protocol demonstrates an alternative, transgene-free method for siRNA delivery, highlighting the functional significance of siRNA length.
siRNA Design and Synthesis:
- Design a 22-nt siRNA guide strand with a 2-nt 3' overhang, targeting a conserved region of the endogenous gene of interest (e.g., CHL-I, CHL-H) [88].
- Synthesize the siRNA duplex commercially with standard purity.
Topical Delivery to Leaves:
- Use an abrasion method. Gently abrade the surface of intact leaves (e.g., of N. benthamiana or Amaranthus cruentus) with fine-grit sandpaper.
- Immediately apply a solution containing the 22-nt siRNA (concentration ~50-100 µM) in a buffer, gently spreading it over the abraded area [88].
Analysis of Silencing Efficacy:
- Monitor treated leaves for local silencing phenotypes (e.g., yellowing for chlorophyll genes).
- Assess the production of transitive siRNAs via small RNA sequencing, a hallmark of the amplified silencing response triggered by 22-nt siRNAs.
- Compare the local silencing strength and systemic movement potential against 21-nt siRNAs targeting the same sequence.
The Scientist's Toolkit: Essential Reagents for VIGS
Table 3: Key Research Reagent Solutions for siRNA VIGS Experiments
| Reagent / Solution | Function & Role in VIGS | Example & Notes |
|---|---|---|
| TRV Viral Vectors (pTRV1, pTRV2) | Backbone for delivering the target gene insert; TRV1 facilitates replication, TRV2 carries the insert. | Available from Addgene; pLX-TRV2-vCHLI (Addgene #239842) is a vsRNAi-ready vector [89]. |
| Agrobacterium tumefaciens | Bacterial vehicle for delivering the viral vectors into plant cells via agroinfiltration. | Strain GV3101 is commonly used for its high transformation efficiency [4]. |
| Acetosyringone | A phenolic compound that induces the Agrobacterium virulence system, crucial for T-DNA transfer. | Typically used at 200 µM during the final bacterial culture step before infiltration [4]. |
| vsRNAi Oligonucleotides | Short, synthetic DNA sequences that, when cloned into the VIGS vector, produce precise siRNAs. | 32-nt inserts offer a simplified, specific, and cost-effective alternative to long fragments [89]. |
| Gu+‑siRNA Nanoparticles | Synthetic nanoparticle system for delivering siRNA; enables species-independent and long-distance silencing. | Gu+‑siRNA NPs protect siRNA from degradation and facilitate vascular movement, useful for topical application [46]. |
The interplay of speed, specificity, and transience positions siRNA-mediated VIGS as a uniquely powerful tool for initial gene function characterization. Its rapid, sequence-specific knockdown capability is ideal for screening and validating gene-phenotype associations in a high-throughput manner. However, researchers must be cognizant of its inherent constraints—the potential for off-target effects and the temporary nature of the silencing—which necessitate careful experimental design and complementary validation with stable genetic models like CRISPR-Cas9 for conclusive findings.
Future advancements are poised to extend the utility of siRNA in VIGS. The development of nanoparticle-based delivery systems (e.g., Gu+-siRNA NPs) promises to overcome limitations in systemic movement and efficiency, potentially enabling robust silencing in a wider range of species [46]. Furthermore, the integration of synthetic biology approaches, such as syn-tasiR-VIGS, which produces highly specific artificial small RNAs from viral vectors, will enhance specificity and multiplexing capabilities [35]. As these technologies mature, the role of siRNA in VIGS will continue to be pivotal, bridging the gap between genomic sequencing and functional annotation to accelerate discovery in both basic and applied research.
Virus-Induced Gene Silencing (VIGS) has established itself as a powerful reverse genetics tool for transiently knocking down targeted gene expression by harnessing the plant's endogenous RNA interference (RNAi) machinery [4] [5]. Traditionally viewed as a transient phenomenon for post-transcriptional gene silencing (PTGS), VIGS is now recognized for its capacity to induce heritable epigenetic modifications, creating a powerful hybrid approach for functional genomics and crop improvement [5]. This paradigm shift positions VIGS not merely as a tool for transient gene knockdown but as a technology capable of creating stable epigenetic variation.
The foundation of this hybrid approach lies in the intersection of VIGS with the RNA-directed DNA methylation (RdDM) pathway. When VIGS vectors are engineered to target promoter sequences rather than coding regions, they can trigger Transcriptional Gene Silencing (TGS) through DNA methylation, leading to stable, mitotically heritable epigenetic marks that can persist even after the viral vector is cleared from the plant [5]. This convergence creates a unique experimental system for dissecting the role of small interfering RNAs (siRNAs) in establishing and maintaining epigenetic states, while simultaneously offering a practical tool for crop enhancement without permanent DNA sequence alteration.
Molecular Mechanisms: From Cytoplasmic Silencing to Nuclear Epigenetics
Foundations of VIGS and siRNA Biogenesis
The classical VIGS mechanism initiates in the cytoplasm as a form of PTGS. The process begins when recombinant viral vectors introduce target gene sequences into plant cells. During viral replication, double-stranded RNA (dsRNA) intermediates are recognized by the host's RNAi machinery. Dicer-like (DCL) enzymes, primarily DCL2 and DCL4, process these dsRNAs into 21-24 nucleotide small interfering RNAs (siRNAs) [4] [13]. These siRNAs are then loaded into ARGONAUTE (AGO) proteins to form the RNA-induced silencing complex (RISC), which guides sequence-specific cleavage of complementary endogenous mRNAs [4] [5].
A critical feature enabling amplification and systemic spread of silencing is the role of RNA-dependent RNA polymerases (RDRs). RDR6 uses the cleaved mRNA fragments as templates to generate secondary dsRNAs, which are subsequently processed into secondary siRNAs through a process called transitivity [5] [13]. This amplification mechanism ensures robust and systemic silencing throughout the plant, making VIGS particularly effective despite its transient nature.
Transition to Epigenetic Modifications
The transition from transient silencing to stable epigenetic changes occurs when siRNAs generated through VIGS are recruited to the nucleus and directed to homologous DNA sequences. This process, known as RNA-directed DNA methylation (RdDM), represents the molecular bridge connecting cytoplasmic RNA silencing to nuclear epigenetics [5].
When VIGS constructs are designed to target promoter sequences rather than coding regions, the resulting siRNAs can guide de novo methylation of cytosines in all sequence contexts (CG, CHG, and CHH). This methylation is established by DOMAINS REARRANGED METHYLTRANSFERASE 2 (DRM2), the primary methyltransferase in the RdDM pathway [93]. The initial methylation can then be maintained through subsequent cell divisions: METHYLTRANSFERASE 1 (MET1) maintains CG methylation, CHROMOMETHYLTRANSFERASE 3 (CMT3) maintains CHG methylation, while CHH methylation requires continuous reinforcement by the RdDM pathway [5] [93].
Table 1: Key siRNA Classes and Their Functions in VIGS and Epigenetic Modification
| siRNA Class | Size (nt) | Generating DCL | Primary Function | Role in VIGS-Epigenetics |
|---|---|---|---|---|
| Primary siRNAs | 21-22 | DCL4, DCL2 | Initial target mRNA cleavage | Trigger sequence-specific silencing |
| Secondary siRNAs | 21-22 | DCL4, DCL2 | Amplify silencing signal | Enhance silencing strength and duration |
| hc-siRNAs | 24 | DCL3 | Transcriptional silencing | Establish DNA methylation via RdDM |
| tasiRNAs | 21 | DCL4 | Endogenous gene regulation | Potential tools for multiplexed targeting |
The following diagram illustrates the complete pathway from initial VIGS inoculation to stable epigenetic modification:
Experimental Protocols: Methodologies for VIGS-Induced Epigenetic Modifications
Vector Design for Epigenetic Targeting
The critical first step in VIGS-mediated epigenetic modification is the strategic design of viral vectors to target regulatory regions rather than coding sequences. Unlike conventional VIGS that targets exons, epigenetic VIGS requires constructs containing promoter-specific fragments of 100-300 base pairs with high density of cytosine residues, particularly in CG contexts, to facilitate efficient methylation establishment and maintenance [5].
Tobacco Rattle Virus (TRV)-based vectors are particularly suitable for this application due to their broad host range and efficient systemic movement [4]. The bipartite TRV system requires two plasmid constructs:
- TRV1: Encodes replicase and movement proteins
- TRV2: Contains the coat protein and cloning site for inserting target sequences
For epigenetic applications, the multiple cloning site of TRV2 is used to insert promoter fragments targeting endogenous genes. The selection of an appropriate promoter fragment is crucial and should be guided by bisulfite sequencing data to identify regions susceptible to methylation changes.
Table 2: Essential Research Reagents for VIGS-Epigenetics Experiments
| Reagent/Vector | Specifications | Function in Protocol | Example Applications |
|---|---|---|---|
| TRV-Based Vectors | Bipartite system (TRV1/TRV2) | Delivery of promoter-targeting sequences | Broad host range Solanaceae species |
| Promoter Inserts | 150-300 bp fragments with high C-content | Trigger RdDM at specific loci | FWA, MEDEA, transgene promoters |
| Agrobacterium Strains | GV3101, LBA4404 | Delivery of viral vectors via agroinfiltration | Transient transformation |
| syn-tasiRNA Precursors | Minimal 54-nt constructs | High-specificity multiplexed targeting | Avoiding off-target effects |
| Methylation Analysis Kits | Bisulfite conversion kits | Detection of DNA methylation changes | Confirming epigenetic modifications |
Plant Inoculation and Growth Conditions
The inoculation protocol follows established VIGS methodologies with specific modifications to enhance epigenetic outcomes:
Agrobacterium Preparation: Transform competent A. tumefaciens GV3101 with TRV1 and TRV2-derived plasmids containing promoter targets. Select positive colonies on appropriate antibiotics and culture overnight in YEP medium with antibiotics and 200 μM acetosyringone.
Agroinfiltration: Resuspend bacterial cultures in infiltration buffer (10 mM MgCl₂, 10 mM MES, 200 μM acetosyringone) to an OD₆₀₀ of 1.0-2.0. Mix TRV1 and TRV2 strains in 1:1 ratio and incubate for 3-4 hours at room temperature before infiltration.
Plant Material and Infiltration: Use 2-3 leaf stage seedlings for infiltration. Apply bacterial suspension to abaxial leaf surface using needleless syringe. For optimal epigenetic effects, maintain plants at 20-22°C post-infiltration, as lower temperatures enhance siRNA accumulation and silencing efficiency [4].
Generational Advancement: After phenotypic assessment in T₀ generation, advance plants to T₁ and subsequent generations while maintaining consistent environmental conditions to assess heritability of epigenetic marks.
Validation and Analysis Methods
Confirmation of successful epigenetic modification requires multiple validation approaches:
DNA Methylation Analysis: Perform bisulfite sequencing of the target promoter region to quantify cytosine methylation in CG, CHG, and CHH contexts. Compare methylation patterns between VIGS-treated plants and controls.
Gene Expression Analysis: Use RT-qPCR to measure transcript levels of the target gene. Successful transcriptional silencing should correlate with promoter methylation.
siRNA Detection: Conduct northern blotting or sRNA sequencing to verify production of promoter-targeting siRNAs.
Phenotypic Assessment: Document visible phenotypes throughout plant development and across generations. For the well-characterized FWA gene, this involves monitoring delayed flowering as a readout of successful epigenetic silencing [5].
Crossing Schemes: Outcross silenced plants with wild-type counterparts to assess meiotic stability of epigenetic states and distinguish between RNA-independent and RNA-dependent maintenance mechanisms.
Applications and Case Studies: From Functional Genomics to Crop Improvement
Model System: Epigenetic Silencing of FWA
The most compelling demonstration of VIGS-induced epigenetic modification comes from studies on the Arabidopsis FWA gene. Normally expressed only in the endosperm, ectopic expression of FWA in vegetative tissues causes a delayed flowering phenotype [5]. Researchers engineered TRV vectors containing a segment of the FWA promoter (TRV:FWAtr) and inoculated wild-type Arabidopsis plants.
The results demonstrated that VIGS targeting could establish de novo methylation in the FWA promoter, resulting in heritable transcriptional silencing that persisted for multiple generations even in the absence of the viral vector [5]. This epigenetic silencing correlated with the expected delayed flowering phenotype, confirming functional gene suppression. Molecular analysis revealed that this process required both Pol IV and Pol V, key components of the canonical RdDM pathway, highlighting the mechanistic overlap between VIGS-induced and endogenous epigenetic pathways.
Applications in Crop Species
The VIGS-epigenetics approach has significant potential for crop improvement, particularly for species with complex genomes or challenging transformation systems:
In pepper (Capsicum annuum L.), where stable transformation remains difficult, VIGS has been successfully employed to study genes involved in fruit quality, pathogen resistance, and abiotic stress tolerance [4]. The extension to epigenetic modifications opens possibilities for creating stable epialleles of agronomically important genes without permanent genetic alteration.
For tomato and potato, VIGS-epigenetics could potentially create stable silencing of susceptibility genes required for pathogen infection, providing durable resistance to bacterial, oomycete, and fungal pathogens [4].
In cereals including rice and maize, where heterosis is commercially important, VIGS-epigenetics approaches could help elucidate how epigenetic modifications contribute to hybrid vigor, potentially leading to strategies for fixing hybrid performance in pure breeding lines [93].
Current Challenges and Future Perspectives
Technical Limitations and Optimization Strategies
Despite its promise, the VIGS-epigenetics approach faces several technical challenges:
Silencing Efficiency and Stability: The efficiency of establishing heritable epigenetic marks varies significantly between target loci and plant species. Current research focuses on optimizing vector systems, including the development of satellite virus-based systems and vectors employing viral suppressors of RNA silencing (VSRs) like P19 and HC-Pro to enhance silencing strength [4].
Species-Specific Limitations: VIGS efficiency is influenced by host factors including Argonaute protein specificity and siRNA movement [4]. Adaptation to monocot species requires specialized vectors such as those based on Barley Stripe Mosaic Virus (BSMV).
Environmental Sensitivity: VIGS efficiency is strongly influenced by environmental conditions including temperature, humidity, and photoperiod [4]. Standardization of growth conditions is essential for reproducible epigenetic outcomes.
Emerging Technologies and Future Directions
Several emerging technologies are poised to enhance the capabilities of VIGS-epigenetics approaches:
syn-tasiRNA Technology: Recent advances in synthetic trans-acting siRNAs enable highly specific multiplexed gene targeting with minimal off-target effects [24]. The development of 54-nt minimal precursors allows expression from size-constrained viral vectors, facilitating non-transgenic delivery.
VIGS-CRISPR Integration: Combining VIGS with CRISPR-based technologies offers exciting possibilities. CRISPR-dCas9 fusion with methyltransferases can target methylation to specific loci, while VIGS can serve as a delivery system for CRISPR components or as a validation tool for candidate genes identified through CRISPR screens.
High-Throughput Epigenetic Screening: The adaptation of VIGS-epigenetics to multi-well formats enables genome-wide screening for phenotypic consequences of epigenetic modifications, potentially identifying novel epialleles with agricultural value.
Multi-Omics Integration: Combining VIGS-epigenetics with transcriptomic, methylomic, and small RNA profiling will provide systems-level understanding of epigenetic regulation networks and their applications in crop improvement [4].
As these technologies mature, VIGS-mediated epigenetic modification is positioned to become an increasingly powerful tool for both basic research and agricultural biotechnology, providing a non-transgenic approach to creating stable phenotypic variation through directed epigenetic change.
The discovery of RNA interference (RNAi) has created a transformative bridge between basic plant science and human therapeutics. Virus-Induced Gene Silencing (VIGS), a technique grounded in the plant antiviral response mechanism of post-transcriptional gene silencing (PTGS), has served as an indispensable reverse genetics tool for functional genomics in plants [5] [4]. This natural cellular defense system shares a fundamental core mechanism with therapeutic RNAi: both utilize small interfering RNAs (siRNAs) to guide the sequence-specific degradation of complementary messenger RNA (mRNA) targets. The transition of this paradigm from a functional genomics tool to an approved drug class represents one of the most significant advancements in modern pharmacotherapy, enabling the precise silencing of disease-causing genes that were previously considered "undruggable" [71] [94].
The clinical success of siRNA therapeutics validates the molecular principles first observed in VIGS and other RNAi systems. Since the initial approval of patisiran in 2018, the field has witnessed rapid expansion with multiple additional approvals, demonstrating that siRNA technology can successfully address unmet medical needs across metabolic, cardiovascular, and genetic disorders [71] [95]. This whitepaper analyzes the key lessons learned from approved siRNA therapeutics, focusing on the technical innovations in delivery, chemical modification, and clinical development that were essential for their translation from research concepts to medicines, while framing these advances within the broader context of VIGS research principles.
The siRNA Therapeutic Landscape: Approved Agents and Targets
The clinical translation of siRNA therapeutics has culminated in several approved agents that demonstrate the technology's versatility. As of 2024, regulatory agencies worldwide have approved six siRNA drugs, with the majority focusing on hepatic targets and rare genetic diseases [95]. These approved drugs have established a strong foundation for the platform, validating key delivery approaches and chemical modification strategies.
Table 1: Approved siRNA Therapeutics and Their Key Characteristics
| Drug (Brand Name) | Target | Indication | Delivery System | Key Innovation |
|---|---|---|---|---|
| Patisiran (Onpattro) | Transthyretin (TTR) | Hereditary transthyretin-mediated amyloidosis | Lipid Nanoparticles (LNPs) | First-approved siRNA therapeutic; established LNP delivery platform [71] |
| Givosiran (Givlaari) | Aminolevulinic acid synthase 1 (ALAS1) | Acute hepatic porphyria | GalNAc conjugation | Demonstrated efficient hepatocyte targeting with enhanced therapeutic index [71] [95] |
| Lumasiran (Oxlumo) | Hydroxyacid oxidase 1 (HAO1) | Primary hyperoxaluria type 1 | GalNAc conjugation | Validated substrate reduction therapy for metabolic disorder [71] [95] |
| Inclisiran (Leqvio) | Proprotein convertase subtilisin/kexin type 9 (PCSK9) | Hypercholesterolemia | GalNAc conjugation | First siRNA for cardiovascular disease with durable effect (dosing every 6 months) [71] [95] |
| Vutrisiran (Amvuttra) | Transthyretin (TTR) | Transthyretin-mediated amyloidosis | GalNAc conjugation | Enhanced stability allowing for subcutaneous quarterly dosing [95] |
| Nedosiran (Rivfloza) | Lactate dehydrogenase A (LDHA) | Primary hyperoxaluria type 1 | GalNAc conjugation | Multi-enzyme targeting for comprehensive oxalate reduction [71] [95] |
The dominance of GalNAc-conjugated siRNAs in recent approvals highlights a key lesson in clinical translation: tissue-specific targeting is paramount for success. The GalNAc moiety binds selectively to the asialoglycoprotein receptor (ASGPR) abundantly expressed on hepatocyte surfaces, facilitating efficient cellular uptake and enabling robust gene silencing at low doses [94]. This targeted approach minimizes systemic exposure and off-target effects, addressing one of the major challenges in oligonucleotide therapeutics.
Molecular Mechanisms: From VIGS to Therapeutic RNAi
The fundamental mechanism of therapeutic RNAi shares conserved features with the Virus-Induced Gene Silencing (VIGS) pathway observed in plants. Both processes utilize small RNA fragments to guide the sequence-specific silencing of complementary gene transcripts through the RNA-induced silencing complex (RISC).
Comparative Mechanisms of VIGS and Therapeutic RNAi
In VIGS, plant infection with recombinant viruses leads to the production of double-stranded RNA (dsRNA) replication intermediates, which are recognized by the plant's antiviral defense system [5] [4]. Dicer-like enzymes (DCL) process these dsRNAs into 21-24 nucleotide small interfering RNAs (siRNAs) that are loaded into RISC. The complex uses the siRNA guide strand to identify and cleave complementary viral mRNA, preventing infection [4].
Therapeutic RNAi follows an analogous pathway but uses exogenously administered synthetic siRNA duplexes. These 21-23 nucleotide duplexes with 3'-dinucleotide overhangs are introduced into the cytosol, where they are recognized and loaded into the multiprotein RNA-induced silencing complex (RISC) [71]. The Argonaute 2 (AGO2) endonuclease selectively retains the guide strand while ejecting the passenger strand. The mature siRNA-RISC complex then engages complementary target mRNA transcripts via Watson-Crick base pairing, predominantly within the seed region (nucleotides 2-8). AGO2 catalyzes site-specific endonucleolytic cleavage of the mRNA between positions corresponding to 10 and 11 from the 5' end of the guide strand, disrupting mRNA stability and abrogating translation [71].
Diagram 1: Therapeutic siRNA Mechanism. The process involves RISC loading, mRNA binding, and AGO2-mediated cleavage leading to gene silencing.
Technical Innovations Enabling Clinical Success
The transition from RNAi concept to approved therapeutics required overcoming substantial technical challenges. Unmodified siRNAs suffer from rapid degradation by nucleases, inefficient cellular uptake due to their negative charge, and potential immunogenicity [71]. The solutions to these problems represent critical lessons for the field.
Chemical Modifications for Enhanced Stability and Specificity
Chemical modifications have been instrumental in transforming siRNAs from research tools to viable therapeutics. These modifications enhance nuclease resistance, improve binding affinity, reduce immunogenicity, and fine-tune pharmacokinetic properties while preserving RNAi activity [71].
Table 2: Key Chemical Modifications in Therapeutic siRNAs
| Modification Type | Examples | Functional Impact |
|---|---|---|
| Sugar (Ribose) Modifications | 2'-O-methyl (2'-OMe), 2'-fluoro (2'-F), 2'-O-methoxyethyl (2'-MOE) | Enhance nuclease resistance, improve binding affinity, reduce immune stimulation [71] |
| Phosphate Backbone Modifications | Phosphorothioate (PS), Phosphorodithioate | Increase resistance to nuclease degradation, improve pharmacokinetic properties [71] |
| Base Modifications | 5-methylcytosine, Pseudouridine | Reduce immune activation, enhance stability while maintaining base pairing [71] |
| Terminal & Conjugate Modifications | GalNAc conjugation, Cholesterol addition | Enable tissue-specific targeting, improve cellular uptake and biodistribution [71] [94] |
Strategic incorporation of these modifications follows pattern-based approaches, with heavy modification of the passenger strand versus minimal modification of the guide strand to bias RISC loading toward the functional strand [71]. Position-specific alterations at critical regions for RISC loading and activity further refine specificity and potency.
Advanced Delivery Systems
Effective delivery represents perhaps the most significant hurdle overcome in siRNA therapeutic development. Without specialized delivery systems, siRNAs cannot reach their intracellular sites of action in therapeutic concentrations [71].
Lipid-Based Nanoparticles (LNPs): Patisiran's success established LNPs as a viable delivery platform for siRNA therapeutics. These systems typically combine ionizable lipids, helper lipids, PEGylated lipids, and cholesterol to form stable particles that protect siRNAs from degradation, facilitate cellular uptake through endocytosis, and enable endosomal escape [71]. The ionizable lipids are particularly crucial as they become positively charged in the acidic endosomal environment, promoting disruption of the endosomal membrane and release of siRNAs into the cytosol.
GalNAc Conjugation: The development of N-acetylgalactosamine (GalNAc) conjugates represents a breakthrough in targeted siRNA delivery. This approach exploits the high-affinity binding of GalNAc to the asialoglycoprotein receptor (ASGPR) abundantly expressed on hepatocytes [94]. The GalNAc-siRNA conjugates are internalized via receptor-mediated endocytosis, achieving highly efficient liver-specific delivery with significantly reduced dosing requirements compared to non-targeted approaches. The success of this platform is evident in its adoption for multiple approved therapeutics, including givosiran, lumasiran, and inclisiran [94] [95].
Diagram 2: siRNA Delivery Platform Comparison. Both LNP and GalNAc platforms must overcome endosomal trapping to achieve cytosolic release.
The Scientist's Toolkit: Essential Reagents and Methodologies
The development of approved siRNA therapeutics has relied on specialized reagents and methodologies that address the unique challenges of oligonucleotide-based drugs.
Table 3: Essential Research Reagents and Platforms for siRNA Therapeutic Development
| Reagent/Platform | Function | Application in Therapeutic Development |
|---|---|---|
| Ionizable Lipids | Enable endosomal escape through pH-dependent structural changes | Core component of LNP delivery systems (e.g., in patisiran) [71] |
| GalNAc Conjugation Chemistry | Facilitates hepatocyte-specific targeting via ASGPR binding | Platform technology for multiple approved siRNAs (givosiran, inclisiran, etc.) [94] |
| Phosphorothioate Linkages | Increase nuclease resistance and plasma protein binding | Backbone modification to improve pharmacokinetic properties [71] |
| 2'-Sugar Modifications | Enhance binding affinity and reduce immune recognition | Ribose modifications (2'-OMe, 2'-F) to improve stability and safety [71] |
| LC-MS/MS Bioanalytical Platforms | Quantify siRNA concentrations in biological matrices | Essential for pharmacokinetic studies; achieved sensitivity to 1-10 ng/mL [94] |
Clinical Development and Regulatory Considerations
The clinical development path for siRNA therapeutics has revealed distinct considerations compared to traditional small molecules. The unique properties of oligonucleotides necessitate specialized approaches to pharmacokinetics, pharmacodynamics, and safety assessment.
Bioanalytical Methodologies
Robust bioanalytical methods are essential for understanding the absorption, distribution, metabolism, and excretion (ADME) properties of siRNA therapeutics. The high molecular weight, negative charge, and potential for nonspecific binding of oligonucleotides present unique challenges [94]. Three major assay platforms have been employed for approved siRNAs:
- Chromatography-based assays (LC-MS/MS): Provide high specificity and ability to differentiate parent drug from metabolites, with sensitivity achieving sub-ng/mL levels for recent approvals [94]
- Ligand-binding assays (LBA): Offer higher sensitivity but lower specificity than LC-MS methods
- PCR-based assays: Deliver high sensitivity and throughput but may lack specificity for intact siRNA
The selection of appropriate bioanalytical methods requires careful consideration of the specific siRNA chemistry, anticipated concentrations, and needed specificity [94].
Clinical Trial Landscape and Trends
Analysis of global clinical trials reveals distinctive patterns between oncology and non-oncology applications of siRNA therapeutics. Non-oncology trials dominate the landscape (90% of 424 trials between 2004-2024), having yielded all six currently approved siRNA drugs for metabolic and genetic diseases [95]. Key non-oncology targets include PCSK9 for hypercholesterolemia and genes associated with hepatitis B infection.
In contrast, oncology applications have progressed more slowly, with trials primarily in Phase I/II stages (60% Phase I) and focusing on solid tumors (40%) and targets like CSF2 [95]. The higher complexity of tumor delivery, heterogeneous target expression, and challenges in penetrating tumor microenvironments have complicated oncology development. However, identified shared targets across multiple tumors (PTGS2, TGFB1) suggest potential for combination approaches [95].
The successful clinical translation of siRNA therapeutics offers compelling lessons for the field. First, solving the delivery challenge through either LNP or targeted conjugation approaches was absolutely essential for therapeutic efficacy. Second, strategic chemical modifications dramatically improved the drug-like properties of siRNAs, enhancing stability while reducing immunogenicity. Third, the tissue-specific approach exemplified by GalNAc conjugation enables favorable therapeutic indices through targeted delivery.
These advances establish siRNA therapeutics as a precise modality that expands the druggable genome. According to Pharmaproject, there are over 260 siRNA drug candidates in preclinical or clinical development, indicating sustained growth in the field [71]. The continuing innovation in delivery technologies, including antibody-oligonucleotide conjugates and novel nanoparticle formulations, promises to expand siRNA applications beyond hepatic targets to additional tissues and disease states.
For researchers working at the intersection of VIGS and therapeutic development, the clinical validation of siRNA technology reinforces the fundamental principles of RNAi while demonstrating the additional sophistication required for human therapeutics. The lessons from approved siRNA therapeutics provide a roadmap for translating basic RNAi research into clinically viable treatments, highlighting the critical importance of delivery, stability, and specificity in achieving successful clinical outcomes.
The field of personalized medicine is undergoing a transformative shift, driven by technological advances in high-throughput screening (HTS) platforms. These platforms enable the rapid testing of thousands of chemical compounds or biological agents against specific disease targets, accelerating the discovery of potential therapies [96]. Contemporary drug discovery is a complex, multidisciplinary process characterized by a high screening attrition rate, where only one viable drug may emerge from millions of screened compounds [96]. This inefficiency has identified the need to screen larger libraries, positioning HTS—which can investigate hundreds of thousands of compounds per day—as a cornerstone of modern discovery pipelines [96]. The growing demand for personalized therapeutic interventions has further propelled innovations in HTS, making these platforms indispensable for identifying patient-specific treatments.
Concurrently, the development of sophisticated molecular tools like small interfering RNA (siRNA) has revolutionized functional genomics and drug target validation. siRNA operates within the RNA interference (RNAi) pathway, a sequence-specific biological mechanism for gene silencing. In plants, this mechanism is harnessed through Virus-Induced Gene Silencing (VIGS), a technique that uses recombinant viral vectors to deliver siRNA precursors, leading to the targeted knockdown of endogenous genes [5]. The principles of siRNA-mediated silencing form a critical technological bridge, connecting basic functional genomics in plants with advanced, targeted therapeutic discovery in humans. This article explores the synergistic future of HTS platforms and siRNA technologies in advancing personalized medicine.
Fundamental Principles of siRNA and VIGS
The Molecular Machinery of siRNA-Mediated Silencing
Virus-Induced Gene Silencing is a powerful reverse genetics technology that exploits the plant's innate post-transcriptional gene silencing (PTGS) antiviral defense system [5]. The process is initiated by the delivery of a recombinant virus carrying a fragment of the host plant's target gene. The molecular mechanism can be summarized in a series of key steps:
- Viral Replication and dsRNA Formation: Following infection, the recombinant virus replicates in the host plant. During replication, double-stranded RNA (dsRNA) molecules—a common viral intermediate—are generated [4].
- Dicer Cleavage and siRNA Biogenesis: The host plant's Dicer-like (DCL) enzymes recognize and cleave the long dsRNA molecules into 21-24 nucleotide small interfering RNA (siRNA) duplexes [5].
- RISC Loading and Target Cleavage: These siRNAs are incorporated into an RNA-Induced Silencing Complex (RISC). The guide strand of the siRNA directs RISC to complementary messenger RNA (mRNA) sequences. The Argonaute (AGO) protein, a catalytic component of RISC, then cleaves the target mRNA, preventing its translation into protein [5].
This pathway is illustrated in the following diagram:
From Transcriptional to Epigenetic Silencing
A significant advancement in VIGS research is its extension beyond post-transcriptional regulation to induce heritable epigenetic modifications. When the viral vector insert is designed to target promoter sequences rather than coding sequences, the siRNA can guide the RNA-directed DNA methylation (RdDM) pathway [5]. In this pathway, a specialized RNA polymerase, Pol V, produces non-coding scaffold RNAs at the target locus. The AGO-siRNA complex binds to these scaffolds, recruiting DNA methyltransferases that deposit methyl groups onto cytosine residues in the promoter region [5]. This methylation leads to transcriptional gene silencing (TGS), which can be stable and inherited over subsequent generations, creating new, stable phenotypes without altering the underlying DNA sequence [5]. This application opens up possibilities for epigenetic breeding and the functional study of heritable gene regulation.
High-Throughput Screening Platforms in Drug Discovery
Core Technologies and Assay Formats
HTS encompasses a range of in vitro, cell-based, or whole-organism-based assays designed to identify "hits"—compounds that produce a desired biological effect [96]. The choice of assay format is critical and depends on the biological question being addressed.
- Biochemical Assays: These are typically used when a specific, purified target (e.g., an enzyme or receptor) is known. The most common readouts are optical, including absorbance, fluorescence, luminescence, and scintillation [96].
- Cell-Based Assays: These are indispensable for studying complex biological processes, pathways, and phenotypic changes within a cellular context. They offer higher biological relevance by accounting for cell permeability, cytotoxicity, and effects on entire signaling networks [96]. A major innovation in this area is the shift from traditional 2D cell monolayers to 3D cell cultures and organoids, which better mimic the in vivo architecture and function of human tissues, providing more predictive data for clinical outcomes [97].
The workflow for a typical HTS campaign in drug discovery, from target identification to lead optimization, is shown below:
Quantitative Data from HTS Implementations
HTS campaigns generate vast amounts of quantitative data. The table below summarizes key metrics and outcomes from recent implementations as reported in the literature.
Table 1: Performance Metrics of Select High-Throughput Screening Platforms
| Platform / Study | Library Size | Screening Throughput | Key Outcome / Hit Rate | Application Context |
|---|---|---|---|---|
| IRB Barcelona Screening Platform [98] | 172,505 compounds | Large-scale, automated | Supports multiple collaborative projects | Personalized medicine, neurodegenerative diseases, cancer |
| Personalized Phage Therapy [99] | Libraries of hundreds of phages | Target identification in 30-120 minutes | Phage-specific lysis detected via ATP release | Rapid identification of therapeutic bacteriophages for antibiotic-resistant infections |
| Standard HTS Campaign [96] | Millions of compounds | ~100,000 compounds per day | One drug from millions screened; hundreds of "hits" may be identified | General drug discovery for neurodegenerative diseases (NDDs) |
Integrated Experimental Protocols
Protocol: A TRV-Based VIGS System in Soybean
This protocol details an efficient Tobacco Rattle Virus (TRV)-based VIGS system for functional gene validation in soybean, adapted from a recent study [100].
1. Vector Construction:
- Primer Design: Design gene-specific primers with added restriction enzyme sites (e.g., EcoRI and XhoI) for directional cloning.
- PCR Amplification: Amplify a 200-300 bp fragment of the target gene (e.g., GmPDS) from soybean cDNA using high-fidelity DNA polymerase.
- Cloning: Ligate the purified PCR product into the corresponding sites of the pTRV2 vector. Transform the ligation product into E. coli DH5α competent cells and select positive clones for sequence verification [100].
2. Agrobacterium Preparation:
- Transform the confirmed recombinant pTRV2 plasmid and the helper pTRV1 plasmid into Agrobacterium tumefaciens strain GV3101.
- Culture a single colony in YEB medium supplemented with appropriate antibiotics (e.g., kanamycin, rifampicin) and 10 mM MES buffer.
- Induce the Agrobacterium culture by adding acetosyringone to a final concentration of 200 μM. Incubate with shaking until the OD₆₀₀ reaches 0.9-1.0 [100].
3. Plant Inoculation via Cotyledon Node Immersion:
- This optimized method overcomes the challenges of soybean's thick cuticle and dense trichomes.
- Surface-sterilize soybean seeds and soak them in sterile water until swollen.
- Bisect the seeds longitudinally to create half-seed explants.
- Immerse the fresh explants in the mixed Agrobacterium suspension (containing both pTRV1 and the recombinant pTRV2) for 20-30 minutes with gentle agitation [100].
4. Post-Inoculation and Analysis:
- Co-cultivate the inoculated explants on sterile medium in the dark for 2-3 days.
- Transfer the plants to soil and maintain under controlled environmental conditions (e.g., 22°C, 16/8 hour light/dark cycle).
- Monitor for silencing phenotypes (e.g., photobleaching for GmPDS) from 14-21 days post-inoculation (dpi).
- Quantify silencing efficiency through molecular analysis (qRT-PCR) of target gene expression.
Essential Research Reagents for VIGS
The successful implementation of a VIGS experiment relies on a suite of specialized reagents and vectors.
Table 2: Key Research Reagent Solutions for VIGS Experiments
| Reagent / Vector | Function and Description | Specific Example |
|---|---|---|
| TRV Viral Vector System | A bipartite vector system; TRV1 encodes replication and movement proteins, while TRV2 carries the target gene insert for silencing. | pTRV1, pTRV2 [100] [4] |
| Agrobacterium tumefaciens | A bacterial strain used as a delivery vehicle to introduce the TRV vectors into plant cells. | Strain GV3101 [100] |
| Induction Media Supplements | Chemical compounds added to the Agrobacterium culture to activate its virulence system for efficient gene transfer. | Acetosyringone (200 μM), MES buffer (10 mM) [100] |
| Marker Gene | A visual reporter used to monitor infection efficiency and silencing spread. | Phytoene Desaturase (PDS); silencing causes photobleaching [100] [4] |
| Viral Suppressor of RNAi (VSR) | A protein co-expressed to temporarily inhibit the plant's silencing machinery, enhancing VIGS efficiency. | P19, HC-Pro [4] |
Future Outlook and Integrated Applications
The convergence of HTS and siRNA-based technologies like VIGS is paving the way for a new era in biomedical research and personalized therapy. Several key directions are emerging:
- High-Throughput Functional Genomics: VIGS is evolving into a high-throughput tool itself, enabling the large-scale functional screening of candidate genes across entire genomes or specific pathways, thus rapidly prioritizing targets for further investigation [5] [4].
- Epigenetic Engineering and Breeding: The ability of VIGS to induce heritable epigenetic modifications offers a novel, non-transgenic strategy for crop improvement. This approach can create stable genotypes with desired traits, such as enhanced stress resistance, by modulating gene expression patterns [5].
- Advanced Personalized Screening Platforms: The concept of personalized HTS is expanding beyond small molecules. For instance, platform technologies that use solid tablets to encapsulate individual bacteriophages alongside detection biochemistry allow for the rapid, high-throughput matching of therapeutic phages to a patient's specific antibiotic-resistant bacterial isolate within hours [99].
- Integration with Multi-Omics and CRISPR: The future lies in combining the high-throughput capabilities of VIGS and HTS with other technologies. Integrating VIGS with multi-omics data (transcriptomics, proteomics) provides a systems-level understanding of gene function [4]. Furthermore, coupling siRNA screens with CRISPR-dCas9 systems allows for seamless transition from transcriptional to epigenetic silencing studies, creating a powerful toolkit for controlling gene expression [5].
In conclusion, the synergistic application of high-throughput screening platforms and siRNA-mediated silencing technologies is fundamentally advancing the capabilities of personalized medicine. By enabling the rapid identification of therapeutic targets and the validation of gene function on an unprecedented scale, these tools are accelerating the development of precise, effective, and patient-tailored treatments for a wide spectrum of diseases.
Conclusion
The integration of siRNA mechanisms within VIGS technology represents a powerful paradigm for functional genomics and therapeutic development. Through understanding the molecular foundations, methodological applications, optimization strategies, and validation frameworks, researchers can harness this technology with increasing precision and efficiency. The future of VIGS lies in its convergence with multi-omics technologies, advanced delivery systems, and computational design tools, potentially enabling not only accelerated crop improvement but also novel therapeutic interventions. As delivery challenges are addressed and specificity enhanced, VIGS and siRNA-based approaches are poised to expand beyond current limitations, offering unprecedented opportunities for targeted gene manipulation in both agricultural and biomedical contexts. The continued refinement of this technology promises to unlock new dimensions in personalized medicine and sustainable agriculture through precise, sequence-specific gene regulation.
References
- 1. RNA Interference (RNAi) - NCBI - NIH [https://www.ncbi.nlm.nih.gov/probe/docs/techrnai/]
- 2. RNA interference [https://en.wikipedia.org/wiki/RNA_interference]
- 3. RNA Silencing in Plants: Mechanisms, Technologies and ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC5108043/]
- 4. Virus-Induced Gene Silencing (VIGS) in Functional Genomics [https://www.mdpi.com/2311-7524/11/11/1297]
- 5. Virus-Induced Gene Silencing (VIGS): A Powerful Tool for ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10057534/]
- 6. Virus-Based MicroRNA Expression for Gene Functional ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC2879806/]
- 7. Syn-tasiR-VIGS: virus-based targeted RNAi in plants by ... [https://pubmed.ncbi.nlm.nih.gov/40105245/]
- 8. Advancing virus-induced gene silencing in sunflower [https://plantmethods.biomedcentral.com/articles/10.1186/s13007-024-01241-z]
- 9. Spray-induced gene silencing for crop protection [https://www.frontiersin.org/journals/plant-science/articles/10.3389/fpls.2025.1527944/full]
- 10. Molecular Mechanisms and Biological Functions of siRNA [https://pmc.ncbi.nlm.nih.gov/articles/PMC5542916/]
- 11. RNAi vs. CRISPR: Guide to Selecting the Best Gene ... [https://www.synthego.com/blog/rnai-vs-crispr-guide/]
- 12. Structural Foundations of RNA Silencing by Argonaute - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC5576611/]
- 13. RNA interference and turnover in plants -a complex ... [https://www.frontiersin.org/journals/plant-science/articles/10.3389/fpls.2025.1608888/full]
- 14. The Argonaute protein family | Genome Biology | Full Text [https://genomebiology.biomedcentral.com/articles/10.1186/gb-2008-9-2-210]
- 15. RNA-induced silencing complex [https://en.wikipedia.org/wiki/RNA-induced_silencing_complex]
- 16. Review RISC-target interaction: Cleavage and translational ... [https://www.sciencedirect.com/science/article/abs/pii/S1874939908001521]
- 17. Molecular Bases of Viral RNA Targeting by Viral Small ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC1866121/]
- 18. Fragile X-related protein and VIG associate with the RNA ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC187452/]
- 19. siRNA 疗法的新突破拓展了药物管线 [https://www.cas.org/zh-hans/resources/cas-insights/sirna-therapeutics]
- 20. RNA interference and turnover in plants -a complex ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12259633/]
- 21. Exogenous dsRNA-Mediated RNAi: Mechanisms ... [https://www.mdpi.com/1999-4915/17/1/49]
- 22. Prediction of VIGS efficiency by the Sfold program and its ... [https://www.sciencedirect.com/science/article/abs/pii/S2095927316301530]
- 23. Antiviral RNA interference in plants: Increasing complexity ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12546453/]
- 24. Effective Gene Silencing in Plants by Synthetic Trans-Acting ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12550521/]
- 25. Enhancing dsRNA/siRNA delivery to combat agricultural ... [https://www.sciencedirect.com/science/article/abs/pii/S0141813024078097]
- 26. Single-cell encoded gene silencing for high-throughput ... [https://www.nature.com/articles/s41467-024-53419-7]
- 27. siRNA vs miRNA: decoding these tiny but mighty molecules [https://www.thermofisher.com/blog/life-in-the-lab/sirna-vs-mirna-decoding-these-tiny-but-mighty-molecules/]
- 28. Differences and relationships between siRNA and miRNA ... [https://www.bocsci.com/resources/differences-and-relationships-between-sirna-and-mirna-in-synthesis-and-function.html?srsltid=AfmBOopVMMm20K9k-KL0aoa4Lzu8PZqGndimuA3ZKdkTkdlZNDrBYa0G]
- 29. A review of MicroRNAs and flavonoids: New insights into ... [https://www.sciencedirect.com/science/article/abs/pii/S0141813025030703]
- 30. Design of therapeutic siRNAs for potential application to ... [https://www.sciencedirect.com/science/article/pii/S240584402500204X]
- 31. Recent advances in miRNA and siRNA approaches ... [https://www.sciencedirect.com/science/article/abs/pii/S0141813025077529]
- 32. siRNAs can function as miRNAs - PMC - PubMed Central - NIH [https://pmc.ncbi.nlm.nih.gov/articles/PMC195999/]
- 33. Therapeutic Approaches of Viral Gene Silencing by Small ... [https://www.mdpi.com/1424-8247/18/7/987]
- 34. Characterizing the role of RSD-6 in the biogenesis of virus ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12424992/]
- 35. Syn-tasiR-VIGS: virus-based targeted RNAi in plants by ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11920798/]
- 36. A rapid, simple, and highly efficient method for VIGS and in ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC8605592/]
- 37. Virus-Induced Gene Silencing (VIGS): A Powerful Tool for ... [https://www.mdpi.com/1422-0067/24/6/5608]
- 38. Virus-induced gene silencing is a versatile tool for ... [https://www.frontiersin.org/journals/plant-science/articles/10.3389/fpls.2014.00323/full]
- 39. Developing and applying a virus-induced gene silencing ... [https://www.sciencedirect.com/science/article/abs/pii/S0378111924009685]
- 40. Establishment of a tobacco rattle virus-induced ... [https://www.sciencedirect.com/science/article/abs/pii/S0304423824008616]
- 41. based virus-induced gene Silencing in Atriplex canescens [https://pmc.ncbi.nlm.nih.gov/articles/PMC12330136/]
- 42. Application and Expansion of Virus-Induced Gene ... [https://www.mdpi.com/2311-7524/9/8/934]
- 43. Virus-Induced Genome Editing (VIGE): One Step Away ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12111327/]
- 44. Agrobacterium-mediated transformation for virus-induced ... [https://pubmed.ncbi.nlm.nih.gov/40730887/]
- 45. An Improved Agrobacterium-Mediated Transformation ... [https://www.mdpi.com/2223-7747/14/15/2353]
- 46. Enabling long‐distance gene silencing in plants via ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11933838/]
- 47. An Efficient Agrobacterium-Mediated Transient ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12388925/]
- 48. Truncated CMV2bN43 enhances virus-induced gene ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12542452/]
- 49. Resistance and tolerance to viroid infection [https://link.springer.com/article/10.1007/s10327-025-01264-x]
- 50. Virus-induced Gene Silencing (VIGS) in Nicotiana ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC2795700/]
- 51. Nanotechnology-driven gene silencing: advancements in ... [https://chembioagro.springeropen.com/articles/10.1186/s40538-025-00738-6]
- 52. Systematic analysis of siRNA and mRNA features impacting ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12205987/]
- 53. analysis of clinical trial registration data from 2004 to 2024 [https://www.frontiersin.org/journals/pharmacology/articles/10.3389/fphar.2025.1637958/full]
- 54. Selection of stable reference genes for accurate reverse- ... [https://www.nature.com/articles/s41598-025-08940-0]
- 55. Engineering siRNA therapeutics: challenges and strategies [https://jnanobiotechnology.biomedcentral.com/articles/10.1186/s12951-023-02147-z]
- 56. Computational Estimation and Experimental Verification of ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC1586062/]
- 57. siRNA potency enhancement via chemical modifications of ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC7812868/]
- 58. Designing & Screening of siRNA molecules for silencing ... [https://www.sciencedirect.com/science/article/abs/pii/S147692712500369X]
- 59. Improving the potency prediction for chemically modified ... [https://www.sciencedirect.com/science/article/pii/S2162253124003020]
- 60. Designing an effective therapeutic siRNA to silence RdRp ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC8170914/]
- 61. Therapeutic potential of chemically modified siRNA [https://pmc.ncbi.nlm.nih.gov/articles/PMC5935465/]
- 62. siRNAmod: A database of experimentally validated ... [https://www.nature.com/articles/srep20031]
- 63. Engineering VIGS Vectors by Modifying Movement ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11664229/]
- 64. Small-interfering RNAs from natural antisense transcripts ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC2603254/]
- 65. Methods for effective real-time RT-PCR analysis of virus- ... [https://www.sciencedirect.com/science/article/abs/pii/S0166093406002722]
- 66. An Optimized Protocol to Increase Virus-Induced Gene ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC3893464/]
- 67. Establishment and application of a root wounding ... [https://www.frontiersin.org/journals/plant-science/articles/10.3389/fpls.2024.1336726/full]
- 68. Construction of Simple and Efficient siRNA Validation ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC7121774/]
- 69. Vacuum and Co-cultivation Agroinfiltration of (Germinated) ... [https://www.frontiersin.org/journals/plant-science/articles/10.3389/fpls.2017.00393/full]
- 70. A novel siRNA validation system for functional screening and ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC7092908/]
- 71. New breakthroughs in siRNA therapeutics expand the drug... | CAS [https://www.cas.org/resources/cas-insights/sirna-therapeutics]
- 72. Therapeutic siRNA: state of the art [https://www.nature.com/articles/s41392-020-0207-x]
- 73. Five Ways to Modify Your siRNA for Improved RNAi [https://bitesizebio.com/47666/five-ways-to-modify-your-sirna-for-improved-rnai/]
- 74. Improving the potency prediction for chemically siRNAs... modified [https://pmc.ncbi.nlm.nih.gov/articles/PMC11960531/]
- 75. The new era of siRNA therapy: Advances in cancer treatment [https://pmc.ncbi.nlm.nih.gov/articles/PMC12148947/]
- 76. Mechanistic intracellular PK/PD modeling to inform ... [https://www.sciencedirect.com/science/article/pii/S2162253125000708]
- 77. RNA Interference Drug Delivery Market Size 2025 to 2034 [https://www.precedenceresearch.com/rna-interference-drug-delivery-market]
- 78. Frontiers | Development of siRNA therapeutics to combat microbial... [https://www.frontiersin.org/journals/cellular-and-infection-microbiology/articles/10.3389/fcimb.2025.1697880/full]
- 79. Real-Time PCR RNA Quantification Workflow [https://www.thermofisher.com/mx/en/home/life-science/pcr/real-time-pcr/real-time-pcr-applications/gene-expression-using-real-time-pcr/real-time-rna-quantification.html]
- 80. Brief guide to RT-qPCR - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC11612376/]
- 81. Comparison of the Phenotypic Performance, Molecular ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC9824520/]
- 82. Development of Virus-Induced Gene Silencing (VIGS) and ... [https://dineshkumar.ucdavis.edu/research/development-virus-induced-gene-silencing-vigs-and-crisprcas9-mediated-genome-editing-approaches]
- 83. Applications and Prospects of CRISPR/Cas9 Technology ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11644675/]
- 84. Efficient generation of stable, heritable gene edits in wheat ... [https://bmcplantbiol.biomedcentral.com/articles/10.1186/s12870-018-1433-z]
- 85. Development of virus-induced genome editing methods in ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10782499/]
- 86. Establishment of a CRISPR/Cas9 gene-editing system for ... [https://www.maxapress.com/article/doi/opr-0024-0012]
- 87. Establishment of a CRISPR/Cas9 gene editing system ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12379479/]
- 88. Topically delivered 22 nt siRNAs enhance RNAi silencing ... [https://link.springer.com/article/10.1007/s00425-021-03708-y]
- 89. Comparative genomics‐driven design of virus‐delivered ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12576428/]
- 90. siRNAs, saRNAs, miRNAs, CRISPR/Cas System [https://nhsjs.com/2025/revolutionizing-gene-editing-techniques-in-medicine-sirnas-sarnas-mirnas-crispr-cas-system/]
- 91. RNA-Targeting Techniques: A Comparative Analysis of ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12563179/]
- 92. Advances in RNA-based therapeutics: current breakthroughs ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12592931/]
- 93. Epigenetic Changes in Hybrids - PMC - PubMed Central - NIH [https://pmc.ncbi.nlm.nih.gov/articles/PMC4528738/]
- 94. Translational and clinical development of therapeutic siRNA ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12477600/]
- 95. Clinical development prospects of siRNA drugs for tumor ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12484142/]
- 96. High-Throughput Screening Platforms in the Discovery of ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC7926814/]
- 97. High-Throughput Screening Market: Transforming Drug ... [https://blog.bccresearch.com/high-throughput-screening-market-transforming-drug-discovery]
- 98. The Screening Platform at IRB Barcelona [https://www.irbbarcelona.org/en/news/institutional/screening-platform-irb-barcelona-key-advancing-personalized-medicine]
- 99. High throughput platform technology for rapid target ... [https://www.nature.com/articles/s41467-024-49710-2]
- 100. Efficient Virus-Induced Gene Silencing (VIGS) Method for ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12115190/]