Agrobacterium-Mediated VIGS: Advanced Infection Methods for Functional Gene Analysis
This article provides a comprehensive resource for researchers on Agrobacterium-mediated Virus-Induced Gene Silencing (VIGS), a powerful reverse genetics tool for rapid functional gene analysis.
Agrobacterium-Mediated VIGS: Advanced Infection Methods for Functional Gene Analysis
Abstract
This article provides a comprehensive resource for researchers on Agrobacterium-mediated Virus-Induced Gene Silencing (VIGS), a powerful reverse genetics tool for rapid functional gene analysis. Covering foundational principles to advanced applications, we detail optimized infection protocols for diverse plant species, including novel methods like root wounding-immersion and seed vacuum infiltration. The content addresses critical troubleshooting factors—genotype dependency, environmental conditions, and vector construction—that dictate silencing efficiency. With a focus on practical validation techniques and comparative method analysis, this guide empowers scientists to implement robust VIGS systems for high-throughput gene function studies in both model and non-model plant species, accelerating research in disease resistance, stress tolerance, and specialized metabolism.
Unlocking Gene Function: The Core Principles of VIGS Technology
Virus-Induced Gene Silencing (VIGS) has emerged as a powerful reverse genetics tool for rapid functional analysis of plant genes. This technology exploits an innate plant defense mechanism—Post-Transcriptional Gene Silencing (PTGS)—which naturally protects plants against viral pathogens. In VIGS, this protective system is co-opted to silence endogenous plant genes by using recombinant viral vectors that carry fragments of host target genes [1] [2]. The application of VIGS is particularly valuable in species where stable genetic transformation is challenging, time-consuming, or inefficient [3] [4]. Within the broader context of Agrobacterium-mediated VIGS infection methods research, understanding the PTGS mechanism is fundamental to optimizing silencing efficiency, developing new vectors, and adapting protocols for recalcitrant species. This article details the molecular basis of this mechanism and provides detailed protocols for its implementation in various plant systems.
The Core Mechanism: From Viral Defense to Functional Genomics
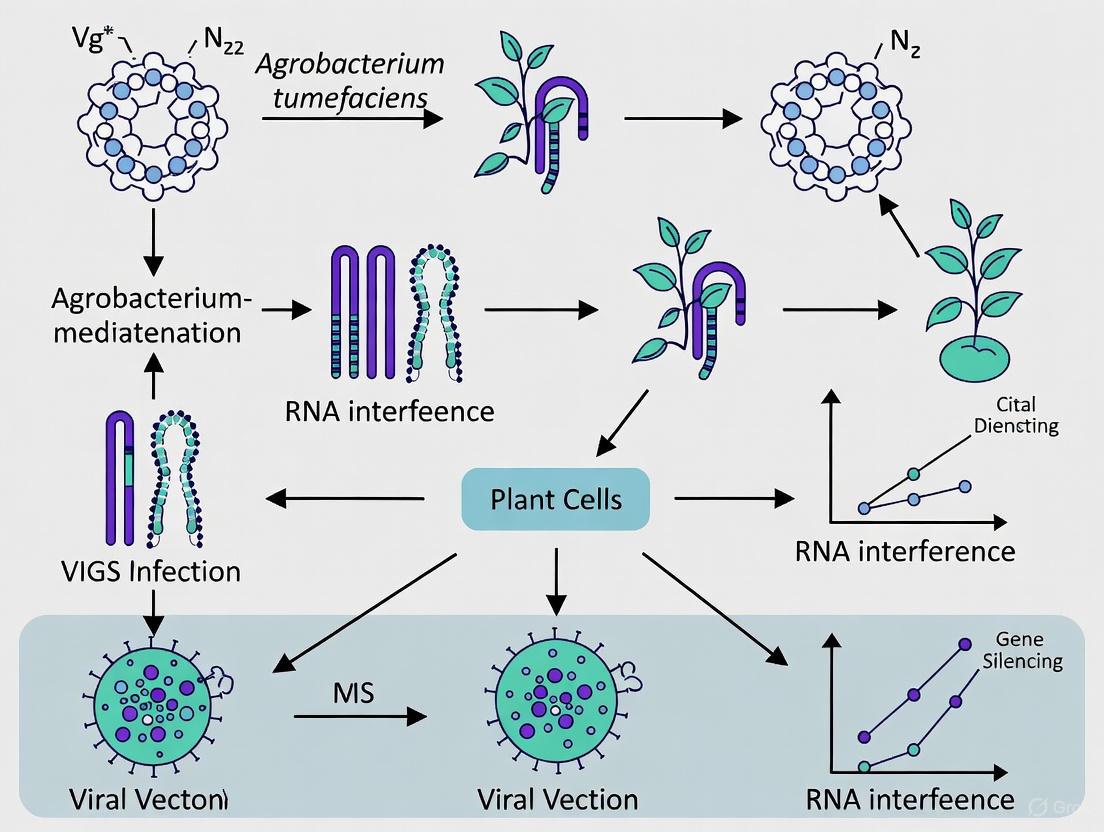
The biological foundation of VIGS is the plant's PTGS machinery, an antiviral defense system [1]. The mechanism can be broken down into a series of sequential steps, as illustrated in the diagram below.
Figure 1. The Molecular Mechanism of VIGS. This diagram illustrates the key steps of Post-Transcriptional Gene Silencing (PTGS) that underpin the VIGS technique, from the initial delivery of the viral vector to the final silencing of the target gene.
- Invasion and Replication: A recombinant virus, most commonly the Tobacco Rattle Virus (TRV), is introduced into the plant cell, often via Agrobacterium tumefaciens delivery. The TRV vector is engineered to carry a fragment (typically 200-500 base pairs) of the plant's endogenous gene targeted for silencing [1] [2]. Once inside, the virus begins to replicate.
- Double-Stranded RNA (dsRNA) Formation: During viral replication, double-stranded RNA (dsRNA) molecules are formed as intermediates. These dsRNAs are the key trigger for the plant's silencing machinery [2].
- DICER Cleavage: The plant recognizes the dsRNA as a foreign molecule and activates DICER-like (DCL) enzymes. These enzymes cleave the long dsRNA into small fragments of 21- to 24-nucleotides in length, known as small interfering RNAs (siRNAs) [1] [2].
- RISC Assembly: The double-stranded siRNAs are then incorporated into a multi-protein complex known as the RNA-Induced Silencing Complex (RISC). One strand of the siRNA duplex (the guide strand) is retained to direct the complex to complementary RNA sequences [2].
- Target mRNA Degradation: The RISC complex, guided by the siRNA, scans the cellular mRNA pool. Upon finding an mRNA sequence with perfect or near-perfect complementarity to its siRNA guide, the "slicer" enzyme Argonaute (AGO) within RISC cleaves the target mRNA [1]. This degradation prevents the mRNA from being translated into a functional protein, thereby silencing the target gene and potentially leading to a observable phenotype.
Key Experimental Protocols in Agrobacterium-Mediated VIGS
Optimizing the delivery of the VIGS construct is critical for high silencing efficiency. Below are detailed protocols for several advanced Agrobacterium-mediated infection methods.
Cotyledon Node Immersion in Soybean
This tissue culture-based method overcomes the challenges posed by soybean's thick leaf cuticle and dense trichomes [3] [4].
- Key Steps:
- Plant Material Preparation: Surface-sterilize soybean seeds and soak in sterile water until swollen. Bisect the seeds longitudinally to obtain half-seed explants.
- Agrobacterium Preparation: Transform the recombinant pTRV1 and pTRV2 vectors (carrying the target gene fragment) into Agrobacterium tumefaciens strain GV3101. Grow cultures to an OD₆₀₀ of ~0.8-1.0 in infiltration medium (10 mM MgCl₂, 10 mM MES, 150-200 μM acetosyringone) [3] [5].
- Inoculation: Mix the pTRV1 and pTRV2 cultures in a 1:1 ratio. Immerse the fresh half-seed explants in the mixed Agrobacterium suspension for 20-30 minutes with gentle agitation.
- Co-cultivation and Growth: Transfer the infected explants to tissue culture media for a brief co-cultivation period before moving them to soil for growth.
- Efficiency Validation: On the fourth day post-infection, excision of the hypocotyl and observation under a fluorescence microscope (if using a TRV2-GFP vector) confirmed infection efficiencies exceeding 80% [3].
Injection of No-Apical-Bud Stem Sections (INABS) in Tomato
The INABS method targets young, actively growing axillary bud tissues for highly efficient and rapid silencing [6].
- Key Steps:
- Plant Material Preparation: Use tomato seedlings with a "Y-type" stem section containing an axillary bud that is about 1–3 cm in length, with the apical bud removed.
- Agrobacterium Preparation: Prepare Agrobacterium cultures carrying pTRV1 and pTRV2 (e.g., pTRV2-SlPDS) as described previously, resuspending to an optimal OD₆₀₀ of 1.0 [6].
- Inoculation: Using a plastic syringe and needle, slowly inject 100–200 μL of the mixed Agrobacterium suspension into the bare stem of the no-apical-bud section. Infiltration is complete when a film of the liquid forms at the top of the stem.
- Phenotype Observation: Silencing phenotypes (e.g., photobleaching from PDS silencing) can appear in the emerging axillary buds as early as 6-8 days post-inoculation (dpi), achieving a silencing efficiency of up to 56.7% [6].
Root Wounding-Immersion for Multiple Species
This robust method is suitable for inoculating large batches of plants and is applicable across species like N. benthamiana, tomato, and pepper [5].
- Key Steps:
- Plant Material Preparation: Grow seedlings to the 3-4 leaf stage (approximately 3 weeks old) and carefully remove them from the soil.
- Root Wounding: Wash roots in pure water to remove soil. Using a sterilized blade, cut approximately one-third of the root system lengthwise.
- Inoculation: Immerse the wounded roots in the mixed Agrobacterium suspension (TRV1:TRV2) for 30 minutes. The "concurrent inoculation" method, where both vectors are mixed before immersion, is highly effective.
- Re-planting and Growth: Re-plant the seedlings into fresh soil or growth medium. Systemic silencing is observed in new leaves, with reported silencing rates of 95-100% in N. benthamiana and tomato [5].
Quantitative Data and Efficiency Metrics
The efficiency of VIGS is influenced by multiple factors. The following tables summarize key optimization parameters and performance metrics from recent studies.
Table 1. Factors Influencing VIGS Efficiency and Typical Optimal Ranges
| Factor | Impact on Efficiency | Typical Optimal Range | References |
|---|---|---|---|
| Agrobacterium OD₆₀₀ | Concentration affects infectivity & symptom severity | OD 0.8 - 1.5 | [6] [5] |
| Acetosyringone Concentration | Induces virulence genes; critical for T-DNA transfer | 150 - 200 μM | [5] [7] |
| Plant Genotype | Susceptibility to viral infection and systemic spread varies | Species and cultivar dependent | [3] [8] |
| Plant Developmental Stage | Younger tissues generally show higher silencing efficiency | Seedling stage (e.g., 1-4 true leaves) | [6] [9] |
| Temperature & Light | Low temperature and specific photoperiods can enhance silencing | e.g., 23°C, 16/8h light/dark | [2] [10] |
Table 2. VIGS Efficiency Metrics Across Different Plant Species and Methods
| Plant Species | Infiltration Method | Target Gene | Reported Silencing Efficiency | References |
|---|---|---|---|---|
| Soybean (Tianlong 1) | Cotyledon Node Immersion | GmPDS | 65% - 95% | [3] [4] |
| Tomato | INABS | SlPDS | 56.7% | [6] |
| Sunflower (various genotypes) | Seed Vacuum Infiltration | HaPDS | 62% - 91% (infection rate) | [8] |
| Camellia drupifera | Pericarp Cutting Immersion | CdCRY1, CdLAC15 | ~69.8% - ~90.91% | [9] |
| Nicotiana benthamiana, Tomato | Root Wounding-Immersion | PDS | 95% - 100% | [5] |
| Styrax japonicus | Vacuum Infiltration | - | 83.33% | [7] |
The experimental workflow for establishing a VIGS system, from design to validation, is summarized below.
Figure 2. VIGS Experimental Workflow. A generalized flowchart for conducting a VIGS experiment, from molecular cloning of the construct to phenotypic and molecular validation of gene silencing.
The Scientist's Toolkit: Essential Research Reagents
A successful VIGS experiment relies on a core set of reagents and vectors. The table below lists the essential components.
Table 3. Essential Reagents for TRV-based VIGS Experiments
| Reagent / Solution | Function / Role in VIGS | Specific Examples / Notes |
|---|---|---|
| TRV Vectors (pTRV1, pTRV2) | Bipartite viral vector system; pTRV2 carries the target gene insert. | pYL192 (TRV1), pYL156 (TRV2); pTRV2-GFP for tracking [8] [5]. |
| Agrobacterium tumefaciens | Delivery vehicle for the TRV DNA construct into plant cells. | Strain GV3101 is most commonly used [3] [5] [10]. |
| Infiltration Buffer | Medium for suspending Agrobacterium during inoculation. | 10 mM MgCl₂, 10 mM MES, 150-200 μM Acetosyringone (induces virulence) [6] [5]. |
| Marker Genes | Positive controls to visually confirm silencing efficiency. | PDS/CLA1 (causes photobleaching), GoPGF (causes gland loss, less lethal) [3] [10]. |
| Antibiotics | Selection for bacterial and plasmid containment. | Kanamycin (for TRV vectors), Rifampicin (for Agrobacterium strain), Gentamicin [6] [8]. |
Advanced Applications and Novel Markers
Beyond routine gene silencing, VIGS is being refined for more advanced applications. A significant development is the identification of superior marker genes. While the phytoene desaturase (PDS) gene, which causes a characteristic photobleaching phenotype when silenced, has been the standard visual marker, it has a major drawback: the silencing phenotype is lethal or severely stunts plant growth, preventing long-term studies [3] [10].
To address this, a novel marker gene, Gossypium PIGMENT GLAND FORMATION GENE (GoPGF), was developed in cotton. Silencing GoPGF reduces the density of pigment glands in cotton tissues without affecting normal plant growth and development. This allows researchers to visually trace silencing efficiency throughout the entire plant life cycle, including during reproductive stages like flowering and boll development, which was not feasible with the lethal PDS marker [10]. This innovation highlights how protocol refinements continue to expand the utility of VIGS in functional genomics.
The synergy between the plant's native PTGS defense mechanism and the engineered VIGS technology creates a uniquely powerful tool for functional genomics. A deep understanding of this mechanism is essential for troubleshooting and optimizing Agrobacterium-mediated VIGS protocols. As evidenced by the continuous development of novel infiltration methods, optimized parameters, and advanced tools like the GoPGF marker, VIGS remains a dynamic and indispensable technique. It enables researchers to rapidly link gene sequences to biological functions, thereby accelerating crop improvement and basic plant science research, particularly in genetically recalcitrant species.
Within plant functional genomics, Virus-Induced Gene Silencing (VIGS) has emerged as a powerful reverse genetics tool for rapidly assessing gene function. This application note details the operational profiles, optimized protocols, and comparative advantages of two predominant VIGS vectors—Tobacco Rattle Virus (TRV) and Bean Pod Mottle Virus (BPMV)—with a specific focus on their implementation via Agrobacterium-mediated delivery. As the demand for high-throughput functional validation grows, understanding the distinct characteristics of these systems is paramount for researchers investigating disease resistance, metabolic pathways, and developmental genetics in both model and non-model plants.
Vector System Profiles and Applications
Tobacco Rattle Virus (TRV)
The TRV system is celebrated for its broad host range and minimal symptomatic interference, which makes it particularly valuable for phenotypic analysis in dicot species.
- Key Features: TRV is a bipartite virus consisting of RNA1 (encoding replication and movement proteins) and RNA2 (encoding the coat protein and containing the insertion site for target genes) [4]. Its ability to spread efficiently throughout the plant, including meristematic tissues, and induce relatively mild symptoms allows for clear observation of silencing phenotypes [4].
- Agro-infiltration Method: The standard delivery method involves using Agrobacterium tumefaciens strains (e.g., GV3101) harboring the binary vectors pTRV1 and pTRV2. The target gene fragment is cloned into the Multiple Cloning Site (MCS) of pTRV2 [4]. An optimized protocol for soybean involves infecting half-seed explants via a 20-30 minute immersion in an Agrobacterium suspension, achieving infection efficiencies of up to 95% [4].
- Silencing Efficacy: In soybean, TRV-mediated silencing can achieve efficiency rates ranging from 65% to 95%, with phenotypes observable within weeks post-infection [4].
Bean Pod Mottle Virus (BPMV)
The BPMV system is a well-established, robust vector specifically optimized for legumes, especially soybean.
- Key Features: BPMV is also a bipartite, positive-strand RNA virus. Its RNA2 is engineered to accommodate gene inserts for silencing or protein expression [11] [12]. The "one-step" BPMV vector allows for direct rub-inoculation of plasmid DNA, bypassing the need for in vitro transcription and simplifying high-throughput studies [12].
- Silencing Patterns: BPMV-induced silencing is potent and widespread. Research on a transgenic GFP soybean line demonstrated that BPMV can achieve near-complete silencing in leaves, stems, flowers, and roots [13]. Silencing was observed as early as 14 days post-inoculation (dpi) in leaves and persisted for up to 7 weeks in flowers, demonstrating its longevity [13]. The orientation and location of the insert are critical; a construct targeting the 3' end of a gene in antisense orientation was found to produce the strongest silencing phenotype [13].
Comparative Analysis of TRV and BPMV
The choice between TRV and BPMV depends on the host plant, target tissue, and specific experimental goals. The following table summarizes their key characteristics for direct comparison.
Table 1: Comparative Analysis of TRV and BPMV VIGS Vector Systems
| Feature | Tobacco Rattle Virus (TRV) | Bean Pod Mottle Virus (BPMV) |
|---|---|---|
| Typical Host Range | Broad (e.g., Tomato, Tobacco, Arabidopsis, Cotton) [4] | Primarily Legumes (Soybean, Common Bean) [12] |
| Key Delivery Method | Agrobacterium-mediated (e.g., shoot apical meristem injection, seed immersion) [14] [4] | Direct plasmid rubbing or Agrobacterium-mediated [12] [4] |
| Silencing Efficiency | 65% - 95% (in soybean) [4] | Near-complete in aerial tissues; strong in roots [13] |
| Onset of Silencing | Within weeks [4] | As early as 14 dpi in leaves [13] |
| Duration of Silencing | Several weeks | Up to 7 weeks (long-lasting) [13] |
| Tissue Coverage | Systemic, including meristems [4] | Systemic; strong in leaves, stems, flowers, roots [13] |
| Ideal Insert Orientation | N/A (typically sense orientation in MCS) | 3'-end antisense orientation is most effective [13] |
| Typical Insert Size | Fragments from 132 bp effective in related systems [12] | 132 bp to 391 bp (as tested in common bean) [12] |
| Visual Symptoms | Mild, minimal interference [4] | Can induce mosaic patterns; milder strains available [13] [12] |
Essential Workflows and Signaling Pathways
The following diagrams illustrate the core experimental workflow for Agrobacterium-mediated VIGS and the subsequent plant RNAi signaling pathway that it hijacks.
VIGS Workflow via Agrobacterium
Diagram 1: VIGS Experimental Workflow. The process from vector construction to phenotypic analysis.
RNAi Signaling in Plant Defense
Diagram 2: RNAi Signaling Pathway. The cellular mechanism of VIGS.
The Scientist's Toolkit: Key Research Reagent Solutions
Successful implementation of VIGS relies on a suite of specialized reagents and vectors. The table below catalogues essential materials for establishing these systems.
Table 2: Essential Research Reagents for Agrobacterium-mediated VIGS
| Reagent / Material | Function / Description | Example Use Cases |
|---|---|---|
| pTRV1 & pTRV2 Vectors | Bipartite TRV system; target gene is cloned into pTRV2 MCS. | General VIGS in solanaceous plants, Arabidopsis, soybean [4]. |
| BPMV IA-R1M & IA-V1 | "One-step" BPMV vectors for direct plasmid rubbing or agro-delivery. | High-throughput silencing and protein expression in soybean [12]. |
| A. tumefaciens GV3101 | Standard disarmed helper strain for plant transformation. | Delivery of TRV and other binary vectors [14] [4]. |
| Phytoene Desaturase (PDS) | A marker gene; silencing causes photobleaching, visually confirming success. | Optimizing protocols and testing silencing efficiency in new systems [12] [4]. |
| Green Fluorescent Protein (GFP) | A visual reporter gene for tracking virus spread and infection efficiency. | Using transgenic GFP plants to map spatial/temporal silencing patterns [13]. |
| Cell Line Development Platform | Automated systems for generating clonal producer cell lines. | Ensuring regulatory compliance by proving clonality in bioproduction [15]. |
Detailed Experimental Protocols
Optimized TRV-VIGS Protocol for Soybean
This protocol, adapted from recent research, achieves high efficiency through cotyledon node transformation [4].
Vector Construction:
- Amplify a 132-391 bp fragment of the target gene (e.g., GmPDS).
- Clone the fragment into the pTRV2-GFP vector using appropriate restriction sites (e.g., EcoRI and XhoI).
- Sequence-verify the recombinant plasmid and transform it into Agrobacterium tumefaciens GV3101.
Plant Material Preparation:
- Surface-sterilize soybean seeds.
- Imbibe seeds in sterile water for 5-6 hours.
- Carefully bisect the swollen seeds longitudinally to create half-seed explants.
Agro-infection and Co-cultivation:
- Prepare suspensions of Agrobacterium containing pTRV1 and the recombinant pTRV2.
- Mix the two suspensions in a 1:1 ratio.
- Immerse the half-seed explants in the Agrobacterium suspension for 20-30 minutes with gentle agitation.
- Blot-dry the explants and co-cultivate them on induction medium in the dark for 2-3 days.
Plant Regeneration and Analysis:
- Transfer explants to light and monitor for GFP fluorescence to confirm infection.
- Select successfully infected seedlings and transplant them into soil.
- Monitor plants for the development of silencing phenotypes (e.g., photobleaching for PDS) and harvest tissue for molecular validation (qPCR) 3-4 weeks post-infection.
BPMV-VIGS Protocol Highlights
For the BPMV system, key optimized parameters in common bean include [12]:
- Plasmid Quantity: Use 5 µg each of BPMV RNA1 and RNA2-derived plasmids for rub-inoculation to achieve >90% infection rates.
- Strain Selection: Employ the IA-Di1 isolate-based vector, which induces very mild symptoms, minimizing interference with silencing phenotypes.
TRV and BPMV are complementary workhorses in the plant VIGS toolkit. TRV offers broad applicability and mild symptoms, while BPMV provides potent, long-lasting silencing specifically in legumes. The continued refinement of Agrobacterium-mediated delivery protocols, as exemplified by the high-efficiency soybean transformation method, is crucial for expanding the frontiers of plant functional genomics. By enabling rapid, high-throughput gene validation, these vector systems accelerate the discovery of agronomically important genes, directly supporting the development of improved crop varieties.
Agrobacterium tumefaciens is a cornerstone tool in plant biotechnology, serving as the principal delivery mechanism for Virus-Induced Gene Silencing (VIGS) constructs. VIGS itself is a powerful reverse genetics technique that leverages the plant's innate post-transcriptional gene silencing (PTGS) machinery to target specific endogenous mRNAs for degradation, enabling rapid functional analysis of plant genes without the need for stable transformation [1]. The TRV (Tobacco Rattle Virus) vector system has emerged as one of the most versatile and widely adopted VIGS platforms due to its broad host range, efficient systemic movement, and mild symptomatic impact on plant hosts [3] [1]. The effectiveness of Agrobacterium-mediated VIGS is influenced by multiple interdependent factors, including the plant genotype, developmental stage at inoculation, Agrobacterium culture density, inoculation methodology, and post-inoculation environmental conditions [16] [1] [8]. This protocol outlines optimized procedures for implementing TRV-based VIGS across diverse plant species, providing a standardized framework for researchers to investigate gene function.
Key Research Reagent Solutions
The following table catalogues essential reagents and materials required for establishing Agrobacterium-mediated VIGS systems.
Table 1: Essential Research Reagents for Agrobacterium-Mediated VIGS
| Reagent/Material | Function/Application | Examples & Notes |
|---|---|---|
| Agrobacterium Strain | Delivery vector for TRV constructs | GV3101 is widely used [3] [8] [17]. |
| TRV Vector System | Bipartite viral vector for silencing | pTRV1 (encodes replication/movement proteins) and pTRV2 (carries target gene insert) [3] [1]. |
| Visual Marker Gene | Silencing efficiency indicator | Phytoene desaturase (PDS); silencing causes photobleaching [3] [6] [17]. |
| Induction Medium Additive | Enhances T-DNA transfer | Acetosyringone; typically used at 100–200 μM [18] [6]. |
| Selection Antibiotics | Maintains plasmid integrity in Agrobacterium | Kanamycin, Gentamicin, Rifampicin [8]. |
Recent research has demonstrated the successful application of Agrobacterium-delivered TRV-VIGS across a phylogenetically diverse range of plant species. The table below summarizes key performance metrics from recent studies, highlighting the protocol efficiency and phenotypic outcomes.
Table 2: Performance Metrics of Agrobacterium-Mediated TRV-VIGS Across Plant Species
| Plant Species | Target Gene | Silencing Efficiency/Infection Rate | Key Phenotypic Observation | Primary Inoculation Method |
|---|---|---|---|---|
| Soybean (Glycine max) | GmPDS | 65% - 95% [3] | Systemic photobleaching [3] | Cotyledon node immersion [3] |
| Tomato (Solanum lycopersicum) | SlPDS | 56.7% [6] | Leaf photobleaching [6] | Injection of no-apical-bud stem section [6] |
| Sunflower (Helianthus annuus) | HaPDS | Up to 91% infection rate [8] | Photo-bleached spots on leaves [8] | Seed vacuum infiltration [8] |
| Periwinkle (Catharanthus roseus) | CrChlH | High (method validated) [17] | Yellow cotyledons [17] | Vacuum infiltration of seedlings [17] |
| Peony (Paeonia ostii) | - | Optimized system [18] | GUS/GFP reporter expression [18] | In vitro embryo-derived seedling infection [18] |
Established Experimental Protocols
Soybean Cotyledon Node Method
An optimized protocol for soybean utilizes the cotyledon node for highly efficient, systemic silencing.
- Vector Construction: Clone a 300-500 bp fragment of the target gene (e.g., GmPDS) into the multiple cloning site of the pTRV2 vector using appropriate restriction enzymes (e.g., EcoRI and XhoI) [3].
- Agrobacterium Preparation: Transform the recombinant pTRV2 and the helper pTRV1 plasmids separately into Agrobacterium tumefaciens strain GV3101. Culture individual colonies in LB medium with appropriate antibiotics and acetosyringone (200 µM) to an OD600 of 0.8-1.0 [3].
- Plant Material Preparation: Surface-sterilize soybean seeds and germinate on moist filter paper. Bisect the swollen seeds longitudinally to create half-seed explants [3].
- Agroinfiltration: Combine the pTRV1 and pTRV2-target Agrobacterium cultures in a 1:1 ratio. Immerse the fresh half-seed explants in the suspension for 20-30 minutes with gentle agitation [3].
- Plant Growth and Analysis: Co-cultivate the infected explants on tissue culture media for 2-3 days before transferring to soil. Silencing phenotypes, such as photobleaching for GmPDS, typically become visible systemically in newly emerged leaves within 21 days post-inoculation (dpi) [3].
Tomato No-Apical-Bud Stem Section (INABS) Method
This method offers a rapid and highly efficient silencing approach for tomato and other solanaceous plants.
- Plant Growth: Grow tomato plants until they develop stems with asymmetric "Y-type" structures containing an axillary bud that is approximately 1-3 cm in length [6].
- Agroinfiltration: Using a plastic syringe and needle, slowly inject 100-200 µL of the Agrobacterium suspension (OD600 = 1.0) directly into the bare stem of the section. Successful infiltration is indicated by a thin film of liquid forming at the top of the stem [6].
- Phenotype Observation: Silencing phenotypes can be observed in the emerging axillary buds as early as 6-8 dpi, with widespread effects in grown leaves by 10 dpi [6].
Sunflower Seed Vacuum Infiltration Protocol
A simple and robust protocol optimized for sunflowers, which are traditionally recalcitrant to transformation.
- Seed Preparation: Peel the seed coats of sunflower seeds. No surface sterilization or in vitro recovery steps are required [8].
- Vacuum Infiltration: Immerse the peeled seeds in the Agrobacterium suspension (OD600 ~0.8). Apply a vacuum of -0.08 MPa for 10 minutes, then slowly release to atmospheric pressure [8].
- Co-cultivation: Subject the infiltrated seeds to a 6-hour co-cultivation period in the dark [8].
- Planting and Evaluation: Sow the seeds directly in soil. This method achieves high infection percentages and allows for extensive viral spread throughout the plant, with TRV detectable in leaves up to node 9 [8].
Workflow and Mechanism of Agrobacterium-Mediated VIGS
The following diagram illustrates the complete experimental workflow and the underlying molecular mechanism of Agrobacterium-mediated VIGS.
Agrobacterium-Mediated VIGS Workflow and Mechanism
The diagram summarizes the integrated biological and experimental process. The yellow nodes trace the key laboratory steps, from construct preparation to final phenotypic analysis. The green nodes depict the core molecular mechanism inside the plant cell: the TRV vector is transcribed to produce RNA, which forms double-stranded RNA (dsRNA), a key trigger for the plant's PTGS system. This dsRNA is processed by Dicer-like enzymes into small interfering RNAs (siRNAs). These siRNAs are loaded into the RNA-induced silencing complex (RISC), which guides the sequence-specific cleavage and degradation of complementary target mRNA, resulting in gene silencing [1]. The entire process is initiated by the plant's innate PTGS machinery, shown in red, which is co-opted by the engineered virus to silence endogenous genes.
Virus-Induced Gene Silencing (VIGS) has emerged as a powerful reverse genetics tool for rapidly analyzing gene function in plants. This Agrobacterium-mediated technology enables transient gene knockdown without the need for stable transformation, significantly accelerating functional genomics studies. A critical component of successful VIGS implementation is the use of visual reporter genes that provide visible markers to monitor silencing efficiency, spatial distribution, and timing of gene knockdown throughout the plant.
Among the various visual reporters available, phytoene desaturase (PDS) and chalcone synthase (CHS) have become the gold standards for optimizing and validating VIGS systems across numerous plant species. These reporters enable researchers to visually assess silencing efficiency before investigating target genes of interest, providing crucial validation of experimental protocols. This application note details the mechanistic basis, implementation protocols, and quantitative assessment methods for using PDS and CHS as visual reporters in Agrobacterium-mediated VIGS systems, with specific frameworks for integration into broader thesis research on VIGS methodology optimization.
Fundamental Principles of PDS and CHS as Visual Reporters
Phytoene Desaturase (PDS): A Vegetative Tissue Reporter
The PDS gene encodes a key enzyme in the carotenoid biosynthesis pathway, catalyzing the conversion of phytoene to ζ-carotene. Carotenoids serve essential functions in photosynthesis, including photoprotection and light-harvesting. When PDS expression is silenced, carotenoid depletion leads to photobleaching—a characteristic white or yellow discoloration of normally green tissues due to chlorophyll degradation under light exposure [16] [19]. This visible phenotype makes PDS an ideal visual marker for monitoring VIGS efficiency in photosynthetic tissues such as leaves, stems, and sepals.
The photobleaching phenotype typically manifests as sectorial patterns following the vasculature, indicating the systemic movement of the silencing signal. The extent and intensity of photobleaching provide semi-quantitative measures of silencing efficiency, with more widespread and severe bleaching correlating with stronger gene knockdown [16]. PDS silencing has been successfully employed as a visual reporter across diverse species, including petunia, soybean, tea plants, and many other crops [16] [3] [19].
Chalcone Synthase (CHS): A Pigmented Tissue Reporter
The CHS gene encodes the first committed enzyme in the flavonoid/anthocyanin biosynthesis pathway, catalyzing the stepwise condensation of 4-coumaroyl-CoA and malonyl-CoA to form naringenin chalcone. Flavonoids and anthocyanins are secondary metabolites responsible for pigmentation in flowers, fruits, and sometimes leaves. Silencing of CHS results in loss of pigmentation, transforming normally pigmented tissues to white or pale colors [16].
This characteristic makes CHS particularly valuable for monitoring VIGS efficiency in floral tissues and pigmented fruits, where the color change from pigmented to white provides a clear visual indicator of successful gene silencing. In petunia, for instance, CHS silencing results in distinctive white sectors on otherwise pigmented petals, enabling quantitative assessment of silencing efficiency through color pattern analysis [16]. The non-lethal nature of CHS silencing makes it especially suitable for studies focused on reproductive tissues and floral biology.
The following diagram illustrates the contrasting mechanisms of PDS and CHS silencing and their resulting visual phenotypes:
Visual Reporter Mechanisms and Phenotypes
Quantitative Comparison of PDS and CHS Visual Reporters
Table 1: Comparative Analysis of PDS and CHS as Visual Reporters in VIGS
| Parameter | PDS (Phytoene Desaturase) | CHS (Chalcone Synthase) |
|---|---|---|
| Biological Pathway | Carotenoid biosynthesis | Flavonoid/anthocyanin biosynthesis |
| Primary Visual Phenotype | Photobleaching (white/yellow tissue) | Loss of pigmentation (white tissue) |
| Optimal Tissue Type | Leaves, stems, green tissues | Flowers, fruits, pigmented tissues |
| Phenotype Onset | 7-14 days post-inoculation | 10-21 days post-inoculation |
| Silencing Duration | 2-4 weeks | 3-5 weeks |
| Quantification Methods | Photobleached area measurement, chlorophyll content assays | Color intensity measurement, anthocyanin extraction |
| Impact on Plant Health | Can be lethal with extensive silencing | Generally non-lethal |
| Reported Silencing Efficiency | 28-95% across species [16] [3] | Up to 69% area in petunia corollas [16] |
| Key Applications | VIGS optimization in vegetative tissues, meristem silencing studies | Floral trait studies, pigmentation genetics |
Table 2: Optimization Parameters for Enhanced Visual Reporter Silencing Efficiency
| Optimization Factor | Optimal Condition for PDS | Optimal Condition for CHS | Effect on Silencing |
|---|---|---|---|
| Temperature Regime | 20°C day/18°C night (petunia) [16] | 20°C day/18°C night (petunia) [16] | Lower temperatures enhance silencing spread |
| Plant Developmental Stage | 3-4 weeks after sowing (petunia) [16] | 3-4 weeks after sowing (petunia) [16] | Younger plants show more efficient silencing |
| Inoculation Method | Mechanically wounded apical meristems [16] | Mechanically wounded apical meristems [16] | Direct meristem access improves systemic spread |
| Agrobacterium OD600 | 0.5-1.0 [7] | 0.5-1.0 [7] | Optimal bacterial density for infection |
| Acetosyringone Concentration | 200 μmol·L⁻¹ [7] | 200 μmol·L⁻¹ [7] | Enhances Agrobacterium virulence |
Experimental Protocol: Agrobacterium-Mediated VIGS Using Visual Reporters
Vector Construction and Agrobacterium Preparation
Research Reagent Solutions:
- pTRV1 and pTRV2 Vectors: TRV-based binary vectors for VIGS [16] [1]
- pTRV2-PDS/pTRV2-CHS: Recombinant vectors containing gene-specific fragments [16] [20]
- Agrobacterium tumefaciens GV3101: Standard strain for plant transformation [3]
- Acetosyringone: Phenolic compound that induces Vir gene expression [7]
Protocol:
- Gene Fragment Selection: Amplify 200-400 bp gene-specific fragments from the target plant's PDS or CHS cDNA using sequence-specific primers with appropriate restriction sites (e.g., EcoRI and XhoI) [3].
- Vector Construction: Ligate the purified PCR fragment into the pTRV2 vector digested with corresponding restriction enzymes. Transform into E. coli DH5α competent cells and verify positive clones by sequencing [3].
- Agrobacterium Transformation: Introduce verified recombinant plasmids into Agrobacterium tumefaciens strain GV3101 through electroporation or freeze-thaw method.
- Culture Preparation: Inoculate single colonies of Agrobacterium harboring pTRV1, pTRV2-PDS, or pTRV2-CHS in 5 mL YEP medium with appropriate antibiotics (kanamycin, rifampicin). Grow overnight at 28°C with shaking at 200 rpm.
- Induction Culture: Subculture the overnight grown Agrobacterium (1:50 ratio) in induction medium (YEP with 10 mM MES, 20 μM acetosyringone) and grow to OD600 = 0.5-1.0 [7].
- Inoculum Preparation: Harvest bacterial cells by centrifugation (3000 × g, 10 min) and resuspend in infiltration buffer (10 mM MgCl2, 10 mM MES, 200 μM acetosyringone) to final OD600 = 0.5-2.0. Incubate the suspension at room temperature for 3-4 hours before inoculation [16] [7].
Plant Inoculation and Incubation
The following workflow outlines the complete experimental procedure for Agrobacterium-mediated VIGS using visual reporters:
VIGS Experimental Workflow
Inoculation Methods:
- Meristem Wounding (Petunia): Mechanically wound shoot apical meristems with a needle and apply 10-20 μL of mixed Agrobacterium suspension (pTRV1 + pTRV2-PDS/CHS in 1:1 ratio) directly to wounded sites [16].
- Cotyledon Node Immersion (Soybean): Bisect swollen, sterilized soybean seeds to obtain half-seed explants and immerse fresh explants in Agrobacterium suspension for 20-30 minutes [3].
- Vacuum Infiltration (Tea, Styrax): Submerge entire seedlings in Agrobacterium suspension and apply vacuum (250-500 mbar) for 2-5 minutes, then slowly release vacuum [7] [19].
- Agroinfiltration (N. benthamiana): Use needleless syringe to infiltrate Agrobacterium suspension into abaxial side of leaves [1].
Post-Inoculation Procedures:
- Maintain inoculated plants at 20°C day/18°C night temperatures with high humidity (70-80%) for 2-3 days to facilitate infection [16].
- Transfer plants to standard growth conditions (species-appropriate temperature, 16/8h photoperiod).
- Monitor for visual phenotypes beginning at 7 days post-inoculation (dpi) for PDS and 10-14 dpi for CHS.
- Document phenotypic progression through photography at regular intervals (3-4 days).
Efficiency Assessment and Troubleshooting
Quantitative Evaluation of Silencing Efficiency
Visual Scoring Method:
- Photobleaching Percentage: Calculate the percentage of total leaf area showing photobleaching symptoms using image analysis software (e.g., ImageJ) [16].
- Pigmentation Loss Index: For CHS, develop a scoring system (0-5) based on the extent of white sector formation on pigmented petals or fruits [16].
- Silencing Propagation Rate: Measure the rate at which silencing spreads from initial sites of inoculation to new growth.
Molecular Validation:
- RNA Extraction: Collect tissue samples from both silenced and non-silenced areas at 14-21 dpi.
- qRT-PCR Analysis: Perform quantitative RT-PCR using gene-specific primers for PDS/CHS and reference genes (e.g., Actin, EF1α).
- Efficiency Calculation: Calculate silencing efficiency as percentage reduction in transcript levels compared to empty vector controls:
Silencing Efficiency = [1 - (2-ΔΔCt)] × 100%
Reported efficiencies range from 28% for PDS to 69% for CHS in optimized petunia systems, with soybean systems achieving 65-95% efficiency [16] [3].
Troubleshooting Common Issues
Table 3: Troubleshooting Guide for Visual Reporter VIGS Experiments
| Problem | Potential Causes | Solutions |
|---|---|---|
| No visual phenotype | Incorrect plant developmental stage; Suboptimal Agrobacterium concentration; Improper inoculation technique | Use younger plants (3-4 weeks); Optimize OD600 (0.5-2.0); Validate inoculation method for specific species |
| Patchy or inconsistent silencing | Incomplete systemic movement; Temperature fluctuations; Genetic variability | Maintain constant optimal temperatures (20°C/18°C); Use uniform plant materials; Employ meristem-targeting inoculation |
| Severe viral symptoms in controls | Empty vector toxicity; High viral titers | Use control vectors with non-plant inserts (e.g., GFP); Optimize Agrobacterium concentration [16] |
| Delayed phenotype appearance | Suboptimal growing conditions; Weak viral replication | Ensure proper temperature and light conditions; Use freshly prepared Agrobacterium cultures |
| Limited silencing duration | Plant recovery from transient silencing; Viral clearance | Harvest tissues at peak silencing (14-21 dpi); Use more aggressive inoculation methods |
Application in Functional Genomics Research
The integration of PDS and CHS visual reporters into VIGS protocols provides critical validation tools for broader functional genomics studies. In gerbera, VIGS with visual reporters enabled the functional analysis of Botrytis cinerea resistance genes, where silenced plants showed significantly delayed lesion growth upon pathogen infection [21]. Similarly, in pepper and tea plants, established VIGS systems using these visual reporters have facilitated the identification of genes controlling fruit quality, disease resistance, and specialized metabolism [1] [19].
For thesis research focused on Agrobacterium-mediated VIGS optimization, systematic evaluation of visual reporter efficiency across parameters such as temperature regimes, developmental stages, inoculation methods, and vector configurations provides robust frameworks for protocol standardization. The quantitative data generated from PDS and CHS silencing not only validates experimental success but also enables statistical comparison of efficiency across different optimization approaches.
The consistent implementation of these visual reporters across plant species—from model organisms to crops—demonstrates their universal utility in VIGS-based functional genomics. Their non-destructive nature allows for longitudinal studies of gene function, while their visible phenotypes enable rapid screening of silencing efficiency before proceeding with target gene analysis, ultimately accelerating the pace of gene discovery and characterization in plant systems.
From Lab to Leaf: A Practical Guide to VIGS Infection Protocols
Agroinfiltration has emerged as a cornerstone technique in plant biotechnology, enabling transient gene expression for functional genomics, protein production, and genetic modification. This Agrobacterium-mediated approach facilitates the introduction of genetic material into plant cells without the need for stable transformation, providing a rapid and versatile platform for research and biopharmaceutical production. The technique's significance continues to grow with advancements in vector design and delivery methodologies, particularly within the context of virus-induced gene silencing (VIGS) and recombinant protein production [22] [23]. This application note provides a comprehensive overview of three fundamental agroinfiltration techniques—leaf injection, spraying, and vacuum infiltration—with detailed protocols optimized for diverse research applications in plant science and drug development.
Core Agroinfiltration Methodologies
Syringe Infiltration (Leaf Injection)
Syringe infiltration represents the most accessible entry point for agroinfiltration studies, requiring minimal specialized equipment while offering precision in localized gene delivery. The technique involves direct pressure-based introduction of Agrobacterium suspension into leaf intercellular spaces through hydraulic force [22] [23].
Experimental Protocol:
- Plant Material Preparation: Grow Nicotiana benthamiana or target species under controlled conditions (25°C, 16/8h light/dark cycle) for 5-6 weeks until fully expanded leaves develop [24] [25].
- Agrobacterium Culture: Inoculate Agrobacterium tumefaciens strains (GV3101, EHA105, or AGL1) harboring binary vectors in YENB or LB medium with appropriate antibiotics [26] [27]. Grow cultures overnight at 28°C with shaking (200-300 rpm) until OD600 reaches 0.4-0.8 [24] [25].
- Bacterial Resuspension: Pellet bacteria by centrifugation (3000 × g, 10 min) and resuspend in infiltration buffer (10 mM MES, 10 mM MgCl2, 100 µM acetosyringone, pH 5.6) to final OD600 of 0.2-1.0 [26] [25]. Incubate suspension at room temperature for 1-4 hours.
- Infiltration Procedure: Using a needleless syringe (1-5 mL), gently press the syringe tip against the abaxial (lower) leaf surface while applying counter-support to the adaxial side. Slowly depress the plunger to infiltrate the bacterial suspension, observing the formation of a water-soaked area [22] [23]. Multiple infiltrations can be performed on a single leaf to test different constructs.
- Post-Infiltration Incubation: Maintain infiltrated plants under normal growth conditions for 2-7 days before analysis [26] [28].
Spraying Techniques
Spray-based infiltration methods offer scalability for larger leaf areas or entire plants, though with potentially reduced efficiency compared to direct injection. Recent advancements have optimized droplet size and pressure parameters to enhance delivery efficiency.
Experimental Protocol:
- Agrobacterium Preparation: Prepare bacterial cultures as described for syringe infiltration, resuspending in buffer containing 0.02% Silwet L-77 surfactant to reduce surface tension [27] [23].
- Spray Apparatus Setup: Utilize a fine-mist spray bottle or agricultural sprayer capable of producing droplets of 50-100 µm diameter. Adjust pressure to 10-15 psi for optimal coverage without leaf damage.
- Application Technique: Uniformly spray the bacterial suspension onto abaxial leaf surfaces until runoff, ensuring complete coverage of stomatal openings.
- Post-Application Handling: Immediately place sprayed plants in high-humidity chambers (≥80% RH) for 24 hours to maintain tissue hydration and facilitate bacterial uptake [23].
Vacuum Infiltration
Vacuum infiltration provides the most uniform and scalable approach for whole-plant or multi-leaf transformation, making it ideal for high-throughput applications and industrial-scale protein production [22] [24]. The process involves submerged infiltration under negative pressure, forcing air from intercellular spaces and replacing it with bacterial suspension upon vacuum release.
Experimental Protocol:
- Plant Preparation: Excise whole plants or leaves and submerge in Agrobacterium suspension (OD600 0.5-1.0) in a vacuum-desiccator [24] [29].
- Vacuum Application: Apply vacuum (25-30 in Hg) for 30-90 seconds, observing bubble formation as air evacuates from intercellular spaces [22] [24].
- Infiltration Cycle: Rapidly release vacuum to allow bacterial suspension penetration. Repeat for 2-3 cycles if necessary for recalcitrant species.
- Post-Infiltration Care: Briefly drain excess suspension and return plants to normal growth conditions [24]. For whole-plant infiltration, maintain high humidity for 24 hours post-treatment to prevent desiccation.
Table 1: Comparative Analysis of Agroinfiltration Methodologies
| Parameter | Syringe Infiltration | Spray Infiltration | Vacuum Infiltration |
|---|---|---|---|
| Equipment Requirements | Needleless syringe | Spray apparatus | Vacuum chamber, pump |
| Scalability | Low (single leaves) | Medium (multiple plants) | High (whole plants, large batches) |
| Transformation Efficiency | Variable (dependent on operator skill) | Moderate | High and consistent |
| Labor Intensity | High | Moderate | Low (once established) |
| Typical Applications | Promoter analysis, protein subcellular localization, small-scale screening | Medium-scale protein production, partial plant transformation | Large-scale recombinant protein production, high-throughput studies |
| Optimal Plant Species | N. benthamiana, tomato, strawberry | N. benthamiana, Arabidopsis | N. benthamiana, lettuce, soybean |
| Reference | [22] [23] | [23] | [22] [24] |
Critical Experimental Parameters and Optimization
Successful agroinfiltration depends on numerous physical, biological, and chemical factors that influence transformation efficiency and transgene expression levels.
Agrobacterium Strain Selection
Strain specificity significantly impacts transformation efficiency across plant species. Comparative studies demonstrate that EHA105 often achieves highest transient expression in dicotyledonous species including strawberry and melon, while GV3101 and AGL1 show superior performance in N. benthamiana and solanaceous plants [26] [27]. For example, in Fragaria vesca, EHA105 yielded approximately 40% higher GUS reporter expression compared to GV3101 and LBA4404 strains [26].
Chemical Additives and Supplements
Strategic inclusion of chemical enhancers in infiltration media dramatically improves T-DNA transfer and transgene expression:
- Acetosyringone: A plant-derived phenolic compound that induces Agrobacterium vir gene expression. Optimal concentrations range from 100-500 µM [27] [25].
- Silwet L-77: A surfactant that reduces surface tension, enhancing suspension penetration through stomata. Use at 0.01-0.02% (v/v) [27].
- Antioxidants: Lipoic acid (5 µM) or ascorbic acid can reduce reactive oxygen species-mediated cell death during infiltration [25].
- Silencing Suppressors: Co-infiltration with vectors expressing viral proteins (p19, HC-Pro, etc.) inhibits post-transcriptional gene silencing, extending transgene expression duration and increasing protein yields up to 50-fold [30] [25].
Physical and Biological Parameters
- Bacterial Density: Optimal OD600 ranges from 0.2 to 1.0, with species-specific optimization required [26] [25].
- Plant Age and Tissue Status: Fully expanded leaves from 5-6 week old N. benthamiana plants typically show highest transformation efficiency [24] [25].
- Co-cultivation Time: Transgene expression typically peaks between 2-4 days post-infiltration (dpi), with protein detection possible for up to 7-10 dpi [26] [28].
- Temperature Optimization: Brief heat treatment (37°C for 15-30 min) 1-2 days post-infiltration can enhance protein expression by activating heat shock proteins [25].
Table 2: Optimization Parameters for High-Efficiency Agroinfiltration
| Parameter | Optimal Range | Effect | Reference |
|---|---|---|---|
| Agrobacterium Strain | EHA105, GV3101, AGL1 | Species-dependent transformation efficiency | [26] [27] |
| OD600 | 0.2-1.0 | Balanced between T-DNA delivery and plant stress response | [26] [25] |
| Acetosyringone | 100-500 µM | Induces vir gene expression, enhances T-DNA transfer | [27] [25] |
| Surfactant (Silwet L-77) | 0.01-0.02% | Reduces surface tension, improves infiltration | [27] |
| Antioxidants | 5 µM lipoic acid | Reduces oxidative stress and cell necrosis | [25] |
| Co-cultivation Time | 2-4 days | Peak transgene expression period | [26] [28] |
| Post-Infiltration Heat Shock | 37°C for 15-30 min | Activates heat shock proteins, enhances expression | [25] |
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Research Reagent Solutions for Agroinfiltration
| Reagent/Vector | Function | Application Notes | Reference |
|---|---|---|---|
| Agrobacterium tumefaciens EHA105 | T-DNA delivery | High virulence, superior for strawberry, melon | [26] [29] |
| Agrobacterium tumefaciens GV3101 | T-DNA delivery | Standard for N. benthamiana, good efficiency | [24] [25] |
| pEAQ-HT Vector | High-yield protein expression | CPMV-based hypertranslation system | [25] |
| p19 Silencing Suppressor | PTGS inhibition | From Tomato bushy stunt virus, boosts protein yield | [30] [25] |
| TRV-based VIGS Vectors | Virus-induced gene silencing | Effective in soybean, tomato, tobacco | [3] |
| Acetosyringone | vir gene inducer | Critical for efficient T-DNA transfer | [27] [25] |
| Silwet L-77 | Surfactant | Enhances tissue penetration, use at 0.01-0.02% | [27] |
Advanced Applications in Plant Biotechnology
Virus-Induced Gene Silencing (VIGS)
Agroinfiltration serves as the primary delivery method for VIGS, enabling rapid functional genomics studies without stable transformation. TRV (Tobacco Rattle Virus)-based vectors have been successfully deployed for high-efficiency gene silencing in soybean (65-95% efficiency), tomato, and tobacco [3]. The modified protocol for soybean involves Agrobacterium infection through cotyledon nodes, with systemic silencing spreading throughout the plant within 2-3 weeks [3].
Recombinant Protein Production
The scalability of vacuum infiltration makes it ideal for pharmaceutical protein production. Plant-based systems offer cost advantages over mammalian cell culture, with yields up to 1.5 g/kg leaf fresh weight reported for some target proteins in N. benthamiana [22] [25]. Recent advances include geminiviral vectors for gene amplification and glycoengineering platforms for humanized protein glycosylation [22] [24].
Genome Editing and Functional Genomics
Agroinfiltration enables transient delivery of CRISPR/Cas9 components for genome editing. In melon, co-expression of developmental regulators (AtGRF5, AtPLT5) with CRISPR/Cas9 vectors significantly improved transformation efficiency in recalcitrant genotypes [29]. Similarly, agroinfiltration facilitates rapid assessment of gene function and regulatory elements in species including strawberry, pigeonpea, and soybean [26] [3] [28].
Workflow Integration and Decision Framework
The following diagram illustrates the agroinfiltration methodology selection process based on research objectives and available resources:
Agroinfiltration Methodology Selection Workflow
Agroinfiltration methodologies provide powerful and versatile tools for plant biotechnology research and applications. Selection of the appropriate technique—syringe, spray, or vacuum infiltration—should be guided by research objectives, scale requirements, and available resources. Through careful optimization of biological, physical, and chemical parameters, researchers can achieve high-efficiency transformation for diverse applications ranging from rapid gene function analysis to large-scale production of pharmaceutical proteins. The continued refinement of these methodologies promises to further expand their utility in both basic and applied plant science research.
In the post-genomic era, functional characterization of genes is essential for advancing plant biology and crop improvement. Virus-Induced Gene Silencing (VIGS) has emerged as a powerful reverse genetics tool that leverages the plant's antiviral RNA silencing machinery to downregulate target gene expression [31]. Unlike stable transformation methods, VIGS offers a rapid, cost-effective alternative that does not require the generation of stable transgenic lines, enabling high-throughput functional screening [5] [31]. Commonly used Agrobacterium-mediated VIGS delivery methods include stem scratching, leaf infiltration, agrodrench, and spray-based applications. However, these techniques often face limitations regarding efficiency, scalability, and applicability across diverse species, particularly for root biology studies and plants resistant to above-ground infection [5].
The root wounding-immersion method represents a significant advancement in VIGS technology. This innovative approach involves partial root excision followed by immersion in Agrobacterium suspension containing tobacco rattle virus (TRV) vectors, enabling highly efficient systemic gene silencing [5] [32]. Developed and optimized for Solanaceous species including Nicotiana benthamiana, tomato (Solanum lycopersicum), pepper (Capsicum annuum L.), and eggplant (Solanum melongena), this method achieves remarkable silencing efficiencies of 95-100% for the marker gene phytoene desaturase (PDS) in N. benthamiana and tomato [5]. This protocol note details the establishment, optimization, and application of this transformative methodology within the broader context of Agrobacterium-mediated VIGS research.
Key Advantages and Performance Metrics
The root wounding-immersion method addresses several limitations associated with conventional VIGS inoculation techniques. Its principal advantages include:
- High Efficiency and Scalability: Enables rapid inoculation of large plant batches with minimal labor [5]
- Broad Host Compatibility: Successfully silences genes across multiple Solanaceous species and Arabidopsis thaliana [5] [32]
- Systemic Silencing: Facilitates uniform gene silencing from roots to aerial tissues [5]
- Resource Economy: Permits reuse of fresh bacterial infusions multiple times without efficiency loss [5]
- Early-Stage Application: Suitable for inoculating seedlings at early developmental stages (3-4 true leaves) [5]
Table 1: Silencing Efficiency of the Root Wounding-Immersion Method Across Plant Species
| Plant Species | Target Gene | Silencing Efficiency | Key Observations |
|---|---|---|---|
| Nicotiana benthamiana | PDS | 95-100% | Systemic photobleaching |
| Tomato (Solanum lycopersicum) | PDS | 95-100% | Systemic photobleaching |
| Tomato (Solanum lycopersicum) | SITL5, SITL6 | High (quantitative data not shown) | Decreased disease resistance |
| Pepper (Capsicum annuum L.) | PDS | Successful silencing | Phenotype observed |
| Eggplant (Solanum melongena) | PDS | Successful silencing | Phenotype observed |
| Arabidopsis thaliana | PDS | Successful silencing | Phenotype observed |
Table 2: Critical Parameters for Optimal Root Wounding-Immersion VIGS
| Parameter | Optimal Condition | Impact on Efficiency |
|---|---|---|
| Root Wounding | Removal of 1/3 root length | Creates infection portals |
| Immersion Duration | 30 minutes | Ensures adequate bacterial uptake |
| Bacterial Density (OD₆₀₀) | 0.8 | Balanced infection and plant viability |
| Plant Developmental Stage | 3 weeks old (3-4 true leaves) | Optimal systemic spread |
| Inoculation Temperature | Same as subsequent growth | Reduces environmental stress |
| Post-inoculation Dark Period | 48 hours | Enhances infection establishment |
Experimental Protocol
Vector Construction and Agrobacterium Preparation
The root wounding-immersion protocol utilizes the Tobacco Rattle Virus (TRV) VIGS system, consisting of two plasmid vectors: pTRV1 (encoding RNA replication proteins) and pTRV2 (carrying the target gene fragment) [5]. The methodology employs pTRV2-GFP as a backbone vector, enabling visual tracking of viral movement via green fluorescent protein expression [5] [32].
Procedure:
- Clone target gene fragments: Design specific primers to amplify approximately 300bp fragments of target genes (PDS, SITL5, SITL6) with appropriate restriction sites [5]
- Ligate into pTRV2-GFP: Directionally clone fragments into the TRV2-GFP binary vector to generate constructs such as TRV2-GFP-NbPDS, TRV2-GFP-SlPDS, etc. [5]
- Transform Agrobacterium: Introduce pTRV1 and recombinant pTRV2 constructs into Agrobacterium tumefaciens strain GV1301 via electroporation [5]
- Culture Agrobacterium: Plate transformed Agrobacterium on LB medium containing kanamycin (50μg/mL) and rifampicin (25μg/mL), incubate at 28°C for 48 hours [5]
- Prepare infiltration suspension:
- Select positive colonies and culture overnight in LB broth with appropriate antibiotics at 28°C, 200rpm
- Resuspend in infiltration solution (10mM MgCl₂, 10mM MES pH5.6, 150μM acetosyringone) to OD₆₀₀=0.8
- Incubate in dark at 28°C for 3 hours for induction [5]
Root Wounding-Immersion Inoculation
The core innovation of this method lies in the strategic combination of root wounding and immersion to achieve highly efficient viral delivery.
Procedure:
- Plant preparation: Grow seedlings until 3-4 true leaf stage (approximately 3 weeks) under controlled conditions (16h light/8h dark, 28°C light/20°C dark) [5]
- Root excision: Carefully remove plants from soil, wash roots with pure water to remove soil particles, and aseptically remove approximately one-third of the root length using a disinfected blade [5] [32]
- Bacterial inoculation: Employ one of two approaches:
- Temperature management: Maintain immersion solutions at temperatures matching subsequent growth conditions to minimize stress [5]
- Post-inoculation care: Transfer inoculated seedlings to sterile soil in trays, maintain in darkness for 48 hours, then return to standard growth conditions [5]
Validation and Efficiency Assessment
Silencing validation:
- Phenotypic monitoring: For PDS silencing, monitor photobleaching symptoms appearing 2-3 weeks post-inoculation [5]
- Molecular verification: Quantify target gene expression reduction via qRT-PCR comparing silenced tissues to controls [33]
- Viral tracking: Monitor GFP fluorescence movement from roots to stems and leaves using fluorescence microscopy [5]
Molecular Mechanism of VIGS
The root wounding-immersion method leverages the well-established molecular pathway of virus-induced gene silencing, with the innovation focusing on delivery efficiency rather than altering the core mechanism.
The diagram above illustrates the molecular events triggered by TRV vector delivery through root wounding-immersion. The process initiates when TRV vectors carrying plant target gene fragments enter root cells through wound sites. Within the plant cell, viral replication produces double-stranded RNA (dsRNA), which the plant's Dicer-like enzymes recognize and cleave into 21-24 nucleotide small interfering RNAs (siRNAs) [31]. These siRNAs integrate into the RNA-induced silencing complex (RISC), guiding it to complementary endogenous mRNA transcripts for sequence-specific degradation [31]. Secondary siRNAs amplified by host RNA-directed RNA polymerases (RDRPs) facilitate systemic spreading of silencing signals throughout the plant, enabling whole-plant gene silencing originating from the inoculated root system [31].
The Scientist's Toolkit: Essential Research Reagents
Table 3: Essential Research Reagents for Root Wounding-Immersion VIGS
| Reagent/Resource | Specification/Function | Application Notes |
|---|---|---|
| TRV Vectors | pTRV1 (RNA replication), pTRV2 (target insert) | Basis for VIGS construct system [5] |
| Agrobacterium Strain | GV1301 or GV3101 | Optimal for plant transformation [5] [3] |
| Antibiotics | Kanamycin (50μg/mL), Rifampicin (25μg/mL) | Selection for transformed Agrobacterium [5] |
| Induction Compounds | Acetosyringone (150μM), MES buffer (10mM, pH5.6) | Vir gene induction in Agrobacterium [5] |
| Plant Material | 3-week-old seedlings with 3-4 true leaves | Optimal developmental stage [5] |
| Marker Genes | PDS, CLA1 | Visual silencing phenotype controls [5] [33] |
| Infiltration Solution | 10mM MgCl₂, 10mM MES, 150μM acetosyringone | Agrobacterium resuspension medium [5] |
Application in Functional Genomics
The root wounding-immersion method has proven particularly valuable for functional analysis of disease resistance genes in Solanaceous crops. Researchers successfully applied this technique to silence two disease-resistance genes, SITL5 and SITL6, in tomato cultivar CLN2037E, resulting in significantly decreased disease resistance [5]. This application demonstrates the method's robustness for studying genes involved in plant-pathogen interactions and validates its utility for rapid assessment of candidate resistance genes without stable transformation.
This VIGS approach is especially advantageous for investigating root-pathogen interactions, as it enables efficient gene silencing in root tissues that are naturally encountered by soil-borne pathogens like Ralstonia solanacearum [34]. The method provides a unique opportunity to study early defense responses during root invasion by pathogens, a critical phase in disease development that is difficult to access with above-ground inoculation methods.
Comparative Analysis with Alternative VIGS Methods
While various VIGS inoculation methods have been developed, the root wounding-immersion approach offers distinct advantages for specific applications:
- Superior to Leaf Infiltration: More efficient for whole-plant silencing, especially for root biology studies [5]
- Advantage Over Agrodrench: Higher efficiency and more consistent results across Solanaceous species [5]
- Alternative to Specialized Methods: Unlike leaf tip injection developed for waxy-leaf species like Lycoris [33] or seed vacuum infiltration optimized for sunflowers [8], root wounding-immersion provides broad compatibility across Solanaceae family
- Complementary to Cotyledon Node Transformation: Unlike the soybean TRV-VIGS method using cotyledon node immersion [3], this technique targets established root systems
Troubleshooting and Technical Considerations
Common challenges and solutions:
- Low silencing efficiency: Optimize root wounding extent (approximately 1/3 length critical) and ensure fresh bacterial preparation [5]
- Plant viability post-inoculation: Maintain appropriate bacterial density (OD₆₀₀=0.8) and avoid excessive root damage [5]
- Inconsistent systemic silencing: Standardize plant developmental stage and environmental conditions (temperature, light) [5] [8]
- Contamination issues: Use sterile tools for root excision and maintain clean growth conditions [5]
The root wounding-immersion method represents a significant advancement in VIGS technology, particularly for Solanaceous plants. Its exceptional efficiency (achieving 95-100% silencing rates), scalability for high-throughput studies, and applicability across multiple species position it as a transformative tool for plant functional genomics. By enabling rapid assessment of gene function—including essential genes whose complete knockout would be lethal in stable lines—this approach accelerates gene discovery and characterization. As plant biotechnology increasingly focuses on root-related traits including nutrient uptake, soil-microbe interactions, and resistance to soil-borne pathogens, the root wounding-immersion method provides an indispensable platform for advancing fundamental knowledge and applied crop improvement strategies.
Within the broader scope of Agrobacterium-mediated Virus-Induced Gene Silencing (VIGS) research, the development of efficient and reproducible infection protocols for recalcitrant species represents a significant challenge. VIGS is a powerful reverse genetics tool that leverages the plant's innate antiviral RNA-silencing machinery to knock down the expression of endogenous genes, enabling rapid functional genomics studies without the need for stable transformation [1]. While routinely used in model plants like Nicotiana benthamiana, the application of VIGS to non-model crops often requires extensive optimization of delivery methods [8] [35]. Among these, seed and sprout vacuum infiltration has emerged as a transformative technique, streamlining the VIGS pipeline for species with difficult transformation landscapes, such as sunflower, soybean, and cereals. This Agrobacterium-mediated approach, utilizing vectors based on the Tobacco Rattle Virus (TRV), offers a pathway to whole-plant systemic silencing by targeting plants at early developmental stages, thereby overcoming barriers posed by thick cuticles, dense trichomes, and genotype-specific resistance [3] [36]. This application note details the key protocols, efficiencies, and critical factors for successfully implementing this method, positioning it as a cornerstone for high-throughput gene validation in agricultural research.
Key Principles and Advantages of the Method
The seed vacuum infiltration protocol fundamentally enhances Agrobacterium-mediated VIGS by exploiting the physiological state of germinating seeds or young sprouts. The application of a vacuum followed by its rapid release facilitates the forced entry of Agrobacterium harboring TRV vectors into the intercellular spaces of susceptible young tissues, leading to a more uniform and widespread infection compared to conventional leaf infiltration [8] [36]. A defining feature of this method is its ability to induce whole-plant level gene silencing shortly after germination, enabling functional studies of genes involved in early developmental processes [36].
The core advantages of this system are multi-faceted, as shown in the following comparison of VIGS delivery methods.
Table 1: Comparison of VIGS Delivery Methods
| Method | Key Procedure | Target Species | Key Advantages | Key Limitations |
|---|---|---|---|---|
| Seed Vacuum Infiltration | Vacuum infiltration of peeled seeds followed by co-cultivation [8]. | Sunflower, Wheat, Maize [8] [36] | Bypasses in vitro culture; applicable to early developmental stages; high silencing efficiency [8]. | Requires optimization of vacuum and co-cultivation duration [8]. |
| Sprout Vacuum Infiltration | Vacuum infiltration of germinated seeds or young sprouts [36]. | Wheat, Maize, Tomato [36] | Whole-plant silencing; avoids particle bombardment; uses simple infiltration solution [36]. | Sensitivity of young sprouts to Agrobacterium overgrowth [36]. |
| Cotyledon Node Immersion | Soaking of bisected seed explants in Agrobacterium suspension [3]. | Soybean [3] | Overcomes barriers of leaf trichomes and thick cuticles; high transformation efficiency (>80%) [3]. | Requires sterile tissue culture procedures and explant preparation [3]. |
| Leaf Agroinfiltration | Needleless syringe infiltration of leaves [37]. | Arabidopsis thaliana, N. benthamiana [37] | Simple and fast for amenable species; no specialized equipment needed [37]. | Inefficient for species with thick or hairy leaves; often results in localized silencing [3]. |
Furthermore, the TRV-based vector system is particularly suited for this application. TRV is a bipartite virus, requiring two plasmids for VIGS: pTRV1, which encodes viral replication and movement proteins, and pTRV2, which carries the capsid protein and a cloning site for inserting a fragment of the target plant gene [8] [1]. The sequence-specific silencing is triggered by double-stranded RNA intermediates of the virus, which are processed into small interfering RNAs (siRNAs) that guide the degradation of complementary endogenous mRNA transcripts [1]. The molecular workflow of TRV-based VIGS is outlined below.
Detailed Experimental Protocols
Optimized Seed Vacuum Infiltration for Sunflower
This protocol, adapted from Mardini et al. (2024), provides a robust and simple method for VIGS in sunflower, achieving infection rates of up to 91% in certain genotypes without the need for in vitro recovery [8] [35].
Research Reagent Solutions:
- Plant Material: Sunflower seeds (e.g., genotype 'Smart SM-64B' for high infection rate).
- VIGS Vectors: Agrobacterium tumefaciens strain GV3101 containing pYL192 (TRV1) and pYL156-derived vector (TRV2 with target gene insert, e.g., HaPDS).
- Infiltration Medium: Liquid LB broth with appropriate antibiotics (e.g., kanamycin, gentamicin, rifampicin) and induction additives (e.g., 10 mM MES, 20 μM acetosyringone).
Step-by-Step Procedure:
- Seed Preparation: Remove the seed coats from dry sunflower seeds carefully to avoid damaging the embryos.
- Agrobacterium Preparation: Inoculate single colonies of Agrobacterium containing pTRV1 and pTRV2-derived vectors in separate liquid cultures. Grow overnight at 28°C with shaking until the optical density at 600 nm (OD600) reaches 1.0-1.5. Pellet the cultures by centrifugation and resuspend in infiltration medium (e.g., 10 mM MgCl₂ with acetosyringone) to a final OD600 of 1.0. Mix the pTRV1 and pTRV2 suspensions in a 1:1 ratio.
- Vacuum Infiltration: Submerge the peeled seeds in the mixed Agrobacterium suspension. Apply a vacuum (e.g., 0.15-0.20 bar) for 2-5 minutes. Rapidly release the vacuum to allow the suspension to infiltrate the seeds.
- Co-cultivation: Transfer the infiltrated seeds to a co-cultivation medium (e.g., moist filter paper or peat-perlite mixture). Incubate in the dark at room temperature for 6 hours.
- Plant Growth: Sow the co-cultivated seeds directly into soil. Maintain plants in a greenhouse under controlled conditions (e.g., 22°C, 16-h light/8-h dark photoperiod). Silencing phenotypes, such as photo-bleaching when targeting PDS, typically become visible within 2-4 weeks [8].
Vacuum and Co-cultivation of Germinated Seeds for Monocots
This protocol, established for wheat and maize, demonstrates the cross-species applicability of the method, even in monocot plants traditionally recalcitrant to VIGS [36].
Research Reagent Solutions:
- Infiltration Solution: A specialized solution containing acetosyringone, cysteine, and Tween 20 is critical for success in monocots [36].
- VIGS Vectors: Agrobacterium tumefaciens containing pTRV1 and pTRV2 with a monocot-optimized target gene fragment (e.g., TaPDS or TaMLO for wheat).
Step-by-Step Procedure:
- Seed Germination: Surface-sterilize wheat or maize seeds and allow them to germinate on moist filter paper until the radicle emerges (approximately 1-2 days).
- Agrobacterium Preparation: Prepare Agrobacterium cultures as described in section 3.1, but resuspend the final pellet in the specialized infiltration solution.
- Vacuum Infiltration: Submerge the germinated seeds (sprouts) in the Agrobacterium suspension and apply a vacuum. The optimal duration may require empirical testing.
- Co-cultivation and Planting: Follow a similar co-cultivation and planting strategy as for sunflower. Successful silencing in wheat is evidenced by photo-bleaching or, in the case of TaMLO silencing, enhanced resistance to powdery mildew [36].
Critical Factors and Efficiency Data
The success of seed and sprout VIGS is highly dependent on several biological and technical parameters. A critical factor is plant genotype, which significantly influences both infection rate and the systemic spread of the silencing signal. For instance, in sunflowers, infection percentages varied from 62% to 91% across different genotypes [8] [35]. Furthermore, the developmental stage of the plant is crucial; younger tissues generally exhibit more active spreading of silencing symptoms, and infiltration at the two-to-three leaf stage in Arabidopsis or the germinated seed stage in cereals yields the highest efficiency [37] [36].
The following table summarizes quantitative data on the efficiency of this method across different crop species.
Table 2: Efficiency of Seed/Sprout VIGS Across Crop Species
| Crop Species | Infiltration Method | Target Gene | Key Optimized Parameter | Reported Efficiency | Reference |
|---|---|---|---|---|---|
| Sunflower (Helianthus annuus) | Seed Vacuum | HaPDS | 6 h co-cultivation | Infection: 62-91% (genotype-dependent); Strong photo-bleaching | [8] |
| Soybean (Glycine max) | Cotyledon Node Immersion | GmPDS, GmRpp6907 | 20-30 min immersion | Silencing efficiency: 65-95% | [3] |
| Wheat (Triticum aestivum) | Sprout Vacuum | TaPDS, TaMLO | Novel infiltration solution (Cys, AS, Tween) | Whole-plant photo-bleaching; Powdery mildew resistance | [36] |
| Maize (Zea mays) | Sprout Vacuum | ZmPDS | Novel infiltration solution (Cys, AS, Tween) | Whole-plant photo-bleaching | [36] |
| Abelmoschus manihot L. | Vacuum Infiltration (Leaf) | AmPDS | Two injections at cotyledon stage | ~60% reduction in AmPDS expression | [38] |
It is also important to note that the presence of TRV RNA, as detected by RT-PCR, is not always confined to tissues showing the visible silencing phenotype, indicating that the virus can spread systemically even without observable effects in all regions [8]. This underscores the necessity of always correlating phenotypic observations with molecular analyses of target gene expression, typically via qRT-PCR.
Essential Research Reagent Solutions
A successful VIGS experiment relies on a standardized toolkit of reagents and vectors. The table below details the core components.
Table 3: Essential Research Reagent Solutions for TRV-VIGS
| Item | Function/Description | Examples & Notes |
|---|---|---|
| TRV Vectors | Bipartite viral genome for VIGS; pTRV1 for replication/movement, pTRV2 for target insert. | pYL192 (TRV1), pYL156 (TRV2); pTRV2-PDS is a common positive control [8] [37]. |
| Agrobacterium Strain | Delivers TRV vectors into plant cells via T-DNA transfer. | GV3101 is widely used for its high transformation efficiency and disarmed pathogenicity [8] [3]. |
| Infiltration Medium | Suspension medium for Agrobacterium, often containing inducters of the Vir genes. | 10 mM MgCl₂, 10 mM MES, 150-200 μM Acetosyringone [8] [37]. |
| Antibiotics | Selective pressure to maintain plasmids in bacterial and plant cultures. | Kanamycin (for TRV vectors), Gentamicin & Rifampicin (for Agrobacterium strain selection) [8]. |
| Marker Gene | A visual reporter for successful VIGS, often causing a photobleaching phenotype. | Phytoene Desaturase (PDS); silencing disrupts carotenoid biosynthesis, leading to chlorophyll photo-oxidation [8] [38]. |
The seed and sprout vacuum infiltration method represents a significant advancement in Agrobacterium-mediated VIGS technology, effectively streamlining functional genomics for a growing list of crop species. By providing a simple, high-throughput, and reproducible protocol that circumvents many of the transformation barriers associated with non-model plants, this approach empowers researchers to rapidly characterize gene function. The robust protocols for sunflower, soybean, and monocots like wheat and maize, supported by a clear understanding of critical success factors such as genotype, developmental stage, and optimized infiltration conditions, establish this technique as an indispensable tool in modern crop improvement and plant biology research. Its integration into a broader thesis on VIGS methodologies highlights a pivotal shift towards more accessible and efficient reverse genetics strategies in agriculture.
Virus-Induced Gene Silencing (VIGS) has emerged as a powerful reverse-genetics tool for rapid functional characterization of plant genes. Unlike stable genetic transformation, VIGS enables rapid, transient silencing of target genes, bypassing the need for time-consuming stable transformation procedures that are particularly challenging in soybean [3]. Among various viral vectors, the Tobacco Rattle Virus (TRV) has gained prominence due to its wide host range, mild symptomology, and efficient systemic movement [3]. However, the application of TRV-VIGS in soybean has been relatively limited compared to other plant species, primarily due to challenges associated with infection efficiency through conventional methods [3]. This application note establishes cotyledon node transformation as a robust, efficient Agrobacterium-mediated delivery system for TRV-VIGS in soybean, providing researchers with a validated protocol for rapid gene function analysis.
Key Advantages of the TRV-VIGS System in Soybean
The established TRV-VIGS system utilizing cotyledon node transformation offers several significant advantages over conventional approaches:
- High Efficiency: Achieves systemic silencing with efficiency ranging from 65% to 95%, significantly higher than traditional methods [3]
- Rapid Results: Induces observable phenotypic changes within 15-21 days post-inoculation (dpi), enabling faster functional analysis [3]
- Minimal Viral Symptoms: TRV vectors produce fewer symptoms compared to other viruses, preventing masking of silencing phenotypes [3]
- Versatility: Successfully validated for silencing multiple gene classes, including defense-related genes and visual markers [3]
Quantitative Performance Data
Table 1: Silencing Efficiency of Key Endogenous Genes in Soybean Using TRV-VIGS
| Target Gene | Gene Function | Silencing Efficiency | Observed Phenotype | Time to Phenotype (dpi) |
|---|---|---|---|---|
| GmPDS | Phytoene desaturase (carotenoid biosynthesis) | 65-95% | Photobleaching (white leaves) | 21 |
| GmRpp6907 | Rust resistance gene | 65-95% | Compromised rust immunity | 21-28 |
| GmRPT4 | Defense-related gene | 65-95% | Altered stress response | 21-28 |
Table 2: Comparison of Agrobacterium Infection Methods for TRV Delivery in Soybean
| Infection Method | Infection Efficiency | Advantages | Limitations |
|---|---|---|---|
| Cotyledon Node Transformation | 80-95% | High systemic spread; Suitable for diverse genotypes; Minimal tissue damage | Requires sterile conditions |
| Conventional Misting | Low | Simple application | Limited penetration due to thick cuticle and dense trichomes |
| Direct Injection | Low | Targeted delivery | Tissue damage; Limited systemic spread |
Detailed Experimental Protocol
Vector Construction and Agrobacterium Preparation
TRV Vector Assembly:
- Amplify target gene fragments (300-400 bp) from soybean cDNA using gene-specific primers with incorporated EcoRI and XhoI restriction sites [3]
- Ligate purified PCR products into the pTRV2-GFP vector pre-digested with EcoRI and XhoI restriction enzymes [3]
- Transform ligation products into DH5α competent cells and select positive clones through antibiotic selection and sequencing verification [3]
- Introduce confirmed recombinant plasmids into Agrobacterium tumefaciens strain GV3101 via freeze-thaw transformation [3]
Agrobacterium Culture Preparation:
- Plate transformed Agrobacterium on YEP agar containing 50 mg/L kanamycin and 50 mg/L rifampicin
- Incubate at 28°C for 48 hours until distinct colonies form [39]
- Inoculate single colonies into 100 mL YEP liquid medium with same antibiotics
- Culture at 28°C with shaking at 200 rpm until mid-logarithmic growth phase (OD600 = 0.6-0.8) [39]
- Pellet bacterial cells by centrifugation at 6000 rpm for 8 minutes
- Resuspend in infiltration buffer (10 mM MES, 200 μM acetosyringone, 10 mM MgCl2, 0.03% Silwet-77) to final OD600 of 0.8-1.0 [39]
- Combine equal volumes of TRV1 and TRV2-derived Agrobacterium suspensions
- Incubate mixture at room temperature in darkness for 3 hours to induce virulence gene expression [39]
Plant Material Preparation and Inoculation
Soybean Seed Preparation:
- Surface-sterilize soybean seeds (cv. Tianlong 1) using standard sterilization protocols
- Imbibe sterilized seeds in sterile water until swollen
- longitudinally bisect seeds to obtain half-seed explants with intact cotyledonary nodes [3]
Cotyledon Node Transformation:
- Immerse fresh half-seed explants in prepared Agrobacterium suspension for 20-30 minutes with gentle agitation [3]
- Alternatively, apply vacuum infiltration (0.5 kPa, 5-10 minutes) for enhanced infection efficiency [39]
- Remove excess bacterial suspension by blotting on sterile filter paper
- Transfer inoculated explants to sterile tissue culture containers with vermiculite or appropriate growth medium [3]
- Maintain plants in growth chamber at 22°C with 16/8 hour light/dark cycle and light intensity of 150 μM m⁻² s⁻¹ [39]
Monitoring and Validation
Infection Efficiency Assessment:
- At 4 days post-infection, examine cotyledonary nodes under fluorescence microscope for GFP signals [3]
- Calculate infection efficiency based on percentage of explants showing fluorescence
- For quantitative assessment, analyze GFP expression via qPCR [3]
Silencing Efficiency Evaluation:
- Monitor plants daily for visual phenotypes (photobleaching for GmPDS)
- Harvest tissue from systemically silenced leaves at 21 dpi for molecular analysis
- Quantify transcript abundance of target genes using qRT-PCR with gene-specific primers [39]
- Calculate silencing efficiency based on percentage reduction in transcript levels compared to empty vector controls [3]
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Reagent Solutions for TRV-VIGS in Soybean
| Reagent/Vector | Function/Application | Key Features |
|---|---|---|
| pTRV1 and pTRV2 Vectors | TRV-based silencing system | Binary vector system; Mild symptoms; Efficient systemic movement |
| Agrobacterium tumefaciens GV3101 | Vector delivery | Disarmed strain; Compatible with plant transformation |
| Infiltration Buffer (MES, AS, MgCl2, Silwet-77) | Enhances Agrobacterium infection | Facilitates bacterial entry; Optimized for cotyledon nodes |
| GmPDS Vector Control | Silencing efficiency validation | Visual photobleaching phenotype |
| Acetosyringone | Vir gene inducer | Enhances T-DNA transfer efficiency |
Workflow and Signaling Pathway
TRV-VIGS Soybean Workflow
This diagram illustrates the optimized workflow for efficient TRV-mediated gene silencing in soybean via cotyledon node transformation, highlighting key techniques that enhance silencing efficiency.
The cotyledon node transformation method for TRV-VIGS delivery in soybean represents a significant advancement in functional genomics for this important crop species. This protocol achieves high silencing efficiency (65-95%), enables rapid phenotypic assessment within 3-4 weeks, and provides a reliable alternative to more tedious stable transformation approaches. The systematic optimization of Agrobacterium strain, inoculation method, and plant developmental stage has addressed previous limitations in soybean VIGS applications. This robust platform accelerates gene function validation, facilitating the identification of candidate genes for soybean improvement programs focused on enhanced disease resistance and stress tolerance.
Agrobacterium-mediated Virus-Induced Gene Silencing (VIGS) has emerged as an indispensable reverse genetics tool for functional genomics in ornamental species that are recalcitrant to stable genetic transformation [1] [40]. This technique leverages the plant's innate post-transcriptional gene silencing machinery, utilizing recombinant viral vectors to transiently suppress target gene expression, enabling rapid phenotypic assessment without the need for stable transformation [41] [1]. While conventional infiltration methods like leaf syringe infiltration have proven effective for model plants, ornamental species often present unique anatomical barriers requiring specialized infiltration approaches such as basal plate infiltration and tepal infiltration to achieve efficient gene silencing [40]. These techniques are particularly valuable for studying genes controlling commercially vital ornamental traits, including flower architecture, color patterning, and pigmentation pathways [42] [40]. This protocol details the optimization and application of these specialized infiltration methods within the broader context of Agrobacterium-mediated VIGS infection methodology for ornamental species.
Technical Data and Comparative Analysis
The efficiency of VIGS in ornamental plants is influenced by multiple factors, including infiltration technique, plant developmental stage, and vector selection. The table below summarizes key quantitative data from established VIGS protocols in ornamental species.
Table 1: Quantitative Parameters for VIGS in Ornamental Species
| Plant Species | Infiltration Method | Target Tissue | Silencing Efficiency | Key Optimized Parameters | Phenotype Observation | Citation |
|---|---|---|---|---|---|---|
| Hydrangea macrophylla | Vacuum Infiltration | Tissue-cultured seedlings, cut flowers | 60% (for HmPDS) | 300-400 bp insert fragment; entire seedling infiltration | Photobleaching (30 dpi); Reduced anthocyanin (15 dpi) | [40] |
| Rosa spp. | Vacuum Infiltration | Not specified | 34% (for RhPDS) | Not specified | Photobleaching | [40] |
| Luffa acutangula | Syringe Infiltration | Cotyledons & true leaves | Confirmed by phenotype & qPCR | ~300 bp insert fragment; OD600 0.8-1.0 | Photobleaching; Shorter tendrils | [43] |
| Nicotiana benthamiana, Tomato | Root Wounding-Immersion | Roots | 95-100% (for PDS) | 1/3 root cut; OD600 >1; 30 min immersion | Photobleaching | [5] |
Table 2: Agrobacterium and Vector System Components
| Component | Type/Strain | Function/Characteristic | Example References |
|---|---|---|---|
| Agrobacterium tumefaciens | GV3101, GV1301, C58C1 | Delivery of recombinant VIGS vectors into plant cells. | [4] [43] [5] |
| VIGS Vector | Tobacco Rattle Virus (TRV) | Bipartite system (TRV1, TRV2); broad host range; mild symptoms. | [1] [5] [40] |
| Reporter Gene | Green Fluorescent Protein (GFP) | Visual tracking of infection and silencing spread. | [5] |
| Marker Gene | Phytoene Desaturase (PDS) | Visual silencing phenotype (photobleaching) to validate system efficiency. | [43] [5] [40] |
| Induction Compound | Acetosyringone | Phenolic compound that induces Vir gene expression in Agrobacterium. | [43] [5] |
Application Notes and Protocols
Basal Plate Infiltration for Bulbous Ornamentals
The basal plate, located at the bottom of bulbs, contains meristematic tissue that gives rise to roots and floral structures, making it an ideal entry point for VIGS vectors to ensure systemic silencing [40].
Protocol:
- Plant Material Preparation: Use healthy bulbs from species like lily or tulip. Carefully remove outer scales and surface-sterilize the bulbs with 70% ethanol for 1 minute, followed by a 2% sodium hypochlorite solution for 15 minutes, and rinse thoroughly with sterile distilled water.
- Agrobacterium Preparation: Transform the pTRV1 and pTRV2-derived vectors (e.g., pTRV2-PDS) into Agrobacterium tumefaciens strain GV3101. Inoculate a single colony in LB medium with appropriate antibiotics (e.g., Kanamycin 50 µg/mL, Rifampicin 25 µg/mL) and incubate overnight at 28°C with shaking at 200 rpm. Sub-culture the bacteria and harvest at OD600 = 0.6-0.8 via centrifugation. Resuspend the pellet in an infiltration buffer (10 mM MgCl₂, 10 mM MES, pH 5.6, and 150 µM acetosyringone) to a final OD600 of 0.8-1.0. Incubate the suspension in the dark at room temperature for 3-4 hours [43] [5].
- Infiltration Procedure: Using a sterile 1 mL syringe without a needle, gently apply 20-50 µL of the mixed Agrobacterium suspension (TRV1:TRV2 = 1:1) directly onto the basal plate of the bulb. Apply gentle pressure to allow the solution to permeate the tissue. Alternatively, a brief vacuum infiltration of the entire bulb can be employed [40].
- Post-Infiltration Cultivation: Place the treated bulbs on sterile moist paper in a tray. Cover the tray with a clear lid or plastic wrap to maintain high humidity. Incubate in a growth chamber at 22°C for 2 days in the dark, then transfer to a standard light cycle (16h light/8h dark) at 22-25°C for growth and phenotype observation [40].
Tepal Infiltration for Flower Color Modification
Tepals, the undifferentiated perianth segments found in plants like hydrangea and magnolia, are direct targets for studying pigment biosynthesis pathways [42] [40].
Protocol:
- Plant and Tissue Selection: Select flower buds at developmental stage 2 (as defined for Hydrangea macrophylla) [40]. For magnolia species, buds at the early tepal primordia stage are optimal [42].
- Agrobacterium Preparation: Prepare the Agrobacterium suspension carrying the pTRV2 vector with a target gene fragment (e.g., CHS, F3'5'H for color genes) as described in section 3.1, resuspending to an OD600 of 0.8-1.0 in infiltration buffer [40].
- Infiltration Procedure:
- For Hydrangea sepals (modified tepals): Use a 1 mL needleless syringe. Gently make one or two small holes on the abaxial side of the sepal and infiltrate the Agrobacterium suspension from the abaxial side. Apply the solution until a water-soaked appearance is visible. Alternatively, for cut inflorescences, submerge the entire flower head in the Agrobacterium suspension and apply a vacuum (approximately 0.8 Bar) for 2-3 minutes [40].
- For Magnolia tepals: Adapt the hydrangea protocol by targeting the inner tepal surfaces during the early developmental stage when tissues are more amenable to infiltration [42].
- Post-Infiltration Care and Analysis: After infiltration, maintain the plants or cut flowers in a growth chamber at 22°C with high humidity for 24 hours in the dark, then under normal light conditions. Silencing phenotypes, such as reduced anthocyanin content or color changes, can be observed as early as 15 days post-infection (dpi) [40]. Validate silencing through RT-qPCR analysis of target gene expression and quantify pigment changes using HPLC [40].
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Research Reagents for VIGS Experiments
| Reagent/Material | Function/Application | Specific Example |
|---|---|---|
| TRV-Based VIGS Vectors | Bipartite viral vector system for inducing silencing. | pTRV1, pTRV2 (e.g., pYL156, pYL279) [41] [1]. |
| Agrobacterium Strains | Delivery vehicle for the T-DNA containing the VIGS vector. | GV3101, GV1301, C58C1 [4] [43] [5]. |
| Antibiotics | Selection for bacterial and plasmid maintenance. | Kanamycin (50 µg/mL), Rifampicin (25 µg/mL) [43] [5]. |
| Infiltration Buffer | Medium for Agrobacterium delivery and plant tissue viability. | 10 mM MgCl₂, 10 mM MES (pH 5.6), 150-200 µM Acetosyringone [43] [5]. |
| Marker Gene Constructs | Positive control for silencing efficiency. | TRV2-PDS (Phytoene Desaturase) for photobleaching [5] [40]. |
| Reporter Gene Constructs | Visual tracking of infection spread. | TRV2-GFP (Green Fluorescent Protein) [5]. |
Workflow and Pathway Visualization
The following diagram illustrates the complete experimental workflow for Agrobacterium-mediated VIGS in ornamental plants, from vector construction to phenotypic analysis, highlighting the specialized basal plate and tepal infiltration techniques.
Maximizing Silencing Efficiency: Critical Optimization and Troubleshooting Strategies
Genotype dependency remains a formidable bottleneck in plant biotechnology, hindering the application of advanced genetic tools like Agrobacterium-mediated virus-induced gene silencing (VIGS) in recalcitrant species. This phenomenon describes the dramatic variation in transformation and regeneration efficiency observed among different genotypes within the same species, where optimized protocols work excellently for some cultivars but fail in others [44]. In the context of Agrobacterium-mediated VIGS, genotype dependency affects the entire process from initial infection to systemic viral spread and silencing efficiency, creating significant obstacles for functional genomics research in non-model plants, medicinal species, and woody perennials [8] [44].
The persistence of genotype dependency directly compromises research reproducibility and scalability, particularly affecting species with high heterozygosity, complex genomes, or limited previous biotechnological characterization [45] [46]. Understanding and addressing this challenge is thus critical for advancing plant functional genomics, especially for species with economic, ecological, or medicinal importance that lack efficient stable transformation systems [47]. This application note synthesizes recent advances in conquering genotype dependency through optimized VIGS methodologies, explant selection, and molecular interventions, providing researchers with practical strategies to enhance experimental success across diverse genetic backgrounds.
Table 1: Documented Genotype-Dependent Responses in VIGS and Transformation Studies
| Plant Species | Nature of Genotype Dependency | Efficiency Range | Citation |
|---|---|---|---|
| Sunflower (Helianthus annuus) | VIGS susceptibility and silencing spread | Infection: 62-91% across genotypes | [8] |
| Soybean (Glycine max) | Transformation and editing efficiency | Highly variable; impacts edited mutant recovery | [46] |
| Walnut (Juglans regia) | VIGS efficiency across cultivars | Up to 48% in optimal cultivar 'Xiangling' | [47] |
| Cannabis (Cannabis sativa) | Regeneration capacity from calli | Transgenics in only 1 of 100 tested cultivars | [44] |
| Cotton (Gossypium hirsutum) | Transformation competence | 'Coker' as primarily amenable cultivar | [44] |
Key Challenges and Underlying Mechanisms
Genotype dependency in VIGS experiments manifests through multiple technical challenges that researchers must recognize and address. The regeneration recalcitrance observed in many species stems from an inability of transformed cells to initiate morphological reprogramming and develop into complete plantlets after foreign DNA integration [44]. This is particularly pronounced in perennial, woody species where complex epigenetic regulation, endogenous hormone homeostasis disruptions, and insufficient expression of key developmental regulatory genes create formidable barriers to efficient biotechnology application [45].
The molecular basis of genotype dependency often involves differential expression of developmental regulatory genes (DEV genes) that control cell fate reprogramming and totipotency [45]. Transcriptomic analyses in species like cotton have revealed substantial differences in transcription factor activity and alternative splicing events between highly embryogenic and recalcitrant genotypes [45]. Additional contributing factors include species-specific traits, genotypic variation in response to plant growth regulators, and the physical-chemical properties of culture environments that interact uniquely with different genetic backgrounds [45].
In practical VIGS applications, genotype dependency affects both viral susceptibility and systemic silencing spread. Research in sunflowers demonstrated that while one genotype ('Smart SM-64B') showed high infection rates (91%), it exhibited limited silencing phenotype spread compared to other genotypes, indicating that viral movement and silencing machinery operations are under genetic control [8]. Similarly, legume crops generally display dramatic transformation efficiency variations, with most species showing less than 15% efficiency compared to transformation-susceptible species like tobacco (100%) or rice (51.77%) [46].
Strategic Approaches and Experimental Solutions
Explant Selection and Transformation Method Optimization
Strategic explant selection forms the foundation for overcoming genotype dependency in VIGS applications. Research consistently demonstrates that tissues with high innate regenerative capacity—particularly embryonic and meristematic tissues—significantly enhance transformation efficiency across diverse genotypes [44]. The developmental stage and type of explant used directly influence cellular totipotency and receptivity to Agrobacterium infection, thereby affecting experimental outcomes in genotype-dependent manners.
Advanced VIGS protocols have successfully employed diverse explant types tailored to specific species requirements. In soybean, the cotyledon node has emerged as an effective target for Agrobacterium-mediated VIGS, facilitating systemic viral spread throughout the plant when processed as half-seed explants [4]. For sunflower and other challenging species, seed vacuum infiltration combined with precisely optimized co-cultivation periods (e.g., 6 hours) has demonstrated remarkable success in achieving high infection rates (up to 77%) while eliminating the need for in vitro recovery steps that often introduce genotype-specific complications [8]. The recently developed root wounding-immersion method offers particular promise for recalcitrant species, achieving 95-100% silencing efficiency in Nicotiana benthamiana and tomato by exploiting the strong infectivity of TRV vectors in root systems and meristematic regions [5].
Table 2: Optimized VIGS Delivery Methods for Recalcitrant Species
| Delivery Method | Key Optimization Parameters | Target Species | Reported Efficiency | Citation |
|---|---|---|---|---|
| Seed Vacuum Infiltration | Vacuum application, 6h co-cultivation, no in vitro recovery | Sunflower | Up to 77% infection | [8] |
| Root Wounding-Immersion | 1/3 root removal, 30min immersion, OD600 = 0.8 | Tomato, Nicotiana, Arabidopsis | 95-100% silencing | [5] |
| Cotyledon Node Transformation | Half-seed explants, 20-30min immersion | Soybean | 65-95% silencing | [4] |
| Spray Infiltration | OD600 = 1.1, 5-10 true leaf stage | Walnut | 48% silencing | [47] |
| Agrodrench | Soil application, repeated dosing | Various Solanaceae | Variable by species | [5] |
Molecular and Vector Design Innovations
Vector design and molecular engineering approaches provide powerful strategies to circumvent genotype dependency. Research systematically investigating parameters for effective virus-induced gene silencing has established that insert length significantly influences silencing efficiency, with optimal fragments ranging between 200-1300 base pairs [48]. Additionally, insert position within the target cDNA proves critical, with middle segments demonstrating superior performance compared to 5' or 3' end fragments, while the inclusion of homopolymeric regions (e.g., poly(A/T) tails) substantially reduces silencing effectiveness and should be avoided in construct design [48].
The strategic deployment of developmental regulatory genes (DEV genes) represents a groundbreaking approach to overcoming regeneration recalcitrance. Studies across multiple species demonstrate that ectopic expression of transcription factors such as WUSCHEL (WUS), BABY BOOM (BBM), and GROWTH-REGULATING FACTOR (GRF)-GIF chimeric complexes can dramatically enhance regeneration capacity in previously recalcitrant genotypes [44] [45]. In cassava, overexpression of Arabidopsis GRF5 and the GRF4-GIF1 chimeric complex substantially increased transformation efficiency, while similar approaches have succeeded in Medicago truncatula, Nicotiana tabacum, and various crop species, highlighting the cross-species potential of DEV gene applications [45].
Protocol: Root Wounding-Immersion for Enhanced VIGS Efficiency
The root wounding-immersion method represents a significant advancement for achieving high-efficiency VIGS in genetically diverse material, particularly effective for solanaceous species and Arabidopsis [5]. This protocol exploits the natural susceptibility of root systems to TRV infection and systemic movement.
Materials and Reagents:
- pTRV1 and pTRV2-derived binary vectors
- Agrobacterium strain GV3101
- LB medium with appropriate antibiotics (kanamycin 50 μg/mL, rifampicin 25 μg/mL)
- Infiltration buffer (10 mM MgCl₂, 10 mM MES pH 5.6, 150 μM acetosyringone)
- Healthy 3-4 week old seedlings with 3-4 true leaves
Procedure:
- Agrobacterium Preparation: Inoculate glycerol stocks of Agrobacterium containing pTRV1 and pTRV2-derived vectors into LB medium with appropriate antibiotics. Incubate overnight at 28°C with shaking at 200 rpm.
- Induction Culture: The next day, transfer 50 μL of the culture to 20 mL of fresh LB medium supplemented with 20 μM acetosyringone and 10 mM MES. Grow overnight under the same conditions.
- Bacterial Resuspension: Centrifuge the induced cultures and resuspend in infiltration buffer to a final OD600 of 0.8. Combine pTRV1 and pTRV2 suspensions in a 1:1 ratio. Incubate in the dark at 28°C for 3 hours without shaking.
- Root Treatment: Carefully remove seedlings from growth medium and gently wash roots to remove soil and impurities. Using a sterilized blade, remove approximately one-third of the root system lengthwise to create wounded surfaces.
- Immersion Inoculation: Immerse the wounded roots in the Agrobacterium suspension for exactly 30 minutes, ensuring complete coverage of wounded root surfaces.
- Recovery and Growth: Transplant treated seedlings to fresh growth medium and maintain under standard growth conditions (16h light/8h dark photoperiod at 25°C). Silencing phenotypes typically become visible within 2-4 weeks post-inoculation.
Critical Optimization Parameters:
- Seedling age: 3-4 weeks post-germination (3-4 true leaves stage)
- Bacterial density: OD600 = 0.8
- Immersion duration: 30 minutes
- Root wounding extent: 1/3 of total root length
- Acetosyringone concentration: 150 μM in infiltration buffer
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Research Reagents for Overcoming Genotype Dependency
| Reagent/Category | Specific Examples | Function/Application | Citation |
|---|---|---|---|
| Agrobacterium Strains | GV3101, C58C1, EHA105 | T-DNA delivery; strain choice affects efficiency | [14] [4] [5] |
| VIGS Vectors | TRV1, TRV2 (pYL192, pYL156) | Viral backbone for silencing construct delivery | [8] [4] [47] |
| DEV Gene Constructs | BBM, WUS, GRF4-GIF1, WOX | Enhanced regeneration in recalcitrant genotypes | [44] [45] |
| Reporter Genes | GUS, GFP, PDS | Transformation efficiency assessment & silencing visualization | [14] [4] [47] |
| Plant Growth Regulators | Synthetic cytokinins, auxins | Modulation of organogenesis and callus proliferation | [44] |
| Chemical Inducers | Acetosyringone (150-200 μM) | Vir gene induction for enhanced T-DNA transfer | [4] [5] |
Conquering genotype dependency in recalcitrant species requires an integrated approach combining strategic explant selection, delivery method optimization, and molecular interventions. The documented success of seed vacuum infiltration in sunflowers, root wounding-immersion in solanaceous species, and DEV gene deployment across diverse crops provides researchers with validated strategies to enhance experimental outcomes across genetic backgrounds [8] [45] [5].
Future advancements will likely involve more sophisticated vector design incorporating tissue-specific promoters and synthetic regulatory elements that function independently of host genetic background. Additionally, the integration of emerging technologies like nanoparticle-mediated delivery and CRISPR-based activation of endogenous DEV genes holds promise for further mitigating genotype-dependent responses [44]. As these tools mature, they will progressively democratize plant biotechnology applications across the full spectrum of genetic diversity, ultimately enabling more precise and reproducible functional genomics research in previously recalcitrant species.
Within functional genomics research, Agrobacterium-mediated Virus-Induced Gene Silencing (VIGS) has emerged as a powerful technique for rapid gene function analysis. While vector design and inoculation methods receive significant attention, the post-inoculation environmental conditions are equally critical yet often underestimated factors determining experimental success. The efficiency of VIGS relies on complex biological interactions between the viral vector and host plant, processes highly sensitive to ambient environmental parameters. This application note systematically examines the impact of temperature, light, and humidity on VIGS efficacy, providing evidence-based protocols to optimize these conditions for enhanced silencing efficiency across diverse plant species, thereby supporting more reliable and reproducible results in Agrobacterium-mediated VIGS research.
Environmental Factors Influencing VIGS Efficiency
The efficacy of VIGS is governed by physiological and molecular processes in the host plant that are exquisitely sensitive to environmental conditions. Temperature directly affects viral replication and movement, plant metabolic rates, and RNA silencing machinery activity. Light intensity and quality influence photosynthesis rates and source-sink relationships that determine resource allocation for defense responses and viral replication. Humidity impacts plant stomatal aperture, transpiration rates, and overall water status, thereby affecting Agrobacterium viability and systemic spread of silencing signals.
Table 1: Optimal Environmental Parameters for VIGS in Different Plant Species
| Plant Species | Temperature (°C) | Light Period (hours) | Relative Humidity (%) | Documented Silencing Efficiency | Citation |
|---|---|---|---|---|---|
| Nicotiana benthamiana | 20-22 | 16 | 45-60% | 65-95% | [3] [4] |
| Soybean (Glycine max) | 22 | 18 | ~45% | 65-95% | [3] [4] |
| Sunflower (Helianthus annuus) | 22 | 18 | ~45% | 62-91% (genotype-dependent) | [8] |
| Pepper (Capsicum annuum) | 20-25 | 16 | 50-70% | Not specified | [1] |
Table 2: Impact of Temperature on Specific VIGS Phenotypes
| Gene Silenced | Plant Species | Temperature Condition | Observed Phenotypic Effect | Citation |
|---|---|---|---|---|
| Sr6 (BED-NLR immune receptor) | Wheat | 20°C (permissive) | Effective resistance to Puccinia graminis | [49] |
| Sr6 (BED-NLR immune receptor) | Wheat | 26°C (restrictive) | Susceptibility to Puccinia graminis | [49] |
| Cte-chlH and Cte-PDS | Castilleja tenuiflora | Not specified | 80% vs. 31% photobleaching efficiency between targets | [14] |
Temperature Optimization
Temperature serves as a critical regulator of VIGS efficiency, influencing both viral replication and the plant's RNA interference machinery. Different plant species exhibit specific temperature optima, with 22°C being frequently reported as ideal for VIGS in several species including soybean and sunflower [3] [8]. Perhaps most notably, some resistance genes cloned and validated using VIGS, such as the wheat Sr6 gene, display temperature-sensitive efficacy, providing functional resistance at 20°C but becoming ineffective at 26°C [49]. This temperature dependence underscores the necessity for precise thermal control during post-inoculation periods.
The molecular basis for temperature sensitivity involves its effect on viral replication rates, systemic movement, and the activity of key RNAi pathway components. Lower temperatures typically slow viral replication, potentially reducing phytotoxicity, while higher temperatures may accelerate viral spread but also trigger plant defense responses that limit infection. The documented genotype-environment interactions in VIGS efficiency highlight the need for species-specific and even genotype-specific temperature optimization [8].
Light and Photoperiod Management
Light intensity and photoperiod significantly influence VIGS efficacy through their effects on plant physiology and development. Research indicates that longer photoperiods (16-18 hours of light) are commonly employed in successful VIGS protocols [3] [4] [8]. The light intensity used in these studies typically ranges from 150-300 μmol m⁻² s⁻¹, sufficient to maintain active photosynthesis without inducing light stress.
Light quality and intensity affect VIGS outcomes through multiple mechanisms: (1) regulation of source-sink relationships that determine photoassimilate allocation to defense versus growth; (2) modulation of phytohormone signaling pathways that interact with RNA silencing machinery; and (3) direct effects on viral replication and movement, which often display light-dependent patterns. The successful application of VIGS to characterize genes involved in photoperception and pigment biosynthesis, such as CdCRY1 in Camellia drupifera, further demonstrates the tight interconnection between light signaling and silencing efficiency [9].
Humidity Control
Relative humidity significantly impacts VIGS success by influencing plant water status, stomatal conductance, and Agrobacterium survival during inoculation. Studies consistently maintain relative humidity at approximately 45% during VIGS experiments [3] [8]. This level appears to balance sufficient hydration for plant growth with minimized risk of fungal contamination or excessive leaf wetness that might impede gas exchange.
Humidity affects VIGS through both direct and indirect mechanisms: optimal humidity maintains turgor pressure necessary for cell expansion and systemic signaling, influences stomatal aperture thereby affecting transpirational pull that may facilitate viral movement, and determines the survival rate of Agrobacterium on leaf surfaces during inoculation procedures. Notably, humidity requirements may vary with plant developmental stage and species-specific morphological characteristics, such as leaf thickness and cuticular wax composition.
Experimental Protocols for Environmental Optimization
Standardized Workflow for Environmental Parameter Testing
(Diagram 1: Environmental parameter testing workflow)
Protocol: Temperature Optimization for VIGS
Objective: To determine the optimal temperature regime for maximizing VIGS efficiency in a target plant species.
Materials:
- Plant materials (soybean cultivar Tianlong 1 or target species)
- TRV-based VIGS vectors (pTRV1, pTRV2 with target gene insert)
- Agrobacterium tumefaciens strain GV3101
- Growth chambers with precise temperature control
- RNA extraction kit and qPCR reagents for silencing verification
Method:
- Inoculate plants using standardized Agrobacterium-mediated protocol (cotyledon node immersion for soybean) [3]
- Divide inoculated plants into three temperature groups:
- Group 1: 18°C constant
- Group 2: 22°C constant
- Group 3: 26°C constant
- Maintain consistent light (16h photoperiod) and humidity (45% RH) across all groups
- Monitor plants daily for symptom development (e.g., photobleaching for PDS silencing)
- At 21 days post-inoculation (dpi), harvest tissue from systemic leaves for molecular analysis
- Extract total RNA and perform qPCR to quantify target gene expression normalized to housekeeping genes
- Score phenotypic manifestations (percentage of leaf area showing silencing symptoms)
- Correlate temperature conditions with silencing efficiency metrics
Expected Results: Temperature will significantly impact silencing efficiency, with 22°C expected to yield optimal results for most species. Some systems may display temperature-sensitive silencing as observed with the Sr6 gene in wheat [49].
Protocol: Light and Photoperiod Optimization
Objective: To establish the ideal light intensity and photoperiod for maximal VIGS efficiency.
Materials:
- Programmable growth chambers with adjustable LED lighting
- Light meter for quantifying photosynthetic photon flux density (PPFD)
- Agrobacterium-inoculated plants
Method:
- Prepare plants as described in Section 3.2 and maintain at optimal temperature (22°C)
- Divide plants into three photoperiod groups:
- Group A: 12h light/12h dark (short day)
- Group B: 16h light/8h dark (long day)
- Group C: 18h light/6h dark (extended long day)
- Maintain light intensity at 200 μmol m⁻² s⁻¹ PPFD across all groups
- For light intensity testing, establish additional groups with varying PPFD (100, 200, 300 μmol m⁻² s⁻¹) at optimal photoperiod
- Monitor silencing progression and quantify efficiency as in Section 3.2
- Document any light-related phenotypic differences beyond silencing symptoms
Expected Results: Longer photoperiods (16-18h) typically enhance VIGS efficiency by supporting increased photosynthetic activity and assimilate production necessary for viral spread and silencing signal amplification [3] [8].
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Research Reagent Solutions for VIGS Optimization
| Reagent/Resource | Function in VIGS | Example Specifications | Citation |
|---|---|---|---|
| TRV Vectors (pTRV1, pTRV2) | Bipartite viral vector system for silencing | pYL192 (TRV1), pYL156 (TRV2) with MCS | [8] [50] |
| Agrobacterium tumefaciens | Vector delivery into plant cells | Strain GV3101 with helper plasmids | [3] [8] |
| Marker Gene Constructs | Silencing efficiency validation | PDS (photobleaching), CHLI (yellowing) | [14] [50] |
| Acetosyringone | Vir gene inducer in Agrobacterium | 100-200 μM in infiltration medium | [9] |
| MES Buffer | pH stabilization in Agrobacterium culture | 10 mM, pH 5.6 | [9] |
| Antibiotics | Selection of transformed Agrobacterium | Kanamycin (50 μg/mL), Rifampicin (50 μg/mL) | [8] [9] |
Environmental parameters—temperature, light, and humidity—are not merely supportive growth conditions but active determinants of VIGS success. The optimal ranges identified through systematic testing (22°C temperature, 16-18h photoperiod, and 45% RH) provide a foundational starting point for researchers implementing Agrobacterium-mediated VIGS across diverse plant systems. The documented genotype-specific responses to these environmental factors necessitate empirical optimization for each new plant system. Furthermore, the discovery of temperature-sensitive VIGS phenotypes, as exemplified by the Sr6 gene characterization, reveals that environmental fine-tuning can unlock functional analysis of previously recalcitrant genes. By integrating these evidence-based environmental optimization strategies, researchers can significantly enhance the efficiency, reproducibility, and applicability of VIGS in functional genomics studies.
In Agrobacterium-mediated Virus-Induced Gene Silencing (VIGS), the precision of agroinoculum preparation—encompassing optimal bacterial density and specific cultivation conditions—is a critical determinant of experimental success. This protocol details standardized methodologies for achieving high-efficiency VIGS across diverse plant species, providing researchers with a framework for reliable gene silencing phenotypes essential for functional genomics research in plant biology and drug discovery pathways.
Quantitative Data on Agroinoculum Parameters
Table 1: Optimized Bacterial Density and Cultivation Conditions for Agrobacterium-mediated VIGS
| Plant Species | Optimal OD₆₀₀ | Agrobacterium Strain | Incubation Conditions | Silencing Efficiency | Citation |
|---|---|---|---|---|---|
| Soybean (Glycine max) | 0.8 | GV3101 | 3 hours, 28°C, dark | 65% - 95% | [4] |
| Sunflower (Helianthus annuus) | N/R | GV3101 | 6 hours co-cultivation | Up to 91% (genotype-dependent) | [8] |
| Castilleja tenuiflora | N/R | C58C1 | Specific to "Tc-AII" method | 65% transformation efficiency | [14] |
| Nicotiana benthamiana | >1.0 (pre-culture), 0.8 (final) | GV1301 | 3 hours, 28°C, dark | ~100% (PDS silencing) | [5] |
| Tomato (Solanum lycopersicum) | >1.0 (pre-culture), 0.8 (final) | GV1301 | 3 hours, 28°C, dark | 95% - 100% | [5] |
N/R: Not explicitly reported in the source material.
Detailed Experimental Protocols
Agrobacterium Culture Preparation for VIGS
This core protocol is foundational for methods described in subsequent sections [4] [5].
- Plate Streaking: Streak frozen glycerol stocks of transformed Agrobacterium (e.g., carrying pTRV1 and pTRV2-derived vectors) onto LB-agar plates containing appropriate antibiotics (e.g., kanamycin 50 µg/mL, rifampicin 25 µg/mL). Incubate at 28°C for 1.5 - 2 days [4] [8].
- Starter Culture: Select two random single colonies and inoculate into 20 mL of LB broth with the same antibiotics. Shake overnight (~16 hours) at 28°C and 200 rpm [5].
- Secondary Culture: The next day, add 50 µL of the starter culture to 20 mL of fresh LB medium supplemented with antibiotics, 20 µM acetosyringone, and 10 mM MES. Culture overnight at 28°C and 200 rpm [5].
- Induction: Harvest the bacterial cells by centrifugation and resuspend in an infiltration solution (10 mM MgCl₂, 10 mM MES, pH 5.6, 150 µM acetosyringone) to a final OD₆₀₀ = 0.8 [4] [5].
- Acetosyringone Induction: Incubate the resuspended Agrobacterium culture in the dark at 28°C for 3-4 hours with gentle agitation to induce virulence genes [5].
- Combining Strains: Mix the induced pTRV1 and pTRV2 (with target insert) cultures in a 1:1 ratio immediately before plant inoculation [5].
Plant Inoculation Methods
Cotyledon Node Immersion for Soybean
This method overcomes challenges posed by soybean's thick cuticle and dense trichomes [4].
- Plant Material: Surface-sterilize soybean seeds and imbibe in sterile water until swollen. Bisect the seeds longitudinally to create half-seed explants.
- Inoculation: Immerse fresh explants in the prepared Agrobacterium suspension (OD₆₀₀ = 0.8) for 20-30 minutes with gentle agitation.
- Co-cultivation & Regeneration: Transfer the infected explants to regeneration media and standard growth conditions.
- Efficiency Validation: On day 4 post-infection, examine the hypocotyl under a fluorescence microscope for GFP signals to confirm successful infection, which can exceed 80% efficiency [4].
Seed Vacuum Infiltration for Sunflower
A robust protocol optimized for recalcitrant species like sunflower that eliminates the need for in vitro recovery [8].
- Seed Preparation: Partially remove the seed coat (peeling) to enhance infiltration.
- Vacuum Infiltration: Submerge seeds in the Agrobacterium suspension and apply a vacuum for a specified duration.
- Co-cultivation: Subject seeds to 6 hours of co-cultivation post-infiltration.
- Planting: Sow seeds directly into soil or growth medium without surface sterilization. Infection rates can reach 77-91%, depending on genotype [8].
Root Wounding-Immersion for Solanaceous Species
This highly efficient method is suitable for high-throughput functional screening [5].
- Plant Preparation: Grow seedlings for 3-4 weeks until they have 3-4 true leaves. Gently remove plants from soil and wash roots with pure water.
- Wounding: Using a disinfected blade, cut approximately one-third of the root length longitudinally to create a wound site.
- Immersion: Immerse the wounded roots in the Agrobacterium suspension (OD₆₀₀ = 0.8) for 30 minutes.
- Re-potting: Transplant treated seedlings into fresh soil. This method achieves a 95-100% silencing rate for PDS in N. benthamiana and tomato [5].
The following diagram illustrates the logical workflow and key decision points for selecting an appropriate VIGS inoculation method based on plant species and research goals.
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for Agroinoculum Preparation and VIGS
| Reagent / Material | Function / Role | Example Usage & Notes |
|---|---|---|
| Agrobacterium tumefaciens Strains | Delivery vector for TRV-based VIGS constructs. | GV3101, GV1301, and C58C1 are common. Strain choice can affect efficiency [14] [4] [5]. |
| TRV Vectors (pTRV1, pTRV2) | Viral backbone for inducing silencing. | pTRV1 contains replication genes. pTRV2 carries the target gene fragment for silencing [4] [8]. |
| Acetosyringone | Phenolic compound that induces Agrobacterium virulence genes. | Critical for efficient T-DNA transfer. Used at 150-200 µM in infiltration buffers [5]. |
| MES Buffer | Maintains stable pH in the infiltration buffer. | Used at 10 mM in culture and infiltration solutions to stabilize pH at ~5.6 [5]. |
| Antibiotics | Selective pressure for plasmid maintenance and contamination control. | Kanamycin (50 µg/mL) for TRV vectors; Rifampicin (25-50 µg/mL) for Agrobacterium strain selection [8] [5]. |
| Infiltration Buffer (MgCl₂/MES) | Solvent for the final Agrobacterium suspension. | Provides osmotic and ionic balance for bacterial viability during plant infection [5]. |
Critical Factors Influencing Agroinoculum Success
- Bacterial Growth Phase: Utilizing bacteria in the mid-exponential growth phase is crucial for optimal virulence. The optical density (OD₆₀₀) serves as a key proxy for this, with an OD₆₀₀ of 0.8 being frequently optimal for the final inoculation suspension [4] [5].
- Genotype Dependence: VIGS efficiency exhibits significant genotype-dependent variation. In sunflower, infection rates ranged from 62% to 91% across different cultivars, highlighting the need for protocol optimization for specific plant lines [8].
- Co-cultivation Conditions: Post-inoculation co-cultivation periods are critical. A 6-hour co-cultivation was identified as optimal for sunflower VIGS, while other species may require different durations [8].
- Plant Growth Environment: Environmental factors post-inoculation significantly impact silencing spread and persistence. Studies indicate that lower temperatures and humidity levels can enhance VIGS efficiency [5].
The precision of agroinoculum preparation, underscored by optimal bacterial density and meticulously controlled cultivation conditions, is the cornerstone of reproducible and high-efficiency VIGS. The protocols and data summarized here provide a actionable roadmap for researchers to implement and further refine these techniques, thereby accelerating functional gene discovery in plant systems.
Within the framework of research on Agrobacterium-mediated Virus-Induced Gene Silencing (VIGS), a fundamental challenge is the host plant's RNA silencing machinery, an adaptive antiviral defense system that rapidly degrades foreign genetic material [51]. This immune response significantly limits the accumulation and persistence of recombinant viral vectors, thereby constraining the efficacy of protein expression and functional genomics studies. Viral Suppressors of RNA Silencing (VSRs) are powerful counter-defense proteins encoded by plant viruses to neutralize this host immunity [52] [53]. This application note details the strategic engineering of expression vectors with heterologous VSRs, providing validated protocols to enhance recombinant protein yields and silencing efficiency in plant systems, with direct relevance to VIGS methodologies.
Recent systematic optimization of Potato virus X (PVX)-derived vectors demonstrates that co-expression of heterologous VSRs can dramatically enhance the accumulation of recombinant proteins.
Table 1: Enhancement of Recombinant Protein Expression via VSRs in PVX Vectors
| VSR Incorporated | Source Virus | Target Protein | Max. Accumulation (mg/g FW) | Fold-Increase vs. Parental PVX | Primary Mechanism of Action |
|---|---|---|---|---|---|
| NSs | Tomato zonate spot virus (TZSV) | GFP | 0.50 | ~3.8-fold | Targets SGS3 for degradation [54] |
| Vaccine Antigen (S2) | 0.017 | >100-fold | |||
| Vaccine Antigen (VP1) | 0.016 | >100-fold | |||
| P38 | Turnip crinkle virus (TCV) | GFP | ~0.50 (High) | ~3-4 fold | Binds to AGO1 via GW/WG motifs [52] [54] |
| P19 | Tomato bushy stunt virus (TBSV) | GFP | Moderate | ~3-4 fold | Sequesters siRNA duplexes [54] |
A critical engineering insight involves transcriptional interference. Initial constructs with VSR cassettes in the same orientation as the target gene showed reduced expression. Simply reversing the orientation of the VSR cassette relative to the target gene markedly improved the expression of both the VSR and the recombinant protein [54].
Detailed Protocol: Engineering a VSR-Enhanced PVX Vector
The following protocol is adapted from Jung et al., outlining the steps to create and evaluate a deconstructed PVX vector (pP2 backbone) harboring a heterologous VSR [55] [54].
Reagents and Equipment
- Plant Material: Nicotiana benthamiana plants, 3-4 weeks old.
- Vector Backbone: Deconstructed PVX vector pP2 (lacking TGB and CP) [54].
- VSR Source: Coding sequences for NSs, P38, or P19.
- Agrobacterium Strain: GV3101.
- Promoters/Terminators: Cauliflower Mosaic Virus (CaMV) 35S promoter, nopaline synthase (NOS) terminator.
- Infiltration Buffer: 10 mM MES, 10 mM MgCl₂, 150 µM acetosyringone, pH 5.6.
Step-by-Step Procedure
Vector Construction: a. Clone your gene of interest (GOI; e.g., GFP, vaccine antigen) into the pP2 PVX backbone under the control of a suitable subgenomic promoter. b. Synthesize or clone the selected VSR (e.g., NSs) coding sequence into an expression cassette containing the CaMV 35S promoter and NOS terminator. c. Insert this entire VSR expression cassette in the reverse orientation downstream of the NOS terminator of the GOI in the pP2 vector, creating the final construct (e.g., pP3NSs:GOI) [54].
Agrobacterium Preparation: a. Transform the final plasmid into Agrobacterium tumefaciens strain GV3101 via electroporation. b. Plate on selective media and incubate at 28°C for 2 days. c. Pick a single colony and inoculate a starter culture (5 mL LB with appropriate antibiotics). Grow overnight at 28°C with shaking. d. Sub-culture into a fresh medium (1:100 dilution) and grow until OD₆₀₀ reaches approximately 0.8. e. Pellet the cells by centrifugation (5000xg for 10 min) and resuspend in infiltration buffer to a final OD₆₀₀ of 0.5. f. Incubate the resuspended culture at room temperature for 3-4 hours with gentle agitation before infiltration.
Plant Infiltration and Analysis: a. Using a needleless syringe, infiltrate the Agrobacterium suspension into the abaxial air spaces of fully expanded N. benthamiana leaves. b. Maintain infiltrated plants under standard growth conditions (e.g., 22-25°C, 16-h light/8-h dark photoperiod). c. Monitor protein expression 3-7 days post-infiltration (dpi): - Visual Assessment: For fluorescent proteins like GFP, use UV illumination. - Protein Quantification: Perform Western blot analysis or ELISA on leaf tissue extracts. - Transcript Analysis: Use qRT-PCR to verify VSR and GOI expression levels.
Signaling Pathways and Experimental Workflow
The following diagrams illustrate the plant antiviral RNA silencing pathway and the strategic workflow for enhancing vector efficacy using VSRs.
Diagram 1: RNA Silencing and VSR Mechanisms. The plant antiviral pathway (green) is targeted at multiple points by specific VSRs (blue) to suppress defense.
Diagram 2: VSR Vector Engineering Workflow. The key step of reverse orientation insertion of the VSR cassette is highlighted for its critical impact on final protein yield.
The Scientist's Toolkit: Essential Research Reagents
Table 2: Key Reagents for VSR-Enhanced Vector Engineering
| Reagent / Tool | Function & Utility in VIGS/VSR Research | Example Source / Strain |
|---|---|---|
| Deconstructed Viral Vectors | Minimal vectors retaining replication elements but lacking movement/coat proteins, reducing pathogenicity and increasing insert capacity [54]. | PVX (pP1, pP2 backbones) |
| Heterologous VSRs | Proteins from diverse viruses used to potently suppress host RNA silencing in trans, boosting recombinant protein yield [55] [54]. | P19 (TBSV), P38 (TCV), NSs (TZSV) |
| Nicotiana benthamiana | A model plant host in molecular farming due to its susceptibility to a wide range of viruses and ease of agroinfiltration [55] [54]. | |
| Agrobacterium tumefaciens | Standard workhorse for delivering DNA constructs into plant cells via transient transformation (agroinfiltration) [54] [8]. | GV3101, C58C1, LBA4404 |
| TRV-based VIGS Vectors | RNA virus vectors widely adopted for efficient and systemic gene silencing in a broad range of plant species [14] [8]. | pYL192 (TRV1), pYL156 (TRV2) |
Within the broader context of optimizing Agrobacterium-mediated Virus-Induced Gene Silencing (VIGS) infection methods, the strategic design of the insert fragment is a critical determinant of experimental success. VIGS functions as a powerful reverse genetics tool, leveraging the plant's innate RNA interference machinery to silence target genes by degrading homologous mRNA sequences [31]. The efficacy of this process hinges on the selection of a target gene fragment that the viral vector will carry. This application note provides a consolidated protocol of evidence-based rules for selecting target fragments with high silencing potential, incorporating quantitative data and detailed methodologies to assist researchers in designing effective VIGS constructs.
Fragment Design Parameters and Experimental Evidence
Core Design Principles
The design of the insert fragment involves several key parameters that directly influence the efficiency and specificity of gene silencing. Adherence to the following rules is essential for maximizing silencing potential:
Optimal Fragment Length: Research across multiple plant species consistently demonstrates that effective silencing fragments typically range from 200 to 400 base pairs (bp) [9] [39]. For instance, studies in Camellia drupifera successfully utilized fragments of 200-300 bp [9], while work in Atriplex canescens selected fragments between 300 and 400 bp [39]. A length within this window is sufficient to trigger a robust RNAi response while minimizing the risk of recombination within the viral vector.
Sequence Specificity and Conservation: The selected fragment must exhibit high specificity for the target gene to prevent unintended off-target silencing. This is verified by using the SGN VIGS Tool (https://vigs.solgenomics.net/) to screen potential fragments [9] [39] and performing a BLAST analysis against the host plant's genome or transcriptome. The chosen fragment should share less than 40% similarity to other non-target genes to ensure specific silencing [9]. Furthermore, for genes belonging to multi-gene families, target highly conserved domains to silence multiple homologous genes, or unique, non-conserved regions to achieve gene-specific knockdown [39].
Fragment Position within the Coding Sequence (CDS): Empirical evidence suggests that fragments from different regions of the CDS can yield varying silencing efficiencies. Research in A. canescens systematically tested three fragments from the 5' end, central region, and 3' end of the AcPDS gene [39]. While multiple positions can be effective, the central region is often preferred as it may be less prone to contain regulatory sequences that could compromise efficiency.
Table 1: Summary of Fragment Design Parameters from Case Studies
| Plant Species | Target Gene | Fragment Length (bp) | Fragment Position | Validated Tool | Citation |
|---|---|---|---|---|---|
| Camellia drupifera | CdCRY1, CdLAC15 | 200-300 | Specific region selected after analysis | SGN VIGS Tool, BLAST | [9] |
| Atriplex canescens | AcPDS | 311, 751, 1221 | 5' end, Central, 3' end | SGN VIGS Tool, BLAST | [39] |
| Luffa acutangula | LaPDS, LaTEN | ~300 | Not Specified | Primer design based on orthologs | [56] |
| Soybean (Glycine max) | GmPDS, GmRpp6907 | Specific amplicon size confirmed via electrophoresis | Not Specified | Primer design and sequencing | [3] |
Experimental Workflow for Fragment Selection and Validation
The following workflow outlines the key steps from initial gene selection to final experimental validation. This process integrates bioinformatic screening with molecular biology techniques to ensure the selection of a high-potency silencing fragment.
Detailed Protocol for Vector Construction
This section provides a step-by-step methodology for cloning the selected target fragment into a VIGS vector, based on established protocols [56] [3] [39].
Primer Design and PCR Amplification
Primer Design: Design gene-specific primers that flank the selected ~300 bp target fragment. The forward and reverse primers must include 5' overhangs containing appropriate restriction enzyme sites (e.g.,
EcoRIandXhoI, orEcoRIandBamHI) for directional cloning [3] [39].- Example Primer Structure:
taaggttaccGAATTCTCTCCGCGTCCTCTAAAAC[3], where the lowercase sequence is the homology arm and the uppercase sequence is theEcoRIsite (GAATTC).
- Example Primer Structure:
PCR Amplification:
- Reaction Setup: Use a high-fidelity DNA polymerase (e.g., Phanta Max Super-Fidelity) in a 50 µL reaction volume. The typical reaction mixture includes:
- 25 µL of 2x Master Mix
- 2 µL of cDNA template
- 2 µL each of forward and reverse primer (10 µM)
- 1 µL of dNTP Mix
- Nuclease-free water to 50 µL [56].
- Thermocycling Conditions:
- Verification: Analyze the PCR product by electrophoresis on a 1.0% agarose gel and purify the band of the correct size using a gel extraction kit [56].
- Reaction Setup: Use a high-fidelity DNA polymerase (e.g., Phanta Max Super-Fidelity) in a 50 µL reaction volume. The typical reaction mixture includes:
Cloning and Agrobacterium Transformation
- Digestion and Ligation: Digest both the purified PCR product and the pTRV2 vector with the selected restriction enzymes (e.g.,
EcoRIandXhoI). Purify the digested products and ligate them using a ligation enzyme mix (e.g., Hieff Clone Enzyme Premix) [3]. Incubate the ligation reaction at 25°C for 30 minutes. - Transformation into E. coli: Transform the ligation product into competent E. coli cells like DH5α. Spread on LB agar plates containing the appropriate antibiotic (e.g., Kanamycin, 50 mg/L) and incubate overnight at 37°C [56] [9].
- Colony Screening and Sequencing: Select individual colonies and screen for positive clones via colony PCR using universal primers for the VIGS vector (e.g., pTRV2 universal primer) [56]. Inoculate positive cultures for plasmid extraction and send the recombinant plasmids for Sanger sequencing to confirm the correct insert sequence and orientation.
- Agrobacterium Transformation: Introduce the sequence-verified plasmid into Agrobacterium tumefaciens strain GV3101 via freeze-thaw transformation [39]. Plate the transformed Agrobacterium on YEP agar containing Kanamycin (50 mg/L) and Rifampicin (25-50 mg/L) and incubate at 28°C for 48 hours [56] [39].
Table 2: Key Research Reagent Solutions for VIGS Vector Construction
| Reagent / Material | Function / Purpose | Example Specifications / Notes |
|---|---|---|
| pTRV1 & pTRV2 Vectors | Binary VIGS vector system; TRV1 encodes viral replication proteins, TRV2 carries the target insert. | The pTRV2 vector is modified for cloning (e.g., pNC-TRV2) [9]. |
| High-Fidelity DNA Polymerase | PCR amplification of target fragment with high accuracy. | E.g., Phanta Max Super-Fidelity [56] or Hieff Robust PCR Master Mix [9]. |
| Restriction Enzymes | Enzymatic digestion of vector and insert for directional cloning. | E.g., EcoRI, XhoI, BamHI [3] [39]. |
| Cloning Kit | Homologous recombination or restriction-ligation based assembly of vector and insert. | E.g., Hieff Clone Kit [56] or Nimble Cloning Kit [9]. |
| E. coli Strain DH5α | Propagation and amplification of recombinant plasmid DNA. | Standard cloning strain [56] [9]. |
| Agrobacterium tumefaciens GV3101 | Delivery of the recombinant VIGS vector into plant cells. | A disarmed strain commonly used for agroinfiltration [56] [3] [39]. |
| SGN VIGS Tool | Online bioinformatic tool for predicting optimal target fragments to maximize silencing efficiency. | https://vigs.solgenomics.net/ [9] [39] |
Concluding Remarks
The meticulous selection of the target fragment is a foundational step in designing a potent VIGS experiment. By adhering to the outlined rules—prioritizing a 200-400 bp fragment that is specific to the target gene and verified through bioinformatic tools—researchers can significantly enhance the probability of achieving high-efficiency gene silencing. The standardized protocols for vector construction and validation ensure the reliability and reproducibility of the VIGS system, thereby accelerating functional genomics studies, particularly in non-model plant species that are recalcitrant to stable transformation.
Ensuring Robust Results: Phenotypic and Molecular Validation of VIGS
Virus-Induced Gene Silencing (VIGS) has established itself as a cornerstone technique in plant functional genomics, enabling rapid, transient knockdown of gene expression without the need for stable transformation. While the photobleaching phenotype resulting from silencing the phytoene desaturase (PDS) gene serves as a convenient visual marker for system optimization, the true power of VIGS is realized in its application to characterize genes controlling agronomically vital traits such as disease resistance and developmental timing [1]. This Application Note details advanced protocols and validation methodologies for employing TRV (Tobacco Rattle Virus)-based VIGS to functionally characterize these critical genes, moving beyond simple visual markers to robust phenotypic and molecular confirmation.
The foundational principle of VIGS leverages the plant's innate post-transcriptional gene silencing (PTGS) machinery. Recombinant viral vectors, typically delivered via Agrobacterium tumefaciens, carry fragments of the plant's endogenous target gene. As the virus replicates and moves systemically, double-stranded RNA intermediates are processed into small interfering RNAs (siRNAs) by Dicer-like enzymes. These siRNAs are incorporated into the RNA-induced silencing complex (RISC), which guides sequence-specific cleavage and degradation of complementary endogenous mRNA transcripts, ultimately leading to a loss-of-function phenotype [1]. This mechanism provides a powerful platform for reverse genetics in a wide range of plant species, especially those recalcitrant to stable transformation.
The utility of VIGS extends far beyond model plants to encompass a diverse array of horticultural and agronomic species. The table below summarizes recent, quantitative evidence of successful VIGS implementation for validating genes involved in development and disease resistance.
Table 1: Quantitative Evidence of VIGS in Functional Gene Validation
| Plant Species | Target Gene | Gene Function | Silencing Efficiency/Impact | Key Phenotypic Outcome | Citation |
|---|---|---|---|---|---|
| Gossypium hirsutum (Cotton) | GhDnaJ316 |
Floral development regulator | Not specified | Accelerated floral transition by 7.7 days (budding) and 9.7 days (flowering) | [57] |
| Glycine max (Soybean) | GmRpp6907 |
Rust resistance | Silencing efficiency: 65-95% | Compromised rust immunity, increased sporulation | [3] |
| Glycine max (Soybean) | GmRPT4 |
Defense-related | Silencing efficiency: 65-95% | Induced significant phenotypic changes | [3] |
| Triticum aestivum (Wheat) | Sr6 (BED-NLR) |
Stem rust resistance | Validated via susceptibility | Increased susceptibility to Puccinia graminis | [49] |
| Gossypium arboreum | GaNBS (OG2) |
Disease resistance (NBS-domain) | Validated via virus tittering | Demonstrated putative role in virus accumulation | [58] |
| Iris japonica | IjPDS (Optimization) |
Carotenoid biosynthesis | 36.67% (optimal in 1-yr seedlings) | Distinct photobleaching, system established | [20] |
| Camellia drupifera | CdCRY1 |
Pericarp pigmentation | ~69.80% (at early capsule stage) | Fading phenotype in exocarps | [9] |
| Camellia drupifera | CdLAC15 |
Pericarp pigmentation | ~90.91% (at mid capsule stage) | Fading phenotype in mesocarps | [9] |
Experimental Protocols for Robust Gene Validation
TRV Vector Construction andAgrobacteriumPreparation
The initial and most critical step is the design and cloning of the VIGS construct. A ~200-500 bp fragment from the 3' untranslated region (UTR) or a non-conserved coding region of the target gene is recommended to ensure specificity and minimize off-target silencing [1] [9].
Detailed Protocol:
- Target Fragment Selection: Use tools like the SGN VIGS Tool to screen for specific sequences. Perform a BLAST search against the plant's genome to ensure the selected fragment has less than 40% similarity to other non-target genes [9].
- PCR Amplification: Amplify the target fragment from cDNA using high-fidelity DNA polymerase. Primers should be designed with appropriate restriction enzyme sites (e.g., EcoRI and XhoI) for directional cloning.
- Example Primer Design for
GmPDS[3]:- Forward: 5'-taaggttaccGAATTCTCTCCGCGTCCTCTAAAAC-3'
- Reverse: 5'-atgcccgggcCTCGAGTCCAGGCTTATTTGGCATAGC-3' (Restriction sites are in bold: EcoRI in forward, XhoI in reverse)
- Example Primer Design for
- Ligation and Transformation: Ligate the purified, digested PCR product into the similarly digested pTRV2 vector. Transform the ligation product into E. coli DH5α competent cells. Select positive clones on kanamycin plates and verify the insert by Sanger sequencing [3] [9].
- Agrobacterium Transformation: Introduce the verified recombinant pTRV2 plasmid and the helper pTRV1 plasmid into Agrobacterium tumefaciens strain GV3101 via electroporation or freeze-thaw transformation.
- Agroinoculum Preparation:
- Inoculate a single colony of Agrobacterium containing pTRV1 or pTRV2-derived vectors into YEB liquid medium supplemented with kanamycin (50 µg/mL) and rifampicin (50 µg/mL).
- Incubate at 28°C with shaking (200-240 rpm) for 24-48 hours.
- Sub-culture the bacteria into fresh induction medium (e.g., YEB with 10 mM MES, 20 µM acetosyringone) and grow until the OD₆₀₀ reaches 0.9-1.0.
- Pellet the cells by centrifugation (5000 rpm for 15 min) and resuspend in an infiltration buffer (10 mM MgCl₂, 10 mM MES, 200 µM acetosyringone) to a final OD₆₀₀ of 1.0-2.0.
- Incubate the resuspended culture in the dark at room temperature for 3-6 hours before use [3] [9].
Plant Inoculation: Optimized Methods for Challenging Tissues
The choice of inoculation method is paramount and depends heavily on the plant species and tissue type.
Table 2: Optimized Agroinfiltration Methods for Different Plant Tissues
| Method | Procedure | Target Species/Tissues | Key Advantage |
|---|---|---|---|
| Cotyledon Node Immersion | Bisect sterilized, swollen seeds; immerse fresh cotyledon node explants in Agrobacterium suspension for 20-30 min [3]. | Soybean; tissues with thick cuticles and dense trichomes. | Achieved >80% infection efficiency, up to 95% in some cultivars. |
| Pericarp Cutting Immersion | Make shallow incisions on the fruit pericarp and immerse the entire fruit in the Agrobacterium suspension under vacuum infiltration [9]. | Recalcitrant, lignified capsules of Camellia drupifera. | Infiltration efficiency of ~93.94% for pericarp pigment genes. |
| Shoot Infusion/Peduncle Injection | Use a needleless syringe to inject Agrobacterium culture into the shoot apex or the fruit peduncle [9]. | Tea oil camellia, other woody plants. | Facilitates systemic viral movement from the plant's apical region. |
Phenotypic and Molecular Validation of Silencing
Confirming successful gene silencing requires a combination of phenotypic assessment and molecular analysis.
Phenotypic Monitoring:
- Developmental Genes: For genes like
GhDnaJ316, carefully monitor and record the timing of key developmental stages, such as days to budding and days to flowering, comparing silenced plants to empty vector controls [57]. - Disease Resistance Genes: Inoculate silenced plants with the relevant pathogen (e.g., Puccinia graminis for
Sr6in wheat). Assess the loss of resistance by quantifying disease symptoms, fungal sporulation, or infection type compared to resistant controls [49].
- Developmental Genes: For genes like
Molecular Verification:
- Quantitative PCR (qPCR): This is the standard method to quantify the reduction in target gene mRNA levels. Total RNA should be extracted from silenced tissues and cDNA synthesized for qPCR analysis. Use stably expressed reference genes (e.g., Actin, UBQ) for normalization. A successful silencing experiment typically shows a >70% reduction in transcript abundance [20].
- Virus Tracking with GFP: Using a pTRV2-GFP vector allows for the visualization of viral spread. Tissues can be observed under a fluorescence microscope or laser confocal microscope to confirm the presence of the virus in the areas where phenotypic changes are observed [20] [3].
- Downstream Marker Analysis: For defense-related genes, validate silencing by examining the expression of known downstream markers. For example, silencing a resistance gene like
Sr6should lead to the downregulation of defense-responsive genes upon pathogen challenge [49].
Diagram Title: VIGS Workflow for Non-PDS Genes
The Scientist's Toolkit: Essential Research Reagents
A successful VIGS experiment relies on a core set of validated reagents and vectors. The following table details essential components for establishing a TRV-based VIGS system.
Table 3: Key Research Reagent Solutions for TRV-VIGS
| Reagent / Solution | Function / Purpose | Example Specifications / Notes |
|---|---|---|
| pTRV1 & pTRV2 Vectors | Bipartite viral vector system. pTRV1 encodes replication/movement proteins; pTRV2 carries the target gene insert. | Available from academic stock centers; pTRV2 is modified with MCS for cloning [1]. |
| pTRV2-GFP Vector | Tracking vector. Allows visual confirmation of viral infection and spread via fluorescence microscopy. | Critical for optimizing inoculation methods in new species/tissues [20] [3]. |
| A. tumefaciens GV3101 | Disarmed Agrobacterium strain. Used for efficient delivery of T-DNA containing TRV vectors into plant cells. | Preferred for its high transformation efficiency and virulence [3] [9]. |
| Acetosyringone | Phenolic compound. Induces the Agrobacterium Vir genes, enhancing T-DNA transfer efficiency. | Used in induction and infiltration buffers at 150-200 µM [3] [9]. |
| Infiltration Buffer | Suspension medium. Maintains Agrobacterium viability and facilitates infiltration. Typically 10 mM MgCl₂, 10 mM MES, pH 5.6. | Optimized osmolarity and pH are critical for infection success. |
| Kanamycin & Rifampicin | Selection antibiotics. Ensure bacterial cultures contain the correct plasmids and are axenic. | Standard concentrations: 50 µg/mL for Kanamycin, 25-50 µg/mL for Rifampicin [9]. |
The protocols and validation strategies outlined herein provide a robust framework for employing VIGS to investigate genes of agronomic importance. By moving beyond the initial optimization phase marked by PDS photobleaching and implementing rigorous, multi-faceted validation techniques, researchers can reliably unravel the function of genes governing complex traits like disease resistance and development. The continuous optimization of delivery methods for recalcitrant species, as demonstrated in woody plants like Camellia drupifera, ensures that VIGS will remain an indispensable tool in the plant functional genomics toolkit, accelerating the pace of discovery and molecular breeding.
In the context of Agrobacterium-mediated Virus-Induced Gene Silencing (VIGS), robust molecular validation is paramount for confirming the silencing of target genes. This Application Note provides a detailed protocol for using quantitative reverse transcription PCR (qRT-PCR) to accurately measure target gene downregulation in VIGS experiments, with a specific focus on the analysis of tomato and related species using available database resources. The guidelines are framed within a broader thesis research on Agrobacterium-mediated VIGS infection methods, providing scientists with standardized procedures for quantifying gene silencing efficiency. We detail the mathematical frameworks for data analysis, experimental design considerations, and implementation protocols to ensure reliable quantification of target gene expression changes, enabling researchers in drug development and plant biotechnology to obtain publication-quality results.
Mathematical Foundations for qRT-PCR Data Analysis
Core Calculation Methods
Two primary mathematical approaches are commonly employed for calculating fold change in qRT-PCR experiments, each with distinct assumptions and applications.
Table 1: Core Mathematical Methods for qRT-PCR Fold Change Calculation
| Method | Formula | Key Assumptions | Application Context |
|---|---|---|---|
| Livak (2-ΔΔCT) [59] | ( FC = 2^{-[ \Delta CT{Treatment} - \Delta CT{Control} ]} )Where ( \Delta CT = CT{target} - CT{ref} ) | Both target and reference genes amplify with near-perfect efficiency (E ≈ 2 or 100%) [59]. | Ideal for well-optimized assays with validated, highly efficient primers. |
| Pfaffl (Efficiency-Adjusted) [59] | ( FC = \frac{(E{target})^{-\Delta CT{target}}}{(E{ref})^{-\Delta CT{ref}}} )Where ( \Delta CT = CT{Treatment} - CT{Control} ) | Incorporates actual amplification efficiencies (E) for both target and reference genes [59]. | Essential when amplification efficiencies deviate from 100% or differ between target and reference genes. |
The Livak method, also known as the 2-ΔΔCT method, is valued for its simplicity but requires the critical assumption that amplification efficiencies for both target and reference genes are approximately 100% [59]. The Pfaffl method offers greater flexibility and accuracy by incorporating experimentally determined amplification efficiencies for both target and reference genes, providing a more reliable fold change calculation when efficiency values deviate from ideal conditions [59].
Advanced Analysis and Statistical Considerations
For comprehensive data analysis, the rtpcr package in R provides a robust framework that accommodates the Pfaffl method and extends to complex experimental designs [59]. This package uses efficiency-weighted ΔCT (wΔCT) values, calculated as:
[ \text{wΔCT} = \log2(E{target}) \cdot CT{target} - \log2(E{ref}) \cdot CT{ref} ]
Fold change is then derived from these weighted values:
[ FC = 2^{-[ \overline{\text{wΔCT}}{Treatment} - \overline{\text{wΔCT}}{Control} ]} ]
This approach leverages the normal distribution of wΔCT values, enabling the application of t-tests for two-group comparisons or analysis of variance (ANOVA) for multi-group experimental designs, providing standard errors, confidence intervals, and statistical mean comparisons for robust hypothesis testing [59].
Experimental Design and Workflow
Reference Gene Selection and Validation
The accuracy of qRT-PCR normalization depends critically on the stability of reference genes. The selection process must be tailored to the specific experimental conditions.
Table 2: Strategies for Reference Gene Selection and Validation
| Strategy | Description | Tools/Resources | Considerations |
|---|---|---|---|
| Housekeeping Genes (HKGs) | Use of classical constitutive genes (e.g., ACT, TUB, GAPDH). | Literature-based selection. | Not universally stable. Expression can vary with tissue type and stress conditions [60] [61]. |
| Lowest Variance Genes (LVGs) | Mining RNA-Seq databases to find genes with minimal expression variance across conditions of interest. | TomExpress (tomato), Genevestigator, public RNA-Seq repositories [60]. | Stability is context-dependent. Requires a comprehensive database [60]. |
| Stable Gene Combinations | Using a geometric mean of multiple non-stable genes whose expressions balance each other. | Custom algorithms applied to RNA-Seq data [60]. | Can outperform single stable genes. Requires computational identification [60]. |
| Experimental Validation | Statistical ranking of candidate reference genes from qPCR data. | geNorm, NormFinder, BestKeeper, RefFinder [61]. | Essential final step. Identifies the most stable genes for a specific experimental setup [61]. |
Research demonstrates that a stable combination of non-stable genes can outperform single reference genes [60]. For tomato studies, the TomExpress database provides a comprehensive resource for in silico identification of stable genes and combinations [60]. For all species, validation with programs like geNorm, NormFinder, and BestKeeper is crucial, as reference gene stability varies significantly across tissues, developmental stages, and stress conditions [61].
qRT-PCR Experimental Protocol
Sample Preparation and RNA Extraction
- Plant Material: Harvest tissue from VIGS-treated and control plants. For temporal studies, ensure consistent harvesting time points post-agroinfiltration (e.g., 32 days for photobleaching observation in Castilleja tenuiflora [14]).
- RNA Extraction: Perform using a standardized kit or method. Include DNase I treatment to eliminate genomic DNA contamination.
- cDNA Synthesis: Synthesize cDNA from 1 μg of total RNA using reverse transcriptase and oligo(dT) or random hexamer primers.
qPCR Setup and Run
- Reaction Mix: Prepare SYBR Green or TaqMan master mix according to manufacturer's instructions. Include no-template controls (NTCs) for each primer set to detect contamination.
- Primer Validation: Use primers with optimized amplification efficiencies (90–110%) and validated specificity via melt curve analysis (for SYBR Green).
- Thermocycling Conditions: Standard cycling conditions include an initial denaturation (95°C for 10 min), followed by 40 cycles of denaturation (95°C for 15 sec) and annealing/extension (60°C for 1 min).
Data Analysis Workflow
The process of analyzing qRT-PCR data to obtain statistically sound fold-change values involves several critical steps, from raw data preprocessing to final statistical comparison. The following workflow outlines this complete pathway:
Critical Pre-Analysis Steps
Baseline Correction: The fluorescence baseline must be correctly defined using early cycles (e.g., cycles 5-15) that precede the exponential amplification phase. Incorrect baseline settings can significantly distort Cq values and subsequent fold-change calculations [62].
Threshold Setting: The threshold should be placed within the exponential phase of all amplification curves, where they are parallel. This ensures that ΔCq values between samples remain constant, regardless of the specific threshold position. When amplification curves are not parallel, ΔCq values become threshold-dependent, compromising data reliability [62].
Implementation with Statistical Software
The rtpcr package in R streamlines this workflow by integrating data preprocessing, efficiency-weighted calculation of ΔCT values, fold-change calculation, and statistical comparison into a cohesive framework [59]. The package supports t-tests for simple treatment-control comparisons and ANOVA or ANCOVA for experiments with multiple factors, providing standard errors and confidence intervals for the calculated expression values [59].
The Scientist's Toolkit
Table 3: Essential Research Reagents and Computational Tools
| Item/Category | Function/Role | Example Application |
|---|---|---|
| qPCR Reagents | Fluorescent detection of amplified DNA. | SYBR Green, TaqMan probes [59]. |
| Reverse Transcriptase | Synthesis of complementary DNA (cDNA) from RNA templates. | First-strand cDNA synthesis for qPCR template. |
| Reference Genes | Internal control for normalization of target gene expression. | AlEF1A, AlTUB6 (stress-specific) [61], ACT.2, ACT.3 (tomato) [60]. |
| VIGS Vectors | Delivery system for triggering gene silencing. | TRV-based pTRV1 and pTRV2 for Agrobacterium-mediated VIGS [14]. |
| Analysis Software | Cq determination, efficiency calculation, and statistical analysis. | rtpcr R package [59], GeneNorm, NormFinder, BestKeeper [61]. |
| RNA-Seq Database | In silico mining of stable reference genes. | TomExpress (tomato) for identifying LVGs and stable combinations [60]. |
Troubleshooting and Validation
Amplification Efficiency Determination
- Calculation: Perform a standard curve with a serial dilution (e.g., 1:10) of cDNA template. Amplification efficiency (E) is calculated from the slope of the standard curve: ( E = 10^{-1/slope} ). Ideal efficiency is 2 (100%), with the acceptable range being 1.8–2.2 [62].
- Impact: Significantly different efficiencies between target and reference genes mandate the use of the Pfaffl method for accurate fold-change calculation [59] [62].
Experimental Validation of Silencing
Beyond qRT-PCR, incorporating a phenotypic marker provides robust validation of VIGS efficacy. For example, in Castilleja tenuiflora, silencing the Cte-chlH or Cte-PDS genes via TRV-based VIGS resulted in visible photobleaching phenotypes in 80% and 31% of plants, respectively, providing a visual correlation with molecular downregulation [14]. This phenotypic evidence, coupled with statistical significance (p ≤ 0.01) between normalized expression in silenced versus control plants, offers compelling confirmation of successful gene silencing [14].
Within the broader scope of Agrobacterium-mediated VIGS (Virus-Induced Gene Silencing) research, tracking the systemic spread of the viral vector is crucial for assessing the efficiency and progression of gene silencing. The use of the Green Fluorescent Protein (GFP) as a reporter enables the non-destructive, real-time visualization of this process. When expressed from a viral vector, GFP serves as a quantitative proxy for viral replication and movement, allowing researchers to monitor the establishment of infection and the progression of silencing throughout the plant [63] [64]. This protocol details the application of GFP-tagged viral vectors for monitoring systemic spread in the context of VIGS, with a focus on Agrobacterium-mediated delivery.
Quantitative Data on Viral Vector and Reporter Systems
The choice of viral vector and reporter system significantly impacts the efficiency and reliability of tracking systemic movement. The table below summarizes key characteristics of different systems based on current research.
Table 1: Comparison of Viral Vectors and Reporter Systems for Tracking Systemic Spread
| Viral Vector | Reporter Protein | Stability of Tag | Primary Application | Key Advantage | Reference |
|---|---|---|---|---|---|
| Cucumber Mosaic Virus (CMV) | iLOV | Stable for >28 days and through serial passages [64]. | Long-term infection dynamics studies; co-infection assays. | Small size ensures genetic stability in the viral genome. | [64] |
| Tobacco Rattle Virus (TRV) | GFP | Stable for at least 21 days post-inoculation [3]. | VIGS in solanaceous plants and soybean; functional genomics. | Induces mild symptoms, minimizing impact on silencing phenotype. | [1] [3] |
| Broad Bean Wilt Virus 2 (BBWV2) | stagRFP | Stable expression in co-infection studies [64]. | Visualizing viral synergies; multi-virus interaction studies. | Provides an orthogonal fluorescent channel for co-infection. | [64] |
GFP is not only a visual marker but also a quantitative reporter. Research demonstrates that GFP fluorescence intensity, when measured via flow cytometry, increases in direct proportion to the GFP gene copy number and mRNA abundance, providing a reliable measure of underlying gene expression driven by the viral vector [63].
Experimental Protocols
Protocol 1: Visualizing Systemic Spread of a GFP-Tagged TRV Vector in Soybean
This optimized protocol uses Agrobacterium-mediated delivery via the cotyledon node for efficient systemic infection of soybean, a challenging host [3].
Key Materials:
- Agrobacterium tumefaciens strain GV3101 harboring pTRV1 and pTRV2-GFP vectors.
- Seeds of soybean (Glycine max) cultivar "Tianlong 1".
- Sterilization solution (e.g., 6% sodium hypochlorite).
- Liquid LB media with appropriate antibiotics (Kanamycin, Rifampicin).
- Induction medium (10 mM MES, 10 mM MgCl₂, 200 µM Acetosyringone, pH 5.6).
Methodology:
- Vector Construction: Clone your gene of interest (for VIGS) or a neutral sequence into the pTRV2-GFP multiple cloning site using standard molecular biology techniques [3].
- Agrobacterium Preparation:
- Transform the pTRV1 and recombinant pTRV2-GFP plasmids into A. tumefaciens GV3101.
- Inoculate single colonies into liquid LB with antibiotics and grow overnight at 28°C with shaking.
- Centrifuge the culture and resuspend the pellet in induction medium to an optical density at 600 nm (OD₆₀₀) of 1.0.
- Incubate the resuspended culture for 3-4 hours at room temperature with shaking.
- Plant Material Preparation:
- Surface-sterilize soybean seeds and soak in sterile water until swollen.
- Critical Step: longitudinally bisect the swollen seeds to create half-seed explants, exposing the cotyledonary node.
- Agroinfiltration:
- Mix the pTRV1 and pTRV2-GFP Agrobacterium suspensions in a 1:1 ratio.
- Immerse the half-seed explants in the mixed Agrobacterium suspension for 20-30 minutes with gentle agitation.
- Blot-dry the explants and transfer to sterile filter paper in Petri dishes for co-cultivation.
- Plant Growth and Monitoring:
- After 3-4 days of co-cultivation in the dark, transfer the explants to soil or appropriate growth medium.
- Visualization: Monitor GFP fluorescence systemically every few days. Using a fluorescence microscope, systemic spread can be confirmed as early as 4 days post-infection in the hypocotyl and cotyledonary node, with fluorescence eventually reaching newly emerged trifoliate leaves [3].
Protocol 2: Tracking Co-infection Dynamics with CMV-iLOV and BBWV2-stagRFP
This protocol allows for the simultaneous visualization of two viruses, revealing complex interactions like synergistic spread [64].
Key Materials:
- Agrobacterium strains harboring CMV RNA1, CMV RNA2-iLOV, and BBWV2-R2-stagRFP infectious cDNA clones.
- Nicotiana benthamiana plants (2-week old).
- Agroinfiltration buffers.
Methodology:
- Agrobacterium Preparation: Prepare cultures for each construct as described in Protocol 1, resuspending to an OD₆₀₀ of 0.5-1.0.
- Plant Infiltration:
- For co-infection, mix the Agrobacterium suspensions containing CMV RNA1, CMV RNA2-iLOV, and BBWV2-R2-stagRFP.
- Using a needleless syringe, infiltrate the mixed culture into the abaxial side of two lower leaves of N. benthamiana.
- Time-Course Imaging:
- Begin imaging at 2-3 days post-inoculation (dpi) and continue at regular intervals.
- Use a fluorescence stereomicroscope with appropriate filter sets for iLOV (green) and RFP (red).
- Document the spatial progression of each virus from the inoculated leaves to the upper, non-inoculated leaves.
- Data Interpretation: This approach can reveal enhanced spread, as demonstrated by BBWV2 facilitating the cell-to-cell movement of CMV in young leaves during co-infection [64].
Visualization and Workflow
The following diagram illustrates the core experimental workflow for monitoring viral systemic spread using a GFP reporter, from vector preparation to final analysis.
Experimental Workflow for Viral Movement Tracking with GFP
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for Viral Movement Tracking with GFP Reporters
| Reagent / Material | Function / Role in Experiment | Example & Notes |
|---|---|---|
| Viral Vectors | Engineered backbones for delivering and expressing the GFP reporter systemically. | pTRV1/pTRV2-GFP [3]: A bipartite system where TRV2 carries the GFP gene. CMV RNA2-iLOV [64]: A genetically stable CMV vector with the iLOV reporter. |
| Fluorescent Reporters | Visual markers for real-time, non-destructive tracking of viral location and spread. | GFP (Green Fluorescent Protein): Standard reporter, quantifiable via flow cytometry [63]. iLOV: A smaller, more stable alternative to GFP for viruses with limited genetic capacity [64]. |
| Agrobacterium Strain | Delivery vehicle for introducing the viral vector into plant cells. | GV3101: A disarmed strain commonly used for agroinfiltration in VIGS protocols [3]. |
| Induction Medium Supplements | Compounds that activate the Agrobacterium Vir genes for efficient T-DNA transfer. | Acetosyringone (200 µM): Essential for inducing virulence in non-wounded plant tissues [3]. MES buffer: Maintains optimal pH for the agroinfiltration process. |
| Visualization Tools | Equipment for detecting and quantifying GFP fluorescence. | Fluorescence Stereomicroscope: For whole-organ or whole-plant imaging of systemic spread. Confocal Microscope: For high-resolution cellular localization. Flow Cytometer: For quantitative analysis of GFP expression levels in single cells [63]. |
Virus-Induced Gene Silencing (VIGS) has emerged as a powerful reverse genetics tool for rapid functional genomics in plants, particularly for species challenging to transform stably. This Agrobacterium-mediated approach leverages recombinant viral vectors to trigger post-transcriptional gene silencing (PTGS) of endogenous plant genes, enabling researchers to study gene function through observable phenotypic changes [1]. The core principle involves engineering viral vectors, most commonly Tobacco Rattle Virus (TRV), to carry fragments of target plant genes, which upon infection, activate the plant's RNA interference machinery, leading to sequence-specific degradation of homologous mRNA transcripts [16] [1].
The efficiency of VIGS is profoundly influenced by the method of inoculation, which determines the initial viral load, tissue specificity, and systemic spread of the silencing signal. Inoculation techniques vary significantly in their technical requirements, efficiency across species, and applicability to different plant developmental stages. While Agrobacterium tumefaciens serves as the primary delivery vehicle for TRV vectors across most protocols, the physical method of introducing this bacterial suspension into plant tissues remains a critical variable requiring optimization for each species, tissue type, and experimental setup [8] [7]. Understanding the comparative advantages and limitations of these techniques is essential for designing effective VIGS experiments, particularly for non-model species and recalcitrant tissues where standard protocols often fail.
Comparative Analysis of Inoculation Techniques
Efficiency Metrics and Performance Indicators
Evaluating VIGS inoculation efficiency involves multiple quantitative and qualitative metrics. Key performance indicators include infection percentage (proportion of treated plants showing viral infection), silencing efficiency (degree of target gene knockdown measured by qRT-PCR), phenotype penetrance (proportion of infected plants showing expected silencing phenotypes), and systemic spread (extent of silencing in tissues distal to inoculation site) [9] [8]. Additional considerations include technical complexity, time requirements, equipment needs, and potential for tissue damage that might confound phenotypic analysis. Different techniques optimize different aspects of these metrics, making technique selection highly dependent on experimental priorities and biological constraints.
Table 1: Comparative Efficiency of VIGS Inoculation Techniques Across Plant Species
| Inoculation Technique | Target Species | Infection Efficiency | Silencing Efficiency | Key Advantages | Major Limitations |
|---|---|---|---|---|---|
| Pericarp Cutting Immersion | Camellia drupifera (woody capsules) | ~94% | ~70-91% (varies with gene) | Effective for recalcitrant woody tissues | Tissue-specific application |
| Cotyledon Node Immersion | Soybean (Glycine max) | Up to 95% | 65-95% | Bypasses leaf trichomes/cuticle; systemic spread | Requires sterile tissue culture |
| Seed Vacuum Infiltration | Sunflower (Helianthus annuus) | 62-91% (genotype-dependent) | High (normalized expression = 0.01) | Simple, no in vitro steps; high-throughput | Genotype-dependent efficiency |
| Shoot Apical Meristem Inoculation | Petunia (Petunia × hybrida) | Significantly improved | 28-69% increase in silencing area | Strongest and most consistent silencing | Technically challenging |
| Vacuum Infiltration (whole plant) | Styrax japonicus | Not specified | 83% | Efficient for whole-plant silencing | Requires specialized equipment |
| Friction-Osmosis | Styrax japonicus | Not specified | 74% | Simpler equipment requirements | Lower efficiency than vacuum |
Technical Specifications and Methodological Considerations
Each inoculation method has specific technical parameters that critically influence efficiency. For vacuum-based methods, key parameters include vacuum pressure intensity, duration of application, and bacterial concentration (OD600). In immersion techniques, critical factors include immersion duration, tissue pre-treatment, and bacterial suspension composition. Mechanical methods like apical meristem inoculation require precision in wounding depth and inoculum application to balance efficiency with tissue damage [16] [7].
Optimal bacterial concentrations (OD600) typically range from 0.5 to 1.0, with specific optima depending on the technique and species. For instance, in Styrax japonicus, vacuum infiltration achieved optimal efficiency at OD600 0.5, while friction-osmosis performed better at OD600 1.0 [7]. The addition of acetosyringone (200 μM), a phenolic inducer of Agrobacterium virulence genes, consistently enhances transformation efficiency across methods [7]. Co-cultivation duration post-inoculation represents another critical parameter, with sunflower protocols optimizing at 6 hours for seed vacuum infiltration [8].
Table 2: Optimized Technical Parameters for VIGS Inoculation Techniques
| Parameter | Pericarp Cutting Immersion | Cotyledon Node Immersion | Seed Vacuum Infiltration | Apical Meristem Inoculation |
|---|---|---|---|---|
| Optimal Developmental Stage | Early to mid capsule development (279 DAP) | 3-4 weeks after sowing | Germinated seeds | 3-4 weeks after sowing |
| Agrobacterium OD600 | 0.9-1.0 | Not specified | 0.9-1.0 | Not specified |
| Acetosyringone Concentration | 200 μM | Not specified | 200 μM | Not specified |
| Infiltration Duration | Immersion time not specified | 20-30 minutes | Vacuum duration not specified; 6h co-cultivation | Not specified |
| Temperature Regime | Not specified | Not specified | 22°C average | 20°C day/18°C night |
Detailed Experimental Protocols
Cotyledon Node Immersion for Soybean
The cotyledon node immersion method has been optimized for soybean to overcome limitations posed by the thick cuticle and dense trichomes on leaves that impede liquid penetration [3]. The protocol begins with surface sterilization of soybean seeds using 70% ethanol for 30 seconds followed by 2% sodium hypochlorite for 6 minutes, with thorough rinsing using sterile water between and after treatments. Sterilized seeds are soaked in sterile water until swollen, typically for 12-24 hours, then longitudinally bisected to obtain half-seed explants, carefully preserving the cotyledonary node region.
Fresh explants are immersed for 20-30 minutes in an Agrobacterium tumefaciens strain GV3101 suspension containing a 1:1 mixture of pTRV1 and pTRV2-derived vectors (e.g., pTRV2-GFP with target gene insert) at optimal density [3]. The infiltration suspension is prepared by resuspending Agrobacterium pellets from overnight cultures in infiltration medium (10 mM MgCl2, 10 mM MES, 200 μM acetosyringone) to an appropriate OD600. Following immersion, explants are blotted dry on sterile filter paper and transferred to co-cultivation medium for 2-3 days in darkness at 22°C. Plants are subsequently transferred to standard growth conditions, with silencing phenotypes typically observable within 2-3 weeks post-inoculation [3].
Seed Vacuum Infiltration for Sunflower
The seed vacuum infiltration method provides a high-throughput approach for sunflower that eliminates requirements for in vitro culture and complex sterile procedures [8]. The protocol begins with careful removal of seed coats from dry sunflower seeds to facilitate infiltration. Prepared seeds are submerged in Agrobacterium tumefaciens strain GV3101 suspension carrying TRV vectors in a vacuum desiccator. The Agrobacterium suspension is prepared by growing bacterial cultures to OD600 0.9-1.0 in YEB medium with appropriate antibiotics, harvesting by centrifugation, and resuspending in infiltration medium (10 mM MgCl2, 10 mM MES, 200 μM acetosyringone).
Application of vacuum (approximately 0.5-0.8 bar) for a specified duration forces the bacterial suspension into intercellular spaces within the seeds. Optimal parameters determined for sunflower include 200 μM acetosyringone concentration and 6 hours of co-cultivation post-infiltration [8]. Following vacuum treatment and co-cultivation, seeds are sown directly in soil mixture (3:1 peat:perlite) and grown under standard greenhouse conditions (22°C, 18h light/6h dark photoperiod, 45% relative humidity). This method achieves infection percentages of 62-91% across different sunflower genotypes, demonstrating both its efficiency and genotype-dependent variability [8].
Apical Meristem Inoculation for Petunia
Shoot apical meristem inoculation represents a highly efficient approach for species like petunia where traditional leaf infiltration methods yield variable results [16]. The protocol involves mechanical wounding of the shoot apical meristem of 3-4 week old plants using a sterile needle or scalblad. Approximately 10-20 μL of Agrobacterium tumefaciens suspension carrying TRV vectors is immediately applied directly to the wounded meristematic tissue.
Critical optimization parameters identified for petunia include plant age (3-4 weeks after sowing superior to 5 weeks) and growth temperature (20°C day/18°C night inducing stronger silencing than higher temperatures) [16]. The bacterial suspension should be prepared from freshly transformed Agrobacterium grown to mid-log phase (OD600 0.6-0.8) in LB medium with appropriate antibiotics, then induced with acetosyringone (200 μM) for several hours before inoculation. This method significantly increases silencing area (69% for CHS, 28% for PDS) compared to other techniques and induces more consistent systemic silencing throughout the plant [16].
Essential Research Reagent Solutions
Core Reagents for VIGS Implementation
Successful implementation of VIGS requires carefully selected reagents and vectors optimized for efficient gene silencing. The core toolkit includes viral vectors, Agrobacterium strains, selection antibiotics, and induction compounds that collectively enable effective delivery and silencing of target genes. These reagents have been refined through extensive optimization across multiple plant species and represent the current standards for reliable VIGS experiments.
Table 3: Essential Research Reagents for VIGS Experiments
| Reagent/Vector | Function/Purpose | Key Features & Optimization Notes |
|---|---|---|
| TRV Vectors (pTRV1/pTRV2) | Bipartite viral vector system | pTRV1: Encodes replication/movement proteins; pTRV2: Contains cloning site for target gene fragments [16] [3] |
| Agrobacterium tumefaciens GV3101 | Delivery vehicle for TRV vectors | Preferred strain for plant transformation; requires rifampicin, gentamicin, and kanamycin selection [3] [8] |
| Acetosyringone | Inducer of Agrobacterium virulence genes | Critical for efficient T-DNA transfer; optimal concentration typically 150-200 μM in infiltration medium [7] |
| pTRV2-sGFP Control Vector | Negative control to eliminate viral symptoms | Contains fragment of green fluorescent protein gene instead of plant gene; minimizes plant necrosis and stunting [16] |
| Infiltration Medium (10 mM MgCl₂, 10 mM MES) | Suspension medium for Agrobacterium | Maintains bacterial viability during inoculation while not inducing plant defense responses [3] [8] |
| Phytoene Desaturase (PDS) Gene Fragment | Visual marker for silencing efficiency | Silencing causes photobleaching; serves as positive control for system functionality across species [16] [3] [8] |
The comparative analysis of VIGS inoculation techniques reveals a complex landscape where optimal method selection depends on multiple factors including target species, tissue type, available resources, and experimental objectives. Seed vacuum infiltration offers exceptional efficiency for high-throughput applications in amenable species like sunflower, while cotyledon node immersion effectively bypasses morphological barriers in challenging species like soybean. For maximum silencing intensity in model systems like petunia, apical meristem inoculation remains superior despite its technical demands.
Critical success factors consistently emerge across techniques: precise developmental staging, optimized Agrobacterium densities (OD600 0.5-1.0), inclusion of acetosyringone (200 μM) in infiltration media, and controlled post-inoculation environments. The significant genotype-dependent efficiency observed in sunflower [8] underscores the necessity for method validation across genetic backgrounds within target species. Future protocol development should address current limitations in woody species and recalcitrant tissues while expanding the vector toolkit beyond TRV to accommodate diverse host-pathogen compatibility requirements.
For researchers implementing these techniques, we recommend initial validation using visual marker genes (PDS, CHS) to establish system efficiency followed by careful optimization of the identified critical parameters for specific experimental systems. The protocols detailed herein provide robust starting points for such optimization across diverse plant species and tissue types.
Cotton leaf curl disease (CLCuD), caused by Begomoviruses, is a devastating disease that severely limits cotton production, particularly in susceptible Gossypium hirsutum varieties [65] [66]. The disease is transmitted by the whitefly (Bemisia tabaci) and leads to characteristic leaf curling symptoms that can drastically reduce yield [65] [66]. In contrast, another cotton species, Gossypium arboreum (desi cotton), exhibits natural resistance to CLCuD, while some G. hirsutum accessions like Mac7 show tolerance compared to highly susceptible varieties like Coker 312 [66].
A key class of plant resistance (R) genes, the nucleotide-binding site leucine-rich repeat (NBS-LRR) genes, plays a crucial role in effector-triggered immunity (ETI) against various pathogens, including viruses [66] [67]. These genes are characterized by a conserved NBS domain and are divided into subclasses such as TIR-NBS-LRR (TNL) and CC-NBS-LRR (CNL) based on their N-terminal domains [66] [67]. Recent genomic studies have identified thousands of NBS-domain-containing genes across plant species, with significant diversity in their domain architecture [66]. This case study details the application of Agrobacterium-mediated virus-induced gene silencing (VIGS) to functionally validate the role of a specific NBS-encoding gene in conferring resistance to CLCuD.
Experimental Design and Key Findings
Identification and Selection of Candidate NBS Genes
The initial phase of the study involved a comparative genome-wide analysis of NBS-encoding genes in resistant and susceptible cotton genotypes. Research identified 12,820 NBS-domain-containing genes across 34 plant species, which were classified into 168 distinct classes based on their domain architecture [66]. In upland cotton (G. hirsutum), 437 NBS-LRR genes were identified, distributed across 26 chromosomes, with 315 belonging to the CNL subclass and 122 to the TNL subclass [67].
- Transcriptomic Profiling: RNA-seq expression analysis of NBS genes in CLCuD-tolerant (G. arboreum, Mac7) and susceptible (Coker 312) cotton genotypes under biotic stress conditions revealed differential expression patterns. Orthogroups OG2, OG6, and OG15 showed putative upregulation in resistant plants, suggesting their potential role in the defense response [66].
- Genetic Variation Analysis: Comparison between susceptible (Coker 312) and tolerant (Mac7) G. hirsutum accessions identified substantial genetic variation within NBS genes, with 6,583 unique variants in Mac7 and 5,173 in Coker 312, further highlighting candidate genes potentially involved in resistance mechanisms [66].
- Candidate Gene Selection: Based on these analyses, a specific NBS gene from Orthogroup 2 (OG2), hereafter referred to as GaNBS, was selected for functional validation due to its strong association with resistance profiles [66].
Functional Validation via VIGS and Key Results
To confirm the functional role of GaNBS in CLCuD resistance, a VIGS-based loss-of-function approach was employed in a resistant cotton background.
- Silencing Efficiency and Phenotype: The VIGS system successfully silenced GaNBS, and the silencing was confirmed via quantitative PCR. The resistant plants with silenced GaNBS showed a significant increase in viral titer compared to control plants, demonstrating that GaNBS is necessary for restricting virus accumulation [66].
- Interaction Studies: Protein-ligand and protein-protein interaction simulations predicted strong binding of the GaNBS protein with ADP/ATP and various core proteins of the cotton leaf curl disease virus, providing a mechanistic insight into its role in pathogen recognition and defense signaling [66].
Table 1: Summary of Key Experimental Findings from the Case Study
| Experimental Stage | Key Finding | Implication |
|---|---|---|
| Genome Identification | 437 NBS-LRR genes identified in G. hirsutum; 12,820 NBS genes across land plants [66] [67] | Reveals extensive diversity and a large pool of candidate resistance genes. |
| Expression Analysis | Upregulation of orthogroups OG2, OG6, and OG15 in resistant genotypes under stress [66] | Prioritizes specific NBS gene clusters for functional studies. |
| Genetic Variation | 6,583 unique NBS gene variants in tolerant Mac7 vs. 5,173 in susceptible Coker 312 [66] | Suggests a genetic basis for resistance in specific accessions. |
| VIGS Validation | Silencing of GaNBS (OG2) led to increased virus titer in a resistant plant [66] | Confirms the putative role of GaNBS in virus resistance. |
Detailed Protocols
Protocol 1: VIGS Vector Construction and Agrobacterium Preparation
This protocol outlines the steps for constructing the Tobacco Rattle Virus (TRV)-based VIGS vector and preparing the Agrobacterium tumefaciens culture for plant transformation [3] [4].
Materials:
- Plasmids: pTRV1 and pTRV2 (or pTRV2-GFP) vectors [3] [4].
- Bacterial Strains: E. coli DH5α competent cells, Agrobacterium tumefaciens strain GV3101 [3] [4].
- Enzymes and Kits: Restriction enzymes (e.g., EcoRI, XhoI), DNA ligase, plasmid extraction kit.
- Primers: Gene-specific primers with added restriction sites (e.g.,
GAATTCfor EcoRI andCTCGAGfor XhoI) [4].
Procedure:
- Amplify Target Gene Fragment: Using cDNA from healthy cotton leaves as a template, amplify a 300-500 base pair fragment of the target GaNBS gene with designed primers [4].
- Digest and Ligate: Digest both the purified PCR product and the pTRV2 vector with the appropriate restriction enzymes. Purify the digested fragments and ligate them using T4 DNA ligase [3] [4].
- Transform and Sequence: Transform the ligation product into E. coli DH5α cells. Select positive clones on LB plates with antibiotics (e.g., kanamycin) and verify the insert by Sanger sequencing [4].
- Mobilize into Agrobacterium: Isolate the correct recombinant plasmid and introduce it into A. tumefaciens GV3101 via electroporation or freeze-thaw method [3] [4].
- Prepare Agrobacterium Culture: Inoculate a single positive Agrobacterium colony into LB medium with appropriate antibiotics (e.g., kanamycin, rifampicin) and incubate at 28°C with shaking for ~16 hours. Centrifuge the culture and resuspend the pellet in an induction medium (e.g., LB with 10 mM MES, 20 μM acetosyringone) to an final optical density at 600 nm (OD₆₀₀) of 1.0-2.0. Incubate the resuspended culture for 4-6 hours at 28°C before use [3].
Protocol 2: Agrobacterium-Mediated VIGS in Cotton
This protocol describes an optimized infiltration method for efficient VIGS in cotton, adapted from successful procedures in soybean and other crops [3] [4].
Materials:
- Plant Material: Seeds of the CLCuD-resistant cotton genotype (e.g., G. arboreum 'Ravi').
- Agrobacterium Cultures: A. tumefaciens GV3101 containing pTRV1 and pTRV2-GaNBS (or empty pTRV2 as a control), prepared as in Protocol 1.
- Infiltration Buffer: 10 mM MgCl₂, 10 mM MES, 150 μM acetosyringone, pH 5.6 [3].
Procedure:
- Germinate Cotton Seeds: Surface-sterilize cotton seeds and germinate them in a sterile tissue culture environment until the cotyledons are fully expanded [4].
- Mix Agrobacterium Cultures: Combine the pTRV1 and pTRV2-GaNBS Agrobacterium cultures in a 1:1 ratio [3].
- Agroinfiltration via Cotyledon Node Injection:
- Acclimatization and Growth: Place the infiltrated plants in a growth chamber with high humidity for 2-3 days. Subsequently, transfer them to standard greenhouse conditions (e.g., 25°C, 16-hour light/8-hour dark photoperiod) [3].
- Monitor Silencing and Challenge with Virus:
- Phenotypic Monitoring: Observe plants for development of silencing symptoms. For conclusive proof, include a positive control where a marker gene like Phytoene Desaturase (PDS) is silenced, resulting in photobleaching [3] [4].
- Molecular Confirmation: At 2-3 weeks post-infiltration, harvest leaf tissue and use reverse transcription quantitative PCR (RT-qPCR) to confirm the downregulation of the target GaNBS gene [66] [3].
- Disease Assay: Inoculate the silenced and control plants with CLCuV via viruliferous whiteflies or agro-infection. Monitor disease symptoms and quantify viral DNA accumulation using PCR or qPCR 3-4 weeks post-inoculation [66].
Table 2: The Scientist's Toolkit: Key Research Reagent Solutions
| Reagent/Resource | Function/Application | Example/Specification |
|---|---|---|
| TRV VIGS Vectors | Bipartite viral vector system for inducing gene silencing; pTRV1 encodes replication/movement proteins, pTRV2 carries the target gene fragment [3] [1]. | pTRV1, pTRV2, pTRV2-GFP [3] [4] |
| Agrobacterium tumefaciens | Delivery vehicle for introducing TRV vectors into plant cells. | Strain GV3101 [3] [4] |
| Acetosyringone | Phenolic compound that induces Agrobacterium virulence genes, crucial for efficient T-DNA transfer. | 150-200 μM in infiltration buffer [3] |
| Marker Genes (e.g., PDS) | Positive control for VIGS; silencing causes a visible photobleaching phenotype, confirming system functionality [3] [4]. | Phytoene Desaturase (GmPDS, CtePDS) [14] [3] |
| NBS Domain HMM Profile | Bioinformatics tool for identifying NBS-encoding genes in genomic sequences. | Pfam accession PF00931 (NBS domain) [66] [68] |
Visualized Workflows and Pathways
The following diagram illustrates the logical workflow and key biological pathways involved in this case study, from gene identification to functional validation.
Conclusion
Agrobacterium-mediated VIGS has firmly established itself as an indispensable, high-speed tool for functional genomics, effectively bypassing the bottlenecks of stable transformation. The continuous development of novel infection methods, such as root wounding-immersion and seed vacuum infiltration, alongside refined optimization strategies for vector design and environmental control, is dramatically expanding its applicability across diverse plant species. The future of VIGS lies in its deeper integration with multi-omics technologies and its growing potential as a vehicle for virus-mediated genome editing. For researchers in plant science and beyond, mastering these advanced VIGS methodologies provides a powerful platform to rapidly decode gene function, accelerating the discovery of genetic traits for enhanced crop resilience, improved nutritional quality, and sustainable agricultural innovation.
References
- 1. Virus-Induced Gene Silencing (VIGS) in Functional Genomics [https://www.mdpi.com/2311-7524/11/11/1297]
- 2. Frontiers | Virus-induced gene silencing is a versatile tool for unraveling... [https://www.frontiersin.org/journals/plant-science/articles/10.3389/fpls.2014.00323/full]
- 3. Efficient Virus-Induced Gene Silencing (VIGS) Method for ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12115190/]
- 4. Efficient Virus-Induced Gene Silencing (VIGS) Method for ... [https://www.mdpi.com/2223-7747/14/10/1547]
- 5. Establishment and application of a root wounding ... [https://www.frontiersin.org/journals/plant-science/articles/10.3389/fpls.2024.1336726/full]
- 6. A rapid, simple, and highly efficient method for VIGS and in ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC8605592/]
- 7. an efficient virus induced gene silencing (VIGS) system [https://www.sciencedirect.com/science/article/pii/S2468014123000833]
- 8. Advancing virus-induced gene silencing in sunflower: key factors of... [https://plantmethods.biomedcentral.com/articles/10.1186/s13007-024-01241-z]
- 9. Development of a robust and efficient virus-induced gene ... [https://plantmethods.biomedcentral.com/articles/10.1186/s13007-024-01320-1]
- 10. Visible gland constantly traces virus-induced gene ... [https://www.frontiersin.org/journals/plant-science/articles/10.3389/fpls.2022.1020841/full]
- 11. Development of Bean pod mottle virus-based vectors for ... [https://www.sciencedirect.com/science/article/pii/S0042682205005611]
- 12. The "one-step" Bean pod mottle virus (BPMV)-derived vector ... [https://bmcplantbiol.biomedcentral.com/articles/10.1186/s12870-014-0232-4]
- 13. Temporal and spatial Bean pod mottle virus‐induced gene ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC6638800/]
- 14. Agrobacterium-mediated transformation for virus-induced ... [https://pubmed.ncbi.nlm.nih.gov/40730887/]
- 15. Viral Vector Production: Applications, Challenges, Advances [https://www.cytena.com/resource-hub/blog/viral-vector-production-for-biomedicine-key-applications-and-industry-trends/]
- 16. An Optimized Protocol to Increase Virus-Induced Gene ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC3893464/]
- 17. A Cotyledon-based Virus-Induced Gene Silencing (Cotyledon ... [https://plantmethods.biomedcentral.com/articles/10.1186/s13007-024-01154-x]
- 18. An Efficient Agrobacterium-Mediated Transient ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12388925/]
- 19. Establishment of a Virus-Induced Gene-Silencing (VIGS) ... [https://www.mdpi.com/1422-0067/24/1/392]
- 20. Developing a homology-driven gene silencing system in < ... [https://www.maxapress.com/article/id/68c8b5d4fa6c58419c48f3c2]
- 21. The use of a candidate gene approach to study Botrytis ... [https://www.frontiersin.org/journals/plant-science/articles/10.3389/fpls.2023.1100416/full]
- 22. Agroinfiltration as an Effective and Scalable Strategy of ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC4113218/]
- 23. Leaf infiltration in plant science: old method, new possibilities [https://plantmethods.biomedcentral.com/articles/10.1186/s13007-021-00782-x]
- 24. Efficient Agroinfiltration of Plants for High-level Transient ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC3846102/]
- 25. Improving agroinfiltration-based transient gene expression in ... [https://plantmethods.biomedcentral.com/articles/10.1186/s13007-018-0343-2]
- 26. Agroinfiltration technique for elucidating the functions of ... [https://www.nature.com/articles/s41598-025-08344-0]
- 27. A simple and efficient in planta transformation method ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11009361/]
- 28. Agroinfiltration-mediated transient assay for rapid ... [https://www.sciencedirect.com/science/article/pii/S266590692500008X]
- 29. Optimised Agrobacterium-Mediated Transformation and ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10378916/]
- 30. Agroinfiltration [https://en.wikipedia.org/wiki/Agroinfiltration]
- 31. Virus-Induced Gene Silencing (VIGS): A Powerful Tool for ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10057534/]
- 32. Establishment and application of a root wounding ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11066161/]
- 33. Establishing a Virus-Induced Gene Silencing System ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10346259/]
- 34. Insights into the Root Invasion by the Plant Pathogenic ... [https://www.mdpi.com/2223-7747/9/4/516]
- 35. Advancing virus-induced gene silencing in sunflower [https://pubmed.ncbi.nlm.nih.gov/39135113/]
- 36. Vacuum and Co-cultivation Agroinfiltration of (Germinated) ... [https://www.frontiersin.org/journals/plant-science/articles/10.3389/fpls.2017.00393/full]
- 37. Efficient Virus-Induced Gene Silencing in Arabidopsis - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC1557620/]
- 38. Establishment of a Virus-Induced Gene Silencing System ... [https://www.mdpi.com/2223-7747/14/2/150]
- 39. based virus-induced gene Silencing in Atriplex canescens [https://plantmethods.biomedcentral.com/articles/10.1186/s13007-025-01427-z]
- 40. Virus-Induced Gene Silencing (VIGS) in Hydrangea ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11644798/]
- 41. Virus-Induced Gene Silencing, a Post Transcriptional ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC2699436/]
- 42. Effects of RNA m6A on the formation of multi-petalization in ... [https://www.sciencedirect.com/science/article/pii/S2468014125000640]
- 43. Establishment of virus-induced gene silencing (VIGS) system ... [https://plantmethods.biomedcentral.com/articles/10.1186/s13007-023-01064-4]
- 44. Improving transformation and regeneration efficiency in ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11659184/]
- 45. Developmental regulatory genes in recalcitrant forest trees [https://www.frontiersin.org/journals/plant-science/articles/10.3389/fpls.2025.1689705/full]
- 46. Recalcitrance to transformation, a hindrance for genome ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10551638/]
- 47. Developing and applying a virus-induced gene silencing ... [https://www.sciencedirect.com/science/article/abs/pii/S0378111924009685]
- 48. Optimized cDNA libraries for virus-induced gene silencing ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC2254408/]
- 49. An optimized disease resistance gene cloning workflow for ... [https://www.nature.com/articles/s41467-025-60033-8]
- 50. Comparative genomics‐driven design of virus‐delivered ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12576428/]
- 51. Viral suppressors from members of the family Closteroviridae ... [https://phytopatholres.biomedcentral.com/articles/10.1186/s42483-021-00104-y]
- 52. Viral suppressors of RNA-based viral immunity: Host targets [https://pmc.ncbi.nlm.nih.gov/articles/PMC2929401/]
- 53. viral silencing suppressors: Tools forged to fine-tune host- ... [https://www.sciencedirect.com/science/article/pii/S0042682215000744]
- 54. Engineering PVX vectors harboring heterologous VSRs to ... [https://www.nature.com/articles/s41598-025-24470-1]
- 55. Engineering PVX vectors harboring heterologous VSRs to ... [https://pubmed.ncbi.nlm.nih.gov/41254168/]
- 56. Establishment of virus-induced gene silencing (VIGS) ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10470258/]
- 57. Genome-Wide Identification of DnaJ Gene Family and ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12609765/]
- 58. Comparative analysis, diversification, and functional ... [https://www.nature.com/articles/s41598-024-62876-5]
- 59. rtpcr: a package for statistical analysis and graphical ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12530193/]
- 60. A stable combination of non-stable genes outperforms ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11682138/]
- 61. Validation of Reference Genes for Accurate RT-qPCR ... [https://www.mdpi.com/2073-4395/15/7/1596]
- 62. PCR/qPCR Data Analysis [https://www.sigmaaldrich.com/US/en/technical-documents/technical-article/genomics/qpcr/data-analysis?srsltid=AfmBOoqBv-FQsOGCKBY08Qp1NxQhaN4tkmYkTet6MSgazmrYt5bY0---]
- 63. Green fluorescent protein is a quantitative reporter of gene ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC1242169/]
- 64. Visual tracking of viral infection dynamics reveals the ... [https://www.nature.com/articles/s41598-023-34553-6]
- 65. Exploration of cotton leaf curl virus resistance genes and ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC6247833/]
- 66. Comparative analysis, diversification, and functional ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11126693/]
- 67. Genome-wide analysis of nucleotide binding site-leucine ... [https://www.sciencedirect.com/science/article/abs/pii/S0885576518301607]
- 68. Identification, characterization, and validation of NBS ... [https://www.frontiersin.org/journals/genetics/articles/10.3389/fgene.2023.1187597/full]